Filter
Associated Lab
- Ahrens Lab (1) Apply Ahrens Lab filter
- Baker Lab (2) Apply Baker Lab filter
- Betzig Lab (3) Apply Betzig Lab filter
- Cardona Lab (6) Apply Cardona Lab filter
- Chklovskii Lab (1) Apply Chklovskii Lab filter
- Dickson Lab (2) Apply Dickson Lab filter
- Druckmann Lab (1) Apply Druckmann Lab filter
- Eddy/Rivas Lab (3) Apply Eddy/Rivas Lab filter
- Fetter Lab (2) Apply Fetter Lab filter
- Gonen Lab (7) Apply Gonen Lab filter
- Grigorieff Lab (6) Apply Grigorieff Lab filter
- Harris Lab (1) Apply Harris Lab filter
- Heberlein Lab (5) Apply Heberlein Lab filter
- Hess Lab (2) Apply Hess Lab filter
- Jayaraman Lab (2) Apply Jayaraman Lab filter
- Ji Lab (1) Apply Ji Lab filter
- Kainmueller Lab (2) Apply Kainmueller Lab filter
- Keller Lab (4) Apply Keller Lab filter
- Lavis Lab (1) Apply Lavis Lab filter
- Lee (Albert) Lab (1) Apply Lee (Albert) Lab filter
- Lippincott-Schwartz Lab (12) Apply Lippincott-Schwartz Lab filter
- Looger Lab (7) Apply Looger Lab filter
- Magee Lab (3) Apply Magee Lab filter
- Pastalkova Lab (1) Apply Pastalkova Lab filter
- Reiser Lab (3) Apply Reiser Lab filter
- Riddiford Lab (2) Apply Riddiford Lab filter
- Romani Lab (1) Apply Romani Lab filter
- Rubin Lab (4) Apply Rubin Lab filter
- Saalfeld Lab (5) Apply Saalfeld Lab filter
- Satou Lab (2) Apply Satou Lab filter
- Scheffer Lab (3) Apply Scheffer Lab filter
- Schreiter Lab (2) Apply Schreiter Lab filter
- Shroff Lab (1) Apply Shroff Lab filter
- Simpson Lab (3) Apply Simpson Lab filter
- Singer Lab (1) Apply Singer Lab filter
- Spruston Lab (2) Apply Spruston Lab filter
- Stern Lab (8) Apply Stern Lab filter
- Sternson Lab (1) Apply Sternson Lab filter
- Svoboda Lab (7) Apply Svoboda Lab filter
- Tjian Lab (2) Apply Tjian Lab filter
- Truman Lab (4) Apply Truman Lab filter
- Turaga Lab (2) Apply Turaga Lab filter
- Turner Lab (1) Apply Turner Lab filter
- Zuker Lab (1) Apply Zuker Lab filter
Associated Project Team
Publication Date
- December 2010 (8) Apply December 2010 filter
- November 2010 (9) Apply November 2010 filter
- October 2010 (15) Apply October 2010 filter
- September 2010 (12) Apply September 2010 filter
- August 2010 (12) Apply August 2010 filter
- July 2010 (8) Apply July 2010 filter
- June 2010 (12) Apply June 2010 filter
- May 2010 (6) Apply May 2010 filter
- April 2010 (12) Apply April 2010 filter
- March 2010 (9) Apply March 2010 filter
- February 2010 (16) Apply February 2010 filter
- January 2010 (42) Apply January 2010 filter
- Remove 2010 filter 2010
Type of Publication
161 Publications
Showing 21-30 of 161 resultsThe Ndc80 complex is a key site of regulated kinetochore-microtubule attachment (a process required for cell division), but the molecular mechanism underlying its function remains unknown. Here we present a subnanometre-resolution cryo-electron microscopy reconstruction of the human Ndc80 complex bound to microtubules, sufficient for precise docking of crystal structures of the component proteins. We find that the Ndc80 complex binds the microtubule with a tubulin monomer repeat, recognizing α- and β-tubulin at both intra- and inter-tubulin dimer interfaces in a manner that is sensitive to tubulin conformation. Furthermore, Ndc80 complexes self-associate along protofilaments through interactions mediated by the amino-terminal tail of the NDC80 protein, which is the site of phospho-regulation by Aurora B kinase. The complex’s mode of interaction with the microtubule and its oligomerization suggest a mechanism by which Aurora B could regulate the stability of load-bearing kinetochore-microtubule attachments.
Proprioceptive sensory signals inform the CNS of the consequences of motor acts, but effective motor planning involves internal neural systems capable of anticipating actual sensory feedback. Just where and how predictive systems exert their influence remains poorly understood. We explored the possibility that spinocerebellar neurons that convey proprioceptive sensory information also integrate information from cortical command systems. Analysis of the circuitry and physiology of identified dorsal spinocerebellar tract neurons in mouse spinal cord revealed distinct populations of Clarke’s column neurons that received direct excitatory and/or indirect inhibitory inputs from descending corticospinal axons. The convergence of these descending inhibitory and excitatory inputs to Clarke’s column neurons established local spinal circuits with the capacity to mark or modulate incoming proprioceptive input. Together, our genetic, anatomical and physiological results indicate that Clarke’s column spinocerebellar neurons nucleate local spinal corollary circuits that are relevant to motor planning and evaluation.
This paper provides a compilation of diagrammatic representations of the expression profiles of epidermal and fat body mRNAs during the last two larval instars and metamorphosis of the tobacco hornworm, Manduca sexta. Included are those encoding insecticyanin, three larval cuticular proteins, dopa decarboxylase, moling, and the juvenile hormone-binding protein JP29 produced by the dorsal abdominal epidermis, and arylphorin and the methionine-rich storage proteins made by the fat body. The mRNA profiles of the ecdysteroid-regulated cascade of transcription factors in the epidermis during the larval molt and the onset of metamorphosis and in the pupal wing during the onset of adult development are also shown. These profiles are accompanied by a brief summary of the current knowledge about the regulation of these mRNAs by ecdysteroids and juvenile hormone based on experimental manipulations, both in vivo and in vitro.
The rapid adoption of high-throughput next generation sequence data in biological research is presenting a major challenge for sequence alignment tools—specifically, the efficient alignment of vast amounts of short reads to large references in the presence of differences arising from sequencing errors and biological sequence variations. To address this challenge, we developed a short read aligner for high-throughput sequencer data that is tolerant of errors or mutations of all types—namely, substitutions, deletions, and insertions. The aligner utilizes a multi-stage approach in which template-based indexing is used to identify candidate regions for alignment with dynamic programming. A template is a pair of gapped seeds, with one used with the read and one used with the reference. In this article, we focus on the development of template families that yield error-tolerant indexing up to a given error-budget. A general algorithm for finding those families is presented, and a recursive construction that creates families with higher error tolerance from ones with a lower error tolerance is developed.
Juvenile hormone (JH) signaling underpins both regulatory and developmental pathways in insects. However, the JH receptor is poorly understood. Methoprene tolerant (Met) and germ cell expressed (gce) have been implicated in JH signaling in Drosophila. We investigated the evolution of Met and gce across 12 Drosophila species and found that these paralogs are conserved across at least 63 million years of dipteran evolution. Distinct patterns of selection found using estimates of dN/dS ratios across Drosophila Met and gce coding sequences, along with their incongruent temporal expression profiles in embryonic Drosophila melanogaster, illustrate avenues through which these genes have diverged within the Diptera. Additionally, we demonstrate that the annotated gene CG15032 is the 5’ terminus of gce. In mosquitoes and beetles, a single Met-like homolog displays structural similarity to both Met and gce, and the intron locations are conserved with those of gce. We found that Tribolium and mosquito Met orthologs are assembled from Met- and gce-specific domains in a modular fashion. Our results suggest that Drosophila Met and gce experienced divergent evolutionary pressures following the duplication of an ancestral gce-like gene found in less derived holometabolous insects.
An estimated 2 million Americans use cocaine, resulting in large personal and societal costs. Discovery of the genetic factors that contribute to cocaine abuse is important for understanding this complex disease. Previously, mutations in the Drosophila LIM-only (dLmo) gene were identified because of their increased behavioral sensitivity to cocaine. Here we show that the mammalian homolog Lmo4, which is highly expressed in brain regions implicated in drug addiction, plays a similar role in cocaine-induced behaviors. Mice with a global reduction in Lmo4 levels show increased sensitivity to the locomotor stimulatory effects of cocaine upon chronic cocaine administration. This effect is reproduced with downregulation of Lmo4 in the nucleus accumbens by RNA interference. Thus, Lmo genes play conserved roles in regulating the behavioral effects of cocaine in invertebrate and mammalian models of drug addiction.
Connections between neurons can be found by checking whether synapses exist at points of contact, which in turn are determined by neural shapes. Finding these shapes is a special case of image segmentation, which is laborious for humans and would ideally be performed by computers. New metrics properly quantify the performance of a computer algorithm using its disagreement with ’true’ segmentations of example images. New machine learning methods search for segmentation algorithms that minimize such metrics. These advances have reduced computer errors dramatically. It should now be faster for a human to correct the remaining errors than to segment an image manually. Further reductions in human effort are expected, and crucial for finding connectomes more complex than that of Caenorhabditis elegans.
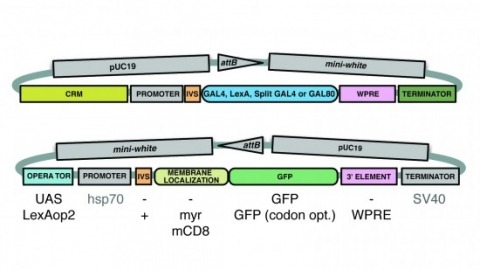
A wide variety of biological experiments rely on the ability to express an exogenous gene in a transgenic animal at a defined level and in a spatially and temporally controlled pattern. We describe major improvements of the methods available for achieving this objective in Drosophila melanogaster. We have systematically varied core promoters, UTRs, operator sequences, and transcriptional activating domains used to direct gene expression with the GAL4, LexA, and Split GAL4 transcription factors and the GAL80 transcriptional repressor. The use of site-specific integration allowed us to make quantitative comparisons between different constructs inserted at the same genomic location. We also characterized a set of PhiC31 integration sites for their ability to support transgene expression of both drivers and responders in the nervous system. The increased strength and reliability of these optimized reagents overcome many of the previous limitations of these methods and will facilitate genetic manipulations of greater complexity and sophistication.
AMP-activated protein kinase (AMPK), a cellular metabolic sensor, is essential in energy regulation and metabolism. Hepatocyte polarization during liver development and regeneration parallels increased metabolism. The current study investigates the effects of AMPK and its upstream activator LKB1 on polarity and bile canalicular network formation and maintenance in collagen sandwich cultures of rat hepatocytes. Immunostaining for the apical protein ABCB1 and the tight junction marker occludin demonstrated that canalicular network formation is sequential and is associated with activation of AMPK and LKB1. AMPK and LKB1 activators accelerated canalicular network formation. Inhibition of AMPK or LKB1 by dominant-negative AMPK or kinase-dead LKB1 constructs blocked canalicular network formation. AICAR and 2-deoxyglucose, which activate AMPK, circumvented the inhibitory effect of kinase-dead LKB1 on canalicular formation, indicating that AMPK directly affects canalicular network formation. After the canalicular network was formed, inhibition of AMPK and LKB1 by dominant-negative AMPK or kinase-dead LKB1 constructs resulted in loss of canalicular network, indicating that AMPK and LKB1 also participate in network maintenance. In addition, activation of AMPK and LKB1 prevented low-Ca(2+)-mediated disruption of the canalicular network and tight junctions. These studies reveal that AMPK and its upstream kinase, LKB1, regulate canalicular network formation and maintenance.
Reconstructing neuronal circuits at the level of synapses is a central problem in neuroscience, and the focus of the nascent field of connectomics. Previously used to reconstruct the C. elegans wiring diagram, serial-section transmission electron microscopy (ssTEM) is a proven technique for the task. However, to reconstruct more complex circuits, ssTEM will require the automation of image processing. We review progress in the processing of electron microscopy images and, in particular, a semi-automated reconstruction pipeline deployed at Janelia. Drosophila circuits underlying identified behaviors are being reconstructed in the pipeline with the goal of generating a complete Drosophila connectome.