A deep understanding of how neural circuits process information will require measurement of neural activity on many different scales, ranging from single synapses to large assemblies of neurons, in the intact brain. To advance this goal we engineer genetically encoded tools for measuring and controlling neuronal activity in the intact brain.
We accelerate the protein engineering and testing cycle for tools for light-based neurophysiology.
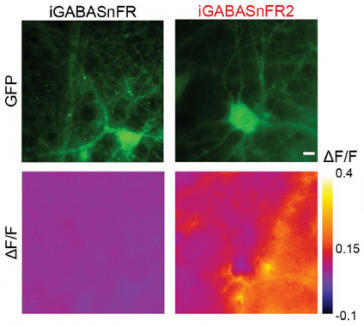
Shortly after it was first cloned, green fluorescent protein was engineered to report changes in cellular calcium and membrane voltage. It took an additional fifteen years of molecular engineering by several research groups on three continents to make protein sensors of calcium useful for brain research. The Genetically-ENcoded Indicator and Effector (GENIE) project has the organization, multidisciplinary know-how, and technology to greatly accelerate optimization of protein sensors for neural function.
We are a diverse group of scientists developing enabling technology for investigating neuronal circuits.
We are invertebrate and vertebrate neurophysiologists, behavioral neuroscientists, biophysicists, microscopists, structural biologists, protein biochemists, cell biologists, genetic engineers, virologists, and molecular biologists. We leverage the shared resources at Janelia, which include experts in instrument design, cell culture, histology, genetic engineering, applied physics, and microscopy. We also partner with collaborators worldwide and participants in the Janelia visitor program.
Our goal is to develop best-in-class, well-characterized reagents and make them widely available to the research community.