Filter
Associated Lab
- Aso Lab (29) Apply Aso Lab filter
- Betzig Lab (1) Apply Betzig Lab filter
- Bock Lab (2) Apply Bock Lab filter
- Branson Lab (7) Apply Branson Lab filter
- Card Lab (5) Apply Card Lab filter
- Clapham Lab (1) Apply Clapham Lab filter
- Dickson Lab (2) Apply Dickson Lab filter
- Druckmann Lab (1) Apply Druckmann Lab filter
- Fetter Lab (1) Apply Fetter Lab filter
- Funke Lab (1) Apply Funke Lab filter
- Harris Lab (3) Apply Harris Lab filter
- Heberlein Lab (2) Apply Heberlein Lab filter
- Hermundstad Lab (2) Apply Hermundstad Lab filter
- Hess Lab (5) Apply Hess Lab filter
- Jayaraman Lab (5) Apply Jayaraman Lab filter
- Lippincott-Schwartz Lab (1) Apply Lippincott-Schwartz Lab filter
- Looger Lab (2) Apply Looger Lab filter
- O'Shea Lab (1) Apply O'Shea Lab filter
- Otopalik Lab (1) Apply Otopalik Lab filter
- Reiser Lab (14) Apply Reiser Lab filter
- Riddiford Lab (1) Apply Riddiford Lab filter
- Romani Lab (1) Apply Romani Lab filter
- Remove Rubin Lab filter Rubin Lab
- Saalfeld Lab (4) Apply Saalfeld Lab filter
- Scheffer Lab (7) Apply Scheffer Lab filter
- Schreiter Lab (1) Apply Schreiter Lab filter
- Simpson Lab (3) Apply Simpson Lab filter
- Singer Lab (1) Apply Singer Lab filter
- Spruston Lab (1) Apply Spruston Lab filter
- Stern Lab (1) Apply Stern Lab filter
- Svoboda Lab (3) Apply Svoboda Lab filter
- Tjian Lab (1) Apply Tjian Lab filter
- Truman Lab (4) Apply Truman Lab filter
- Turaga Lab (1) Apply Turaga Lab filter
- Turner Lab (5) Apply Turner Lab filter
- Zuker Lab (1) Apply Zuker Lab filter
Associated Project Team
Publication Date
- 2024 (2) Apply 2024 filter
- 2023 (6) Apply 2023 filter
- 2022 (1) Apply 2022 filter
- 2021 (4) Apply 2021 filter
- 2020 (9) Apply 2020 filter
- 2019 (6) Apply 2019 filter
- 2018 (7) Apply 2018 filter
- 2017 (15) Apply 2017 filter
- 2016 (3) Apply 2016 filter
- 2015 (16) Apply 2015 filter
- 2014 (9) Apply 2014 filter
- 2013 (5) Apply 2013 filter
- 2012 (8) Apply 2012 filter
- 2011 (4) Apply 2011 filter
- 2010 (4) Apply 2010 filter
- 2009 (2) Apply 2009 filter
- 2008 (4) Apply 2008 filter
- 2007 (2) Apply 2007 filter
- 2006 (1) Apply 2006 filter
- 2002 (1) Apply 2002 filter
- 2000 (2) Apply 2000 filter
- 1999 (1) Apply 1999 filter
- 1997 (1) Apply 1997 filter
- 1995 (2) Apply 1995 filter
- 1994 (2) Apply 1994 filter
- 1993 (2) Apply 1993 filter
- 1992 (1) Apply 1992 filter
- 1991 (2) Apply 1991 filter
- 1990 (3) Apply 1990 filter
- 1989 (2) Apply 1989 filter
- 1987 (2) Apply 1987 filter
- 1986 (1) Apply 1986 filter
- 1985 (1) Apply 1985 filter
- 1984 (1) Apply 1984 filter
- 1983 (1) Apply 1983 filter
- 1982 (2) Apply 1982 filter
- 1981 (1) Apply 1981 filter
- 1979 (1) Apply 1979 filter
- 1973 (1) Apply 1973 filter
Type of Publication
138 Publications
Showing 61-70 of 138 resultsDrosophila central neurons arise from neuroblasts that generate neurons in a pair-wise fashion, with the two daughters providing the basis for distinct A and B hemilineage groups. Thirty three postembryonically-born hemilineages contribute over 90% of the neurons in each thoracic hemisegment. We devised genetic approaches to define the anatomy of most of these hemilineages and to assessed their functional roles using the heat-sensitive channel dTRPA1. The simplest hemilineages contained local interneurons and their activation caused tonic or phasic leg movements lacking interlimb coordination. The next level was hemilineages of similar projection cells that drove intersegmentally coordinated behaviors such as walking. The highest level involved hemilineages whose activation elicited complex behaviors such as takeoff. These activation phenotypes indicate that the hemilineages vary in their behavioral roles with some contributing to local networks for sensorimotor processing and others having higher order functions of coordinating these local networks into complex behavior.
When navigating in their environment, animals use visual motion cues as feedback signals that are elicited by their own motion. Such signals are provided by wide-field neurons sampling motion directions at multiple image points as the animal maneuvers. Each one of these neurons responds selectively to a specific optic flow-field representing the spatial distribution of motion vectors on the retina. Here, we describe the discovery of a group of local, inhibitory interneurons in the fruit fly Drosophila key for filtering these cues. Using anatomy, molecular characterization, activity manipulation, and physiological recordings, we demonstrate that these interneurons convey direction-selective inhibition to wide-field neurons with opposite preferred direction and provide evidence for how their connectivity enables the computation required for integrating opposing motions. Our results indicate that, rather than sharpening directional selectivity per se, these circuit elements reduce noise by eliminating non-specific responses to complex visual information.
•Discovery of bi-stratified glutamatergic lobula plate-intrinsic (LPi) interneurons•LPi neurons provide visual null direction inhibition to wide-field tangential cells•Blocking LPi activity leads to target neurons responding to inadequate motion cues•Motion opponency thus increases flow-field selectivity
Newly identified inhibitory neurons are central to an integrative circuit that enables Drosophila to process visual cues with opposite motions generated during flight. The neurons are required to discriminate between distinct complex motion patterns, indicating that neural processing of opposing cues can yield outcomes beyond the simple sum of two inputs.
Taste memories allow animals to modulate feeding behavior in accordance with past experience and avoid the consumption of potentially harmful food [1]. We have developed a single-fly taste memory assay to functionally interrogate the neural circuitry encoding taste memories [2]. Here, we screen a collection of Split-GAL4 lines that label small populations of neurons associated with the fly memory center-the mushroom bodies (MBs) [3]. Genetic silencing of PPL1 dopamine neurons disrupts conditioned, but not naive, feeding behavior, suggesting these neurons are selectively involved in the conditioned taste response. We identify two PPL1 subpopulations that innervate the MB α lobe and are essential for aversive taste memory. Thermogenetic activation of these dopamine neurons during training induces memory, indicating these neurons are sufficient for the reinforcing properties of bitter tastant to the MBs. Silencing of either the intrinsic MB neurons or the output neurons from the α lobe disrupts taste conditioning. Thermogenetic manipulation of these output neurons alters naive feeding response, suggesting that dopamine neurons modulate the threshold of response to appetitive tastants. Taken together, these findings detail a neural mechanism underlying the formation of taste memory and provide a functional model for dopamine-dependent plasticity in Drosophila.
We describe the development and application of methods for high-throughput neuroanatomy in Drosophila using light microscopy. These tools enable efficient multicolor stochastic labeling of neurons at both low and high densities. Expression of multiple membrane-targeted and distinct epitope-tagged proteins is controlled both by a transcriptional driver and by stochastic, recombinase-mediated excision of transcription-terminating cassettes. This MultiColor FlpOut (MCFO) approach can be used to reveal cell shapes and relative cell positions and to track the progeny of precursor cells through development. Using two different recombinases, the number of cells labeled and the number of color combinations observed in those cells can be controlled separately. We demonstrate the utility of MCFO in a detailed study of diversity and variability of Distal medulla (Dm) neurons, multicolumnar local interneurons in the adult visual system. Similar to many brain regions, the medulla has a repetitive columnar structure that supports parallel information processing together with orthogonal layers of cell processes that enable communication between columns. We find that, within a medulla layer, processes of the cells of a given Dm neuron type form distinct patterns that reflect both the morphology of individual cells and the relative positions of their arbors. These stereotyped cell arrangements differ between cell types and can even differ for the processes of the same cell type in different medulla layers. This unexpected diversity of coverage patterns provides multiple independent ways of integrating visual information across the retinotopic columns and implies the existence of multiple developmental mechanisms that generate these distinct patterns.
Insects exhibit an elaborate repertoire of behaviors in response to environmental stimuli. The central complex plays a key role in combining various modalities of sensory information with an insect's internal state and past experience to select appropriate responses. Progress has been made in understanding the broad spectrum of outputs from the central complex neuropils and circuits involved in numerous behaviors. Many resident neurons have also been identified. However, the specific roles of these intricate structures and the functional connections between them remain largely obscure. Significant gains rely on obtaining a comprehensive catalog of the neurons and associated GAL4 lines that arborize within these brain regions, and on mapping neuronal pathways connecting these structures. To this end, small populations of neurons in the Drosophila melanogaster central complex were stochastically labeled using the multicolor flip-out technique and a catalog was created of the neurons, their morphologies, trajectories, relative arrangements, and corresponding GAL4 lines. This report focuses on one structure of the central complex, the protocerebral bridge, and identifies just 17 morphologically distinct cell types that arborize in this structure. This work also provides new insights into the anatomical structure of the four components of the central complex and its accessory neuropils. Most strikingly, we found that the protocerebral bridge contains 18 glomeruli, not 16, as previously believed. Revised wiring diagrams that take into account this updated architectural design are presented. This updated map of the Drosophila central complex will facilitate a deeper behavioral and physiological dissection of this sophisticated set of structures. J. Comp. Neurol. 523:997-1037, 2015. © 2014 Wiley Periodicals, Inc.
We describe an engineered family of highly antigenic molecules based on GFP-like fluorescent proteins. These molecules contain numerous copies of peptide epitopes and simultaneously bind IgG antibodies at each location. These 'spaghetti monster' fluorescent proteins (smFPs) distributed well in neurons, notably into small dendrites, spines and axons. smFP immunolabeling localized weakly expressed proteins not well resolved with traditional epitope tags. By varying epitope and scaffold, we generated a diverse family of mutually orthogonal antigens. In cultured neurons and mouse and fly brains, smFP probes allowed robust, orthogonal multicolor visualization of proteins, cell populations and neuropil. smFP variants complement existing tracers and greatly increase the number of simultaneous imaging channels, and they performed well in advanced preparations such as array tomography, super-resolution fluorescence imaging and electron microscopy. In living cells, the probes improved single-molecule image tracking and increased yield for RNA-seq. These probes facilitate new experiments in connectomics, transcriptomics and protein localization.
Ends-out gene targeting allows seamless replacement of endogenous genes with engineered DNA fragments by homologous recombination, thus creating designer "genes" in the endogenous locus. Conventional gene targeting in Drosophila involves targeting with the preintegrated donor DNA in the larval primordial germ cells. Here we report G: ene targeting during O: ogenesis with L: ethality I: nhibitor and C: RISPR/Cas (Golic+), which improves on all major steps in such transgene-based gene targeting systems. First, donor DNA is integrated into precharacterized attP sites for efficient flip-out. Second, FLP, I-SceI, and Cas9 are specifically expressed in cystoblasts, which arise continuously from female germline stem cells, thereby providing a continual source of independent targeting events in each offspring. Third, a repressor-based lethality selection is implemented to facilitate screening for correct targeting events. Altogether, Golic+ realizes high-efficiency ends-out gene targeting in ovarian cystoblasts, which can be readily scaled up to achieve high-throughput genome editing.
Drosophila melanogaster plays an important role in molecular, genetic, and genomic studies of heredity, development, metabolism, behavior, and human disease. The initial reference genome sequence reported more than a decade ago had a profound impact on progress in Drosophila research, and improving the accuracy and completeness of this sequence continues to be important to further progress. We previously described improvement of the 117-Mb sequence in the euchromatic portion of the genome and 21 Mb in the heterochromatic portion, using a whole-genome shotgun assembly, BAC physical mapping, and clone-based finishing. Here, we report an improved reference sequence of the single-copy and middle-repetitive regions of the genome, produced using cytogenetic mapping to mitotic and polytene chromosomes, clone-based finishing and BAC fingerprint verification, ordering of scaffolds by alignment to cDNA sequences, incorporation of other map and sequence data, and validation by whole-genome optical restriction mapping. These data substantially improve the accuracy and completeness of the reference sequence and the order and orientation of sequence scaffolds into chromosome arm assemblies. Representation of the Y chromosome and other heterochromatic regions is particularly improved. The new 143.9-Mb reference sequence, designated Release 6, effectively exhausts clone-based technologies for mapping and sequencing. Highly repeat-rich regions, including large satellite blocks and functional elements such as the ribosomal RNA genes and the centromeres, are largely inaccessible to current sequencing and assembly methods and remain poorly represented. Further significant improvements will require sequencing technologies that do not depend on molecular cloning and that produce very long reads.
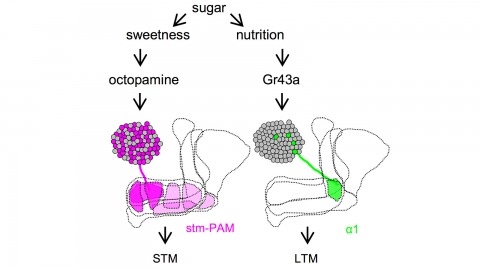
Drosophila melanogaster can acquire a stable appetitive olfactory memory when the presentation of a sugar reward and an odor are paired. However, the neuronal mechanisms by which a single training induces long-term memory are poorly understood. Here we show that two distinct subsets of dopamine neurons in the fly brain signal reward for short-term (STM) and long-term memories (LTM). One subset induces memory that decays within several hours, whereas the other induces memory that gradually develops after training. They convey reward signals to spatially segregated synaptic domains of the mushroom body (MB), a potential site for convergence. Furthermore, we identified a single type of dopamine neuron that conveys the reward signal to restricted subdomains of the mushroom body lobes and induces long-term memory. Constant appetitive memory retention after a single training session thus comprises two memory components triggered by distinct dopamine neurons.
Animals discriminate stimuli, learn their predictive value and use this knowledge to modify their behavior. In Drosophila, the mushroom body (MB) plays a key role in these processes. Sensory stimuli are sparsely represented by ∼2000 Kenyon cells, which converge onto 34 output neurons (MBONs) of 21 types. We studied the role of MBONs in several associative learning tasks and in sleep regulation, revealing the extent to which information flow is segregated into distinct channels and suggesting possible roles for the multi-layered MBON network. We also show that optogenetic activation of MBONs can, depending on cell type, induce repulsion or attraction in flies. The behavioral effects of MBON perturbation are combinatorial, suggesting that the MBON ensemble collectively represents valence. We propose that local, stimulus-specific dopaminergic modulation selectively alters the balance within the MBON network for those stimuli. Our results suggest that valence encoded by the MBON ensemble biases memory-based action selection.