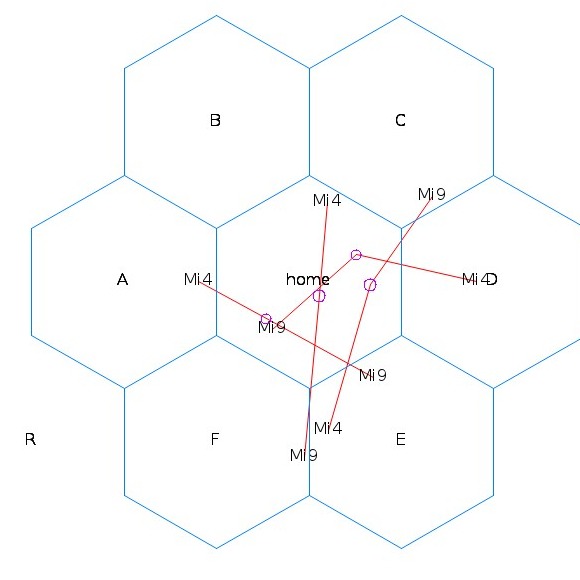
Lou Scheffer is extending techniques developed for electrical engineering and the design of chips to the understanding of the construction, development, and operation of the nervous system.
The main goal of our lab is to be able to quickly and reliably extract the detailed structure and connections from a volume of neural tissue. Biologically, this capability could be used to understand how the nervous system works, in detail; to figure out how it develops into its adult form; and to compare different animals and genetic variants.
The reconstruction consists of automated analysis of electron microscopy (EM) photographs, followed by manual proofreading. The proofreading step involves considerable manual work and represents the bulk of the person-hours expended in reconstruction. This makes it the limiting factor in the size of organism we can reconstruct.
Several steps are needed to improve the reconstruction of large biological neural systems:
Replace hard decisions on limited scope with soft decisions and global optimization, based on techniques from communication and coding theory.
Incorporate biological knowledge into the recjfonstruction process. Much of this information comes from techniques of genetics and light microscopy, which are very different in terms of the types, scales, and resolution of the data returned. This disparate data must be reconciled before it can help in reconstruction. Improve the user interface and productivity of manual proofreading, based on engineering experience with graphical systems for integrated circuit design and verification.
Adopt and adapt techniques developed for chip design to neural reconstruction. A project with many people working on a common problem, with millions of components, will involve many factors other than just algorithms. Methodologies that use existing tools, the databases used, the division of labor, and even mundane details like naming conventions, can have a large effect on a project of the scale of the proposed reconstruction. Chip designers have considerable experience with these issues, and we should take advantage of this experience. In addition, there are many potential avenues that have not yet been explored well enough to determine whether they would be helpful.
All these improvements, and more not yet thought of, will be needed, as the overall goal is the reconstruction and understanding of nervous systems, many of which are orders of magnitude larger than anything yet attempted. There is some hope for such efforts, since gains in productivity have been made in electrical engineering (EE), dealing with broadly similar structures of components and connections between them. In the EE field, large gains in productivity (50x or more) were obtained as a sequence of smaller innovations, and I expect a similar course of events here.
Rebuild Reconstruction Infrastructure Based on Probability
Right now all reconstruction decisions are binary—either two regions are part of the same cell, or they are not. Either two features detected in adjacent sections are connected, or they are not. The rest of the reconstruction process proceeds from the results of these decisions.
However, this approach throws away considerable information. Finding the underlying structure will require revisiting some of the early classification decisions. Intuitively, it's much better to overturn a dubious decision than one that is very solid. In a field such as communication theory, this intuition can be quantified and analyzed. As a result, your cell phone, for example, uses techniques such as Viterbi decoding that propagate the uncertainties throughout the process to find the most likely overall result.
The first step is to express the relationships between the detected objects in probabilistic terms. Then algorithms such as belief propagation, used in error-correcting codes, can be used to find the most likely underlying structure that would yield the known measurement. This can be applied at all stages of the reconstruction: What are the odds that two adjacent features are part of the same neuron? What are the odds that a feature detected in two consecutive sections belongs to the same biological structure? Also, the probabilistic structure provides an easy way to add biological and human input. It also provides a way to concentrate proofreading on the regions where the uncertainty is greatest, or where the gain from disambiguation is highest.
The primary mechanism that must be developed is incremental and real-time maintenance of the linkage map, which defines how features detected in each image are combined into biological elements such as neurons. For example, when a human proofreader states that two structures are definitely part of the same cell, these changes should then propagate through the rest of the reconstruction, as the system tries to find a configuration that incorporates this new data.
Incorporate Biological Knowledge
Many improvements to the initial reconstruction can be driven by the incorporation of biological knowledge into the reconstruction process. The different topologies, sizes, and shapes of neuron types have been known since the days of Golgi. This knowledge is rapidly improving, from both recent EM reconstructions and genetic techniques combined with sophisticated optical microscopy. Descriptions of these known neuron types must be entered in a machine-readable form so the software can automatically classify many of the neurons it encounters and identify the ones it cannot recognize. Before proofreading, each neuron then gets a tag, depending on the closest match (and the confidence of that match). After proofreading, we have much more quantitative data about each neuron—this will allow us to update the templates and make future matches that are more accurate.
Furthermore, from partial reconstructions we know some of which types of neurons are associated in various ways (e.g., adjacent in the same bundle, terminating in the same layers). This too can be used to help reconstruction. There are also more general biological principles—mitochondria are entirely contained within cells, each cell contains exactly one nucleus—that can help in reconstruction. These can also be handled within the probabilistic framework.
The next step, once we have identified and labeled all the neurons, is to find all connections between them (the synapses). This is now done by biologists and proofreaders working from existing EM photographs, and the appearance of a synapse depends on the animal studied. This step needs to be automated to tackle the larger systems of interest.
The final linkage should include all the biological elements we can detect. Although the main goal is neurons and synapses, location of the other machinery within the cell may be significant as well. Rather than just finding and removing mitochondria and vesicles, we should link these additional structures to the final neuron configuration.
Increase Productivity in Proofreading
Proofreading is currently the rate-limiting step in reconstruction and likely to remain so, since other tasks such as image acquisition and computer analysis are easily parallelized (and thus can be sped up as needed, subject only to budget constraints). There is a considerable body of knowledge and experience in organizing and improving interactive graphical tasks performed on large data sets. This can be applied to the proofreading software. Techniques include recording and reconstruction of user sessions, statistical methods to determine time spent on various tasks, measurement of the associated error rates, and ergonomic studies examining the effect of different input and display devices. Areas that can be improved include the use of visual clues such as color, brightness, spacing, text versus graphical input, customization, realistic versus symbolic displays, and menu and command structure. Both ease of learning by a new user and productivity of an expert user are important for this application.
Even without formal study, it is clear the process can be much improved by, for example, easy tagging of neuron types and use of color to indicate reconstruction status. Also, the probabilistic infrastructure should be used to order user decisions by overall impact and re-evaluate decisions on areas not yet proofread based on the best available information.
Use Engineering Experience from Chip Design
A project the size of reconstructing a fly's brain (perhaps 100 million connections) requires more than just algorithms. It is going to need many people, probably distributed over multiple sites. This raises a number of engineering questions, largely unrelated to the scientific problems.
How is the work to be divided? How and where are the data stored, and how do you prevent two users from making conflicting changes to the same data? How are software revisions handled? What network connectivity is required, and what happens if the network goes down? When one collaborator needs the result from another's work, what do they see—the latest version, which might be in progress, or some previously stable version? How can they choose, if they need to? Even seemingly minor issues, such as naming conventions, can create havoc if not considered early.
Chip design has run into all these problems, and many other related ones. A good fraction of the 3–4 billion dollar EDA (electronic design automation) industry is devoted to solving problems of this nature. As the reconstruction projects scale up, they should study the experience of those who have struggled to work with systems of similar size and complexity. Many of the solutions to these problems that have been adapted in the world of integrated circuits will probably be applicable to neural reconstruction, either directly or by analogy.
More-Speculative Improvements
The above improvements are almost certain to work, and we are attacking them first. However, there are many other possibilities for improving the reconstruction process that are more speculative but may be helpful, and we are investigating these as well. Many of these have been discussed in Dmitri Chklovskii's (HHMI, Janelia) group, and include tomographic or slant views, combined optical and EM imaging of the same samples, and tagging of synapses by type. Each of these, if adopted, will require research into the best way to combine the new data into the reconstruction process.
