Filter
Associated Lab
- Ahrens Lab (7) Apply Ahrens Lab filter
- Druckmann Lab (2) Apply Druckmann Lab filter
- Dudman Lab (1) Apply Dudman Lab filter
- Freeman Lab (2) Apply Freeman Lab filter
- Harris Lab (7) Apply Harris Lab filter
- Hermundstad Lab (1) Apply Hermundstad Lab filter
- Hess Lab (2) Apply Hess Lab filter
- Jayaraman Lab (10) Apply Jayaraman Lab filter
- Karpova Lab (2) Apply Karpova Lab filter
- Keller Lab (5) Apply Keller Lab filter
- Lavis Lab (8) Apply Lavis Lab filter
- Leonardo Lab (4) Apply Leonardo Lab filter
- Remove Looger Lab filter Looger Lab
- Podgorski Lab (6) Apply Podgorski Lab filter
- Rubin Lab (2) Apply Rubin Lab filter
- Schreiter Lab (24) Apply Schreiter Lab filter
- Simpson Lab (1) Apply Simpson Lab filter
- Spruston Lab (1) Apply Spruston Lab filter
- Sternson Lab (2) Apply Sternson Lab filter
- Svoboda Lab (20) Apply Svoboda Lab filter
- Tervo Lab (1) Apply Tervo Lab filter
- Tillberg Lab (1) Apply Tillberg Lab filter
- Turner Lab (1) Apply Turner Lab filter
- Zlatic Lab (1) Apply Zlatic Lab filter
Associated Project Team
Publication Date
- 2024 (1) Apply 2024 filter
- 2023 (5) Apply 2023 filter
- 2022 (7) Apply 2022 filter
- 2021 (11) Apply 2021 filter
- 2020 (7) Apply 2020 filter
- 2019 (15) Apply 2019 filter
- 2018 (8) Apply 2018 filter
- 2017 (6) Apply 2017 filter
- 2016 (10) Apply 2016 filter
- 2015 (9) Apply 2015 filter
- 2014 (11) Apply 2014 filter
- 2013 (10) Apply 2013 filter
- 2012 (13) Apply 2012 filter
- 2011 (7) Apply 2011 filter
- 2010 (7) Apply 2010 filter
- 2009 (7) Apply 2009 filter
- 2008 (3) Apply 2008 filter
Type of Publication
137 Publications
Showing 111-120 of 137 resultsRecording activity from identified populations of neurons is a central goal of neuroscience. Changes in membrane depolarization, particularly action potentials, are the most important features of neural physiology to extract, although ions, neurotransmitters, neuromodulators, second messengers, and the activation state of specific proteins are also crucial. Modern fluorescence microscopy provides the basis for such activity mapping, through multi-photon imaging and other optical schemes. Probes remain the rate-limiting step for progress in this field: they should be bright and photostable, and ideally come in multiple colors. Only protein-based reagents permit chronic imaging from genetically specified cells. Here we review recent progress in the design, optimization and deployment of genetically encoded indicators for calcium ions (a proxy for action potentials), membrane potential, and neurotransmitters. We highlight seminal experiments, and present an outlook for future progress.
Genetically encoded calcium indicators (GECIs), together with modern microscopy, allow repeated activity measurement, in real time and with cellular resolution, of defined cellular populations. Recent efforts in protein engineering have yielded several high-quality GECIs that facilitate new applications in neuroscience. Here, we summarize recent progress in GECI design, optimization, and characterization, and provide guidelines for selecting the appropriate GECI for a given biological application. We focus on the unique challenges associated with imaging in behaving animals.
The phnD gene of Escherichia coli encodes the periplasmic binding protein of the phosphonate (Pn) uptake and utilization pathway. We have crystallized and determined structures of E. coli PhnD (EcPhnD) in the absence of ligand and in complex with the environmentally abundant 2-aminoethylphosphonate (2AEP). Similar to other bacterial periplasmic binding proteins, 2AEP binds near the center of mass of EcPhnD in a cleft formed between two lobes. Comparison of the open, unliganded structure with the closed 2AEP-bound structure shows that the two lobes pivot around a hinge by \~{}70° between the two states. Extensive hydrogen bonding and electrostatic interactions stabilize 2AEP, which binds to EcPhnD with low nanomolar affinity. These structures provide insight into Pn uptake by bacteria and facilitated the rational design of high signal-to-noise Pn biosensors based on both coupled small-molecule dyes and autocatalytic fluorescent proteins.
We describe the generation of a family of high-signal-to-noise single-wavelength genetically encoded indicators for maltose. This was achieved by insertion of circularly permuted fluorescent proteins into a bacterial periplasmic binding protein (PBP), Escherichia coli maltodextrin-binding protein, resulting in a four-color family of maltose indicators. The sensors were iteratively optimized to have sufficient brightness and maltose-dependent fluorescence increases for imaging, under both one- and two-photon illumination. We demonstrate that maltose affinity of the sensors can be tuned in a fashion largely independent of the fluorescent readout mechanism. Using literature mutations, the binding specificity could be altered to moderate sucrose preference, but with a significant loss of affinity. We use the soluble sensors in individual E. coli bacteria to observe rapid maltose transport across the plasma membrane, and membrane fusion versions of the sensors on mammalian cells to visualize the addition of maltose to extracellular media. The PBP superfamily includes scaffolds specific for a number of analytes whose visualization would be critical to the reverse engineering of complex systems such as neural networks, biosynthetic pathways, and signal transduction cascades. We expect the methodology outlined here to be useful in the development of indicators for many such analytes.
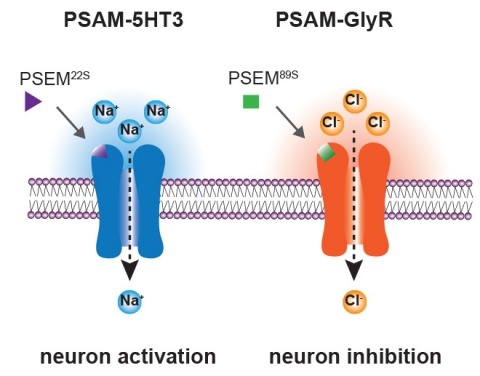
Ionic flux mediates essential physiological and behavioral functions in defined cell populations. Cell type-specific activators of diverse ionic conductances are needed for probing these effects. We combined chemistry and protein engineering to enable the systematic creation of a toolbox of ligand-gated ion channels (LGICs) with orthogonal pharmacologic selectivity and divergent functional properties. The LGICs and their small-molecule effectors were able to activate a range of ionic conductances in genetically specified cell types. LGICs constructed for neuronal perturbation could be used to selectively manipulate neuron activity in mammalian brains in vivo. The diversity of ion channel tools accessible from this approach will be useful for examining the relationship between neuronal activity and animal behavior, as well as for cell biological and physiological applications requiring chemical control of ion conductance.
Multiphoton imaging (MPI) is widely used for recording activity simultaneously from many neurons in superficial cortical layers in vivo. We combined regenerative amplification multiphoton microscopy (RAMM) with genetically encoded calcium indicators to extend MPI of neuronal population activity into layer 5 (L5) of adult mouse somatosensory cortex. We found that this approach could be used to record and quantify spontaneous and sensory-evoked activity in populations of L5 neuronal somata located as much as 800 μm below the pia. In addition, we found that RAMM could be used to simultaneously image activity from large (80) populations of apical dendrites and follow these dendrites down to their somata of origin.
We developed a multicolor neuron labeling technique in Drosophila melanogaster that combines the power to specifically target different neural populations with the label diversity provided by stochastic color choice. This adaptation of vertebrate Brainbow uses recombination to select one of three epitope-tagged proteins detectable by immunofluorescence. Two copies of this construct yield six bright, separable colors. We used Drosophila Brainbow to study the innervation patterns of multiple antennal lobe projection neuron lineages in the same preparation and to observe the relative trajectories of individual aminergic neurons. Nerve bundles, and even individual neurites hundreds of micrometers long, can be followed with definitive color labeling. We traced motor neurons in the subesophageal ganglion and correlated them to neuromuscular junctions to identify their specific proboscis muscle targets. The ability to independently visualize multiple lineage or neuron projections in the same preparation greatly advances the goal of mapping how neurons connect into circuits.
Decoding the wiring diagram of the retina requires simultaneous observation of activity in identified neuron populations. Available recording methods are limited in their scope: electrodes can access only a small fraction of neurons at once, whereas synthetic fluorescent indicator dyes label tissue indiscriminately. Here, we describe a method for studying retinal circuitry at cellular and subcellular levels combining two-photon microscopy and a genetically encoded calcium indicator. Using specific viral and promoter constructs to drive expression of GCaMP3, we labeled all five major neuron classes in the adult mouse retina. Stimulus-evoked GCaMP3 responses as imaged by two-photon microscopy permitted functional cell type annotation. Fluorescence responses were similar to those measured with the small molecule dye OGB-1. Fluorescence intensity correlated linearly with spike rates >10 spikes/s, and a significant change in fluorescence always reflected a significant change in spike firing rate. GCaMP3 expression had no apparent effect on neuronal function. Imaging at subcellular resolution showed compartment-specific calcium dynamics in multiple identified cell types.
Genetically encoded calcium indicators (GECIs), which are based on chimeric fluorescent proteins, can be used to monitor calcium transients in living cells and organisms. Because they are encoded by DNA, GECIs can be delivered to the intact brain noninvasively and targeted to defined populations of neurons and specific subcellular compartments for long-term, repeated measurements in vivo. GECIs have improved iteratively and are becoming useful for imaging neural activity in vivo. Here we summarize extrinsic and intrinsic factors that influence a GECI's performance and provides guidelines for selecting the appropriate GECI for a given application. We also review recent progress in GECI design, optimization, and standardized testing protocols.