Representative hepatocytes from five distinct categories identified by organelle architecture. Credit: Daniel Feliciano / HHMI Janelia Research Campus
As we go about our day, the trillions of cells in our bodies run like well-oiled machines: continually sensing what’s happening around them and making modifications to keep us humming along.
Like adjusting a gear in a car based on the terrain, the cell modifies compartmentalized structures called organelles to change its metabolic state in response to its environment.
Now, researchers at HHMI’s Janelia Research Campus have devised a new, low-cost way to look under the hood and examine these organelles and how they change in every cell within a tissue.
These insights could help scientists better understand how diseases develop and, potentially, how to treat them.
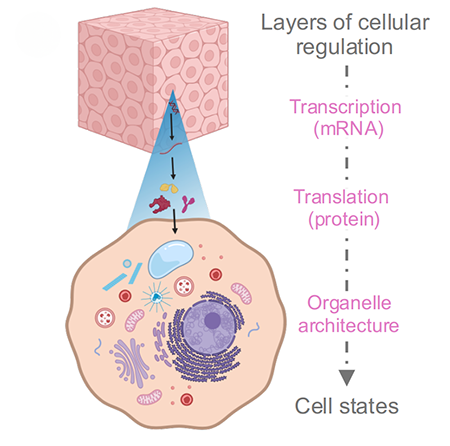
“The organelles carry information about how the cell ‘feels’ in response to cues from its environment, so by evaluating how organelles are changing, we could extract clues about the state of the cell,” says Janelia Fellow Daniel Feliciano, who heads a lab that led the new research. “By connecting the subcellular scale to the tissue scale in a complex organism, you can see how structural changes inside cells shape the behavior of entire tissues.”
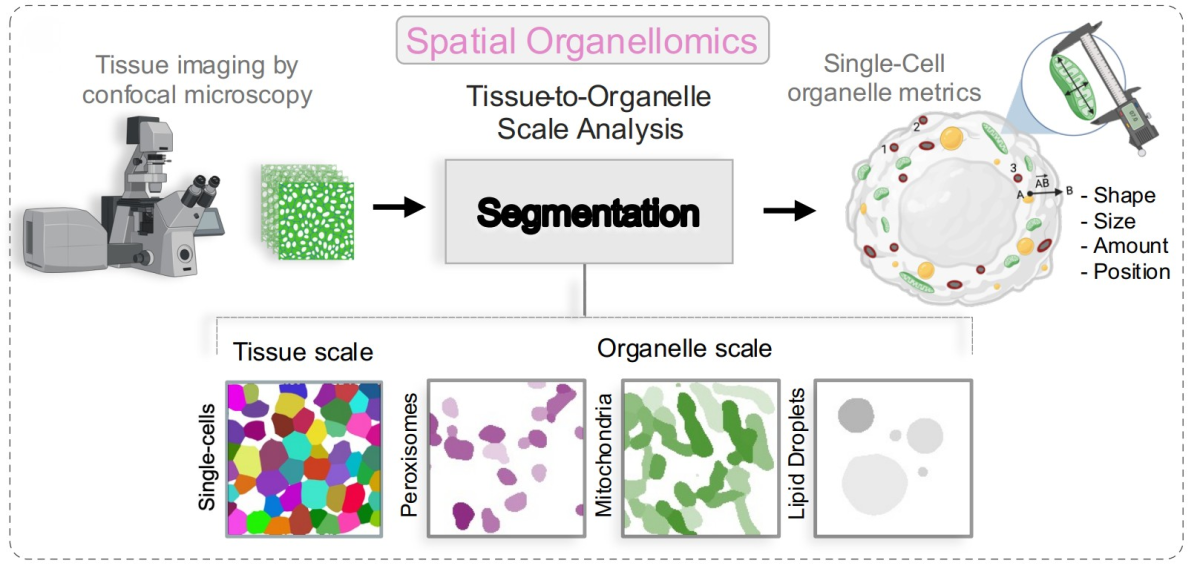
Understanding cells within tissues – and what organelles reveal
Feliciano and the team developed Spatial Organellomics or sOrganellomics, a technique to extract information from key organelles to assess the state of cells within a tissue.
First, they image tissue samples using a 3D fluorescence microscope. Then, they identify individual organelles within each cell using “Liv-Zones”, a segmentation and analysis pipeline develop in collaboration with Jan Funke, a former Janelia Group Leader. This pipeline integrates Cellpose, software developed by Janelia Group Leaders Carsen Stringer and Marius Pachitariu, to identify cells and organelles.
Next, they extract dozens of features for each organelle of interest, including number, position, shape, and size. By compiling these features, they create an “organelle signature” for each cell. Mapping these signatures across the entire tissue allows researchers to visualize different cellular states and track how they change over time.
Unlike other methods that examine all the RNA molecules or proteins in cells, sOrganellomics does not require specialized equipment. It also doesn’t destroy the sample so it can be combined with these other techniques to extract additional layers of information.
Comparative 3D visualization of hepatocyte organelle architecture in control, fasted, and Western diet–fed mice. Credit: Daniel Feliciano / HHMI Janelia Research Campus
Examining organelles in liver cells
The team used the new method to look at cells called hepatocytes, which perform different metabolic tasks in the liver.
They classified hepatocytes into five categories based on their organelle signatures. Surprisingly, when they mapped the cells’ locations, they found that different types of hepatocytes were intermixed throughout the liver, rather than being confined to specific zones, as previously thought.
The team hypothesized that this intermingling of hepatocyte subtypes might help the liver better adapt to environmental changes.
“One possibility is that this sub-zonal heterogeneity increases the plasticity of the liver allowing it to better adapt to things like toxins in the blood, pathogens, or even changes in diet.” Feliciano says.
Seeing how cells and tissues change
To test their hypothesis, the team exposed animals to different diets: one with a reduced level of nutrients and one high in fat and carbohydrates. For both experimental conditions, the five previously observed hepatocyte categories disappeared, and new types representing different hepatocyte states emerged. Each diet generated unique hepatocyte categories that were less diverse than before.
“What that means is that the hepatocytes are adapting -- they’re changing and configuring everything inside so they can actually handle that perturbation, that stress, and they handle it in different ways,” Feliciano says. “They have different strategies depending on the nutritional challenge.”
Because the cells become more homogeneous, the findings suggest that an animal’s diet may reduce the liver’s ability to respond to future stresses, potentially shedding light on how disease vulnerability might develop.
“The tissue becomes less resistant to anything that comes after that,” Feliciano says. “If something comes, like an infection or some of the cells become cancerous, there may be fewer ways for the tissue to counter that challenge.”
Predicting disease progression from organelle signatures
In addition to these biological insights, the team implemented AI models that use organelle signatures to predict the different hepatocyte states as well as the animal’s diet with more than 95 percent accuracy. This strategy can also track the progression of fatty liver disease in mice, suggesting that organelle signatures could eventually serve as diagnostic indicators.
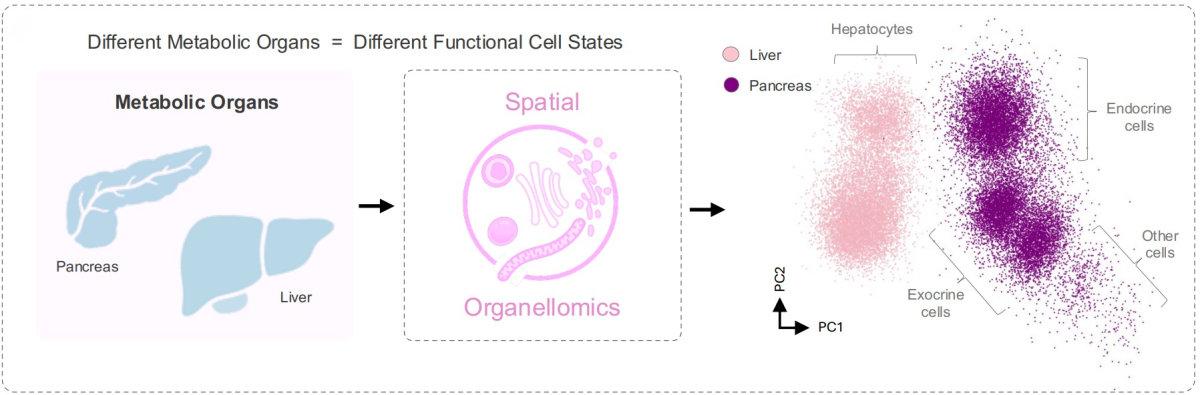
Beyond the liver, sOrganellomics could be used to examine cells in different organs and diseases. The team is currently collaborating with other Janelia labs to apply the method in the gut and the pancreas.
Key Takeaways
- A team led by Janelia Fellow Daniel Feliciano developed a new, cost-effective technique that uses information about the structures inside cells to reveal the condition of tissues and organs.
- Using their new method, the team assessed the cellular makeup of liver tissue and how it changes under different diets. They also implemented AI tools that use these subcellular features to track changes in cell states and disease progression.
- Understanding how cells and tissues respond to their environment could help scientists gain insight into how diseases develop and, potentially, how to treat them.