We pioneer 3D isotropic high resolution volume Electron Microscopy (vEM) technology to enable discoveries in connectomics, tissue biology, and cell biology.
We aim to transcend the limitations of Focused Ion Beam Scanning Electron Microscopy (FIB-SEM) technology to enable large volume [1.0], high throughput [2.0], and artifact-free [3.0], isotropic high-resolution 3D imaging.
Enhanced FIB-SEM 1.0: Enable large volume high resolution 3D imaging, available for Open Science
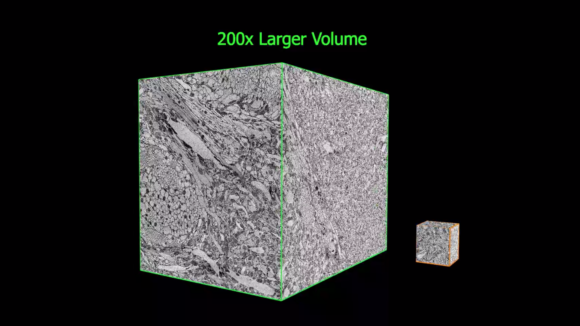
Isotropic high-resolution imaging of large volumes provides unprecedented opportunities to advance connectomics and cell biology research. Conventional Focused Ion Beam Scanning Electron Microscopy (FIB-SEM) offers unique benefits such as high resolution (< 10 nm in x, y, and z), minimal defects, and robust image alignment, well suited for superior tracing of neuronal processes and automated segmentation. However, its prevailing deficiencies in imaging speed and duration cap the maximum possible image volume. We developed innovative solutions to overcome these barriers and transformed FIB-SEM from a conventional lab tool lacking long term reliability to a robust imaging platform capable of years of continuous imaging without defects in the final image stack [Xu et al. 2017; Xu et al. 2020a; Xu et al. 2020b].
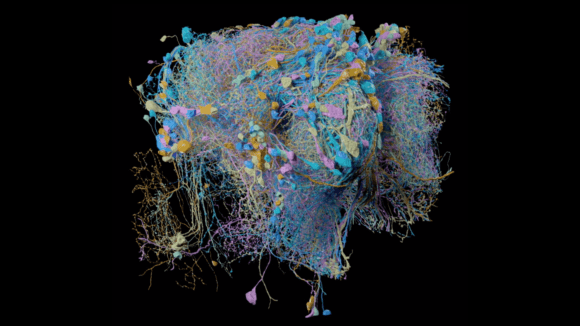
These improvements have enabled the extension of the continuous imaged volume by more than four orders of magnitude from 103 µm3 to at least 107 µm3, while maintaining an isotropic sampling of 8 x 8 x 8 nm3 voxels. Moreover, by trading off imaging speed, the system can readily be operated at even higher resolutions achieving voxel sizes of 4 x 4 x 4 nm3. The expanded volumes enabled by this enhanced FIB-SEM technology 1.0 enable a vast new regime in scientific learning, where nano scale resolution coupled with meso and even macro scale volumes is critical. The largest connectome to date has been generated through this enhanced FIB-SEM platform [Xu et al. 2020c], where the superior z resolution empowers automated tracing of neuronal processes and reduces the time-consuming human proofreading effort. The journey of enhanced FIB-SEM development and Drosophia hemibrain endeavor was featured in Nature & New York Times.
Drosophila hemibrain generated by two Enhanced FIB-SEM systems through 13 hot-knife tabs.
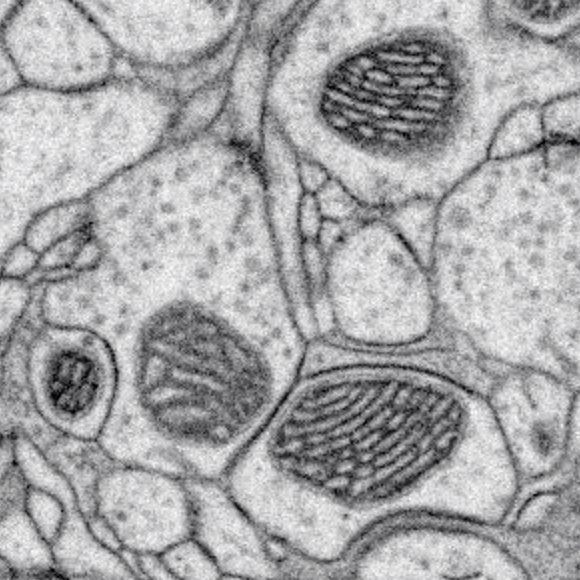
Higher resolution further improves the interpretation of otherwise ambiguous details. Nearly all organelles can be resolved and classified with whole cell imaging at 4 x 4 x 4 nm3 nm voxels. We use this approach to establish an isotropic 3D reference library of whole cells and tissues [Xu et al. 2021], and open access to all datasets in OpenOrganelle [Heinrich et al. 2021]. Together, this represents the beginning of a new field of high-resolution whole-cell 3D EM, and the story was featured in Nature and TheScientist. Additionally, by combining with super-resolution fluorescence imaging, CLEM applications unleash the full potential of intracellular organelle identification with labeling insights.
"I will ask you to mark again that rather typical feature of the development of our subject; how so much progress depends on the interplay of techniques, discoveries and new ideas, probably in that order of decreasing importance."
Sydney Brenner
(1927-2019)