Filter
Associated Lab
- Baker Lab (1) Apply Baker Lab filter
- Betzig Lab (2) Apply Betzig Lab filter
- Harris Lab (3) Apply Harris Lab filter
- Hess Lab (2) Apply Hess Lab filter
- Rubin Lab (2) Apply Rubin Lab filter
- Spruston Lab (2) Apply Spruston Lab filter
- Stern Lab (2) Apply Stern Lab filter
- Tjian Lab (1) Apply Tjian Lab filter
- Truman Lab (1) Apply Truman Lab filter
Publication Date
- December 1994 (2) Apply December 1994 filter
- November 1994 (1) Apply November 1994 filter
- July 1994 (1) Apply July 1994 filter
- June 1994 (3) Apply June 1994 filter
- May 1994 (2) Apply May 1994 filter
- April 1994 (2) Apply April 1994 filter
- January 1994 (1) Apply January 1994 filter
- Remove 1994 filter 1994
Type of Publication
12 Publications
Showing 1-10 of 12 resultsThe gall-forming aphidCerataphis fransseni produces soldiers that defend against predators. Soldiers are produced soon after colony foundation and the number of soldiers increases nonlinearly during colony growth. The number of soldiers scales to the square-root of the number of non-soldiers and linearly to the surface area of the gall. This suggests that soldiers are produced to defend an area, for example the perimeter of the colony or the surface of the gall, rather than individual aphids.
In insects, the neuropeptide eclosion hormone (EH) acts on the CNS to evoke the stereotyped behaviors that cause ecdysis, the shedding of the cuticle at the end of each molt. Concomitantly, EH induces an increase in cyclic GMP (cGMP). Using antibodies against this second messenger, we show that this increase is confined to a network of 50 peptidergic neurons distributed throughout the CNS. Increases appeared 30 min after EH treatment, spread rapidly throughout these neurons, and were extremely long lived. We show that this response is synaptically driven, and does not involve the soluble, nitric oxide (NO)-activated, guanylate cyclase. Stereotyped variations in the duration of the cGMP response among neurons suggest a role in coordinating responses having different latencies and durations.
Given a relatively short query stringW of lengthP, a long subject stringA of lengthN, and a thresholdD, theapproximate keyword search problem is to find all substrings ofA that align withW with not more than D insertions, deletions, and mismatches. In typical applications, such as searching a DNA sequence database, the size of the “database”A is much larger than that of the queryW, e.g.,N is on the order of millions or billions andP is a hundred to a thousand. In this paper we present an algorithm that given a precomputedindex of the databaseA, finds rare matches in time that issublinear inN, i.e.,N c for somec<1. The sequenceA must be overa. finite alphabet σ. More precisely, our algorithm requires 0(DN pow(ɛ) logN) expected-time where ɛ=D/P is the maximum number of differences as a percentage of query length, and pow(ɛ) is an increasing and concave function that is 0 when ɛ=0. Thus the algorithm is superior to current O(DN) algorithms when ɛ is small enough to guarantee that pow(ɛ) < 1. As seen in the paper, this is true for a wide range of ɛ, e.g., ɛ. up to 33% for DNA sequences (¦⌆¦=4) and 56% for proteins sequences (¦⌆¦=20). In preliminary practical experiments, the approach gives a 50-to 500-fold improvement over previous algorithms for prolems of interest in molecular biology.
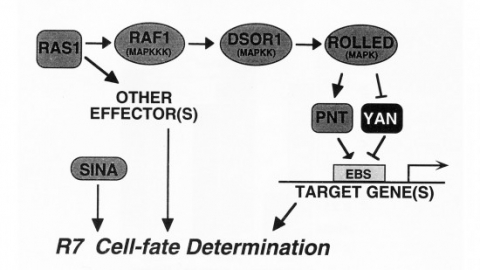
We show that the activities of two Ets-related transcription factors required for normal eye development in Drosophila, pointed and yan, are regulated by the Ras1/MAPK pathway. The pointed gene codes for two related proteins, and we show that one form is a constitutive activator of transcription, while the activity of the other form is stimulated by the Ras1/MAPK pathway. Mutation of the single consensus MAPK phosphorylation site in the second form abrogates this responsiveness. yan is a negative regulator of photoreceptor determination, and genetic data suggest that it acts as an antagonist of Ras1. We demonstrate that yan can repress transcription and that this repression activity is negatively regulated by the Ras1/MAPK signal, most likely through direct phosphorylation of yan by MAPK.
Luminescent centers with sharp (<0.07 millielectron volt), spectrally distinct emission lines were imaged in a GaAs/AIGaAs quantum well by means of low-temperature near-field scanning optical microscopy. Temperature, magnetic field, and linewidth measurements establish that these centers arise from excitons laterally localized at interface fluctuations. For sufficiently narrow wells, virtually all emission originates from such centers. Near-field microscopy/spectroscopy provides a means to access energies and homogeneous line widths for the individual eigenstates of these centers, and thus opens a rich area of physics involving quantum resolved systems.
Luminescent centers with sharp (<0.07 millielectron volt), spectrally distinct emission lines were imaged in a GaAs/AIGaAs quantum well by means of low-temperature near-field scanning optical microscopy. Temperature, magnetic field, and linewidth measurements establish that these centers arise from excitons laterally localized at interface fluctuations. For sufficiently narrow wells, virtually all emission originates from such centers. Near-field microscopy/spectroscopy provides a means to access energies and homogeneous line widths for the individual eigenstates of these centers, and thus opens a rich area of physics involving quantum resolved systems.
Commentary: Harald Hess and I joined forces, combining my near-field optical technology with his cryogenic scanned probe microscope to produce the first paper on high resolution spectroscopy beyond the diffraction limit. We discovered that the broad luminescence spectrum traditionally observed from quantum well heterostructures reflects a resolution-limited ensemble average of emission from numerous discrete sites of exciton recombination occurring at atomic-scale corrugations in the confining interfaces. With the combination of high spatial resolution from near-field excitation and high spectral resolution from cryogenic operation, we were able to isolate these emission sites in a multidimensional space of xy position and wavelength, even though their density was too great to isolate them on the basis of spatial resolution alone. This insight was very influential in the genesis of the concept (see above) that would eventually lead to far-field superresolution by PALM.
Excitatory postsynaptic currents in neurones of the central nervous system have a dual-component time course that results from the co-activation of AMPA/kainate-type and NMDA-type glutamate receptors. New approaches in electrophysiology and molecular biology have provided a better understanding of the factors that determine the kinetics of excitatory postsynaptic currents. Recent studies suggest that the time course of neurotransmitter concentration in the synaptic cleft, the gating properties of the native channels, and the glutamate receptor subunit composition all appear to be important factors.
Aphid soldiers, altruistic larvae that protect the colony from predators, are an example of highly social behaviour in an insect group with a natural history different from the eusocial Hymenoptera and Isoptera. Aphids therefore allow independent tests of theory developed to explain the evolution of eusociality. Although soldiers have been discovered in five tribes from two families, the number and pattern of origins and losses of soldiers is unknown due to a lack of phylogenetic data. Here I present a mtDNA based phylogeny for the Hormaphididae, and test the hypothesis that soldiers in the tribe Cerataphidini produced during two points in the life cycle represent independent origins. The results support this hypothesis. In addition, a minimum of five evolutionary events, either four origins and one loss or five origins, are required to explain the distribution of soldiers in the family. The positions of the origins and losses are well resolved, and this phylogeny provides an historical framework for studies on the causes of soldier aphid evolution.
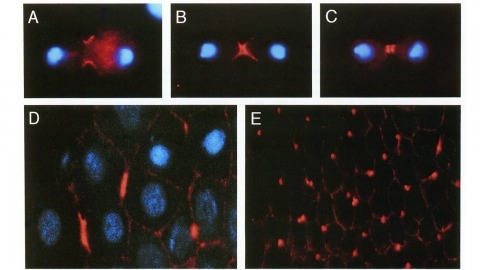
We have identified a Drosophila gene, peanut (pnut), that is related in sequence to the CDC3, CDC10, CDC11, and CDC12 genes of S. cerevisiae. These genes are required for cytokinesis, and their products are present at the bud neck during cell division. We find that pnut is also required for cytokinesis: in pnut mutants, imaginal tissues fail to proliferate and instead develop clusters of large, multinucleate cells. Pnut protein is localized to the cleavage furrow of dividing cells during cytokinesis and to the intercellular bridge connecting postmitotic daughter cells. In addition to its role in cytokinesis, pnut displays genetic interactions with seven in absentia, a gene required for neuronal fate determination in the compound eye, suggesting that pnut may have pleiotropic functions. Our results suggest that this class of proteins is involved in aspects of cytokinesis that have been conserved between flies and yeast.
X-ray absorption measurements from H-passivated porous Si and from oxidized Si nanocrystals, combined with electron microscopy, ir absorption, α recoil, and luminescence emission data, provide a consistent structural picture of the species responsible for the visible luminescence observed in these samples. The mass-weighted average structures in por-Si are particles, not wires, with dimensions significantly smaller than previously reported or proposed.