Filter
Associated Lab
- Aguilera Castrejon Lab (2) Apply Aguilera Castrejon Lab filter
- Ahrens Lab (55) Apply Ahrens Lab filter
- Aso Lab (40) Apply Aso Lab filter
- Baker Lab (19) Apply Baker Lab filter
- Betzig Lab (101) Apply Betzig Lab filter
- Beyene Lab (9) Apply Beyene Lab filter
- Bock Lab (14) Apply Bock Lab filter
- Branson Lab (51) Apply Branson Lab filter
- Card Lab (37) Apply Card Lab filter
- Cardona Lab (45) Apply Cardona Lab filter
- Chklovskii Lab (10) Apply Chklovskii Lab filter
- Clapham Lab (14) Apply Clapham Lab filter
- Cui Lab (19) Apply Cui Lab filter
- Darshan Lab (8) Apply Darshan Lab filter
- Dickson Lab (32) Apply Dickson Lab filter
- Druckmann Lab (21) Apply Druckmann Lab filter
- Dudman Lab (38) Apply Dudman Lab filter
- Eddy/Rivas Lab (30) Apply Eddy/Rivas Lab filter
- Egnor Lab (4) Apply Egnor Lab filter
- Espinosa Medina Lab (15) Apply Espinosa Medina Lab filter
- Feliciano Lab (8) Apply Feliciano Lab filter
- Fetter Lab (31) Apply Fetter Lab filter
- FIB-SEM Technology (1) Apply FIB-SEM Technology filter
- Fitzgerald Lab (16) Apply Fitzgerald Lab filter
- Freeman Lab (15) Apply Freeman Lab filter
- Funke Lab (39) Apply Funke Lab filter
- Gonen Lab (59) Apply Gonen Lab filter
- Grigorieff Lab (34) Apply Grigorieff Lab filter
- Harris Lab (53) Apply Harris Lab filter
- Heberlein Lab (13) Apply Heberlein Lab filter
- Hermundstad Lab (24) Apply Hermundstad Lab filter
- Hess Lab (74) Apply Hess Lab filter
- Ilanges Lab (2) Apply Ilanges Lab filter
- Jayaraman Lab (42) Apply Jayaraman Lab filter
- Ji Lab (33) Apply Ji Lab filter
- Johnson Lab (1) Apply Johnson Lab filter
- Karpova Lab (13) Apply Karpova Lab filter
- Keleman Lab (8) Apply Keleman Lab filter
- Keller Lab (61) Apply Keller Lab filter
- Koay Lab (2) Apply Koay Lab filter
- Lavis Lab (139) Apply Lavis Lab filter
- Lee (Albert) Lab (29) Apply Lee (Albert) Lab filter
- Leonardo Lab (19) Apply Leonardo Lab filter
- Li Lab (4) Apply Li Lab filter
- Lippincott-Schwartz Lab (100) Apply Lippincott-Schwartz Lab filter
- Liu (Yin) Lab (2) Apply Liu (Yin) Lab filter
- Liu (Zhe) Lab (59) Apply Liu (Zhe) Lab filter
- Looger Lab (137) Apply Looger Lab filter
- Magee Lab (31) Apply Magee Lab filter
- Menon Lab (12) Apply Menon Lab filter
- Murphy Lab (6) Apply Murphy Lab filter
- O'Shea Lab (6) Apply O'Shea Lab filter
- Otopalik Lab (1) Apply Otopalik Lab filter
- Pachitariu Lab (36) Apply Pachitariu Lab filter
- Pastalkova Lab (5) Apply Pastalkova Lab filter
- Pavlopoulos Lab (7) Apply Pavlopoulos Lab filter
- Pedram Lab (4) Apply Pedram Lab filter
- Podgorski Lab (16) Apply Podgorski Lab filter
- Reiser Lab (45) Apply Reiser Lab filter
- Riddiford Lab (20) Apply Riddiford Lab filter
- Romani Lab (31) Apply Romani Lab filter
- Rubin Lab (107) Apply Rubin Lab filter
- Saalfeld Lab (46) Apply Saalfeld Lab filter
- Satou Lab (1) Apply Satou Lab filter
- Scheffer Lab (36) Apply Scheffer Lab filter
- Schreiter Lab (51) Apply Schreiter Lab filter
- Sgro Lab (1) Apply Sgro Lab filter
- Shroff Lab (31) Apply Shroff Lab filter
- Simpson Lab (18) Apply Simpson Lab filter
- Singer Lab (37) Apply Singer Lab filter
- Spruston Lab (58) Apply Spruston Lab filter
- Stern Lab (73) Apply Stern Lab filter
- Sternson Lab (47) Apply Sternson Lab filter
- Stringer Lab (33) Apply Stringer Lab filter
- Svoboda Lab (131) Apply Svoboda Lab filter
- Tebo Lab (9) Apply Tebo Lab filter
- Tervo Lab (9) Apply Tervo Lab filter
- Tillberg Lab (18) Apply Tillberg Lab filter
- Tjian Lab (17) Apply Tjian Lab filter
- Truman Lab (58) Apply Truman Lab filter
- Turaga Lab (40) Apply Turaga Lab filter
- Turner Lab (28) Apply Turner Lab filter
- Vale Lab (8) Apply Vale Lab filter
- Voigts Lab (3) Apply Voigts Lab filter
- Wang (Meng) Lab (22) Apply Wang (Meng) Lab filter
- Wang (Shaohe) Lab (6) Apply Wang (Shaohe) Lab filter
- Wu Lab (8) Apply Wu Lab filter
- Zlatic Lab (26) Apply Zlatic Lab filter
- Zuker Lab (5) Apply Zuker Lab filter
Associated Project Team
- CellMap (12) Apply CellMap filter
- COSEM (3) Apply COSEM filter
- FIB-SEM Technology (3) Apply FIB-SEM Technology filter
- Fly Descending Interneuron (11) Apply Fly Descending Interneuron filter
- Fly Functional Connectome (14) Apply Fly Functional Connectome filter
- Fly Olympiad (5) Apply Fly Olympiad filter
- FlyEM (54) Apply FlyEM filter
- FlyLight (49) Apply FlyLight filter
- GENIE (47) Apply GENIE filter
- Integrative Imaging (6) Apply Integrative Imaging filter
- Larval Olympiad (2) Apply Larval Olympiad filter
- MouseLight (18) Apply MouseLight filter
- NeuroSeq (1) Apply NeuroSeq filter
- ThalamoSeq (1) Apply ThalamoSeq filter
- Tool Translation Team (T3) (27) Apply Tool Translation Team (T3) filter
- Transcription Imaging (45) Apply Transcription Imaging filter
Associated Support Team
- Project Pipeline Support (5) Apply Project Pipeline Support filter
- Anatomy and Histology (18) Apply Anatomy and Histology filter
- Cryo-Electron Microscopy (39) Apply Cryo-Electron Microscopy filter
- Electron Microscopy (17) Apply Electron Microscopy filter
- Gene Targeting and Transgenics (11) Apply Gene Targeting and Transgenics filter
- High Performance Computing (7) Apply High Performance Computing filter
- Integrative Imaging (17) Apply Integrative Imaging filter
- Invertebrate Shared Resource (40) Apply Invertebrate Shared Resource filter
- Janelia Experimental Technology (37) Apply Janelia Experimental Technology filter
- Management Team (1) Apply Management Team filter
- Molecular Genomics (15) Apply Molecular Genomics filter
- Primary & iPS Cell Culture (14) Apply Primary & iPS Cell Culture filter
- Project Technical Resources (50) Apply Project Technical Resources filter
- Quantitative Genomics (19) Apply Quantitative Genomics filter
- Scientific Computing (93) Apply Scientific Computing filter
- Viral Tools (14) Apply Viral Tools filter
- Vivarium (7) Apply Vivarium filter
Publication Date
- 2025 (160) Apply 2025 filter
- 2024 (213) Apply 2024 filter
- 2023 (158) Apply 2023 filter
- 2022 (166) Apply 2022 filter
- 2021 (175) Apply 2021 filter
- 2020 (177) Apply 2020 filter
- 2019 (177) Apply 2019 filter
- 2018 (206) Apply 2018 filter
- 2017 (186) Apply 2017 filter
- 2016 (191) Apply 2016 filter
- 2015 (195) Apply 2015 filter
- 2014 (190) Apply 2014 filter
- 2013 (136) Apply 2013 filter
- 2012 (112) Apply 2012 filter
- 2011 (98) Apply 2011 filter
- 2010 (61) Apply 2010 filter
- 2009 (56) Apply 2009 filter
- 2008 (40) Apply 2008 filter
- 2007 (21) Apply 2007 filter
- 2006 (3) Apply 2006 filter
2721 Janelia Publications
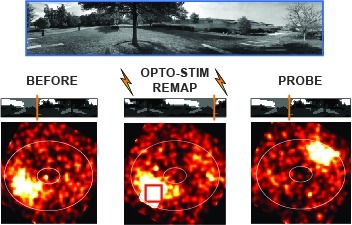
Showing 1111-1120 of 2721 resultsMany animals rely on an internal heading representation when navigating in varied environments. How this representation is linked to the sensory cues that define different surroundings is unclear. In the fly brain, heading is represented by 'compass' neurons that innervate a ring-shaped structure known as the ellipsoid body. Each compass neuron receives inputs from 'ring' neurons that are selective for particular visual features; this combination provides an ideal substrate for the extraction of directional information from a visual scene. Here we combine two-photon calcium imaging and optogenetics in tethered flying flies with circuit modelling, and show how the correlated activity of compass and visual neurons drives plasticity, which flexibly transforms two-dimensional visual cues into a stable heading representation. We also describe how this plasticity enables the fly to convert a partial heading representation, established from orienting within part of a novel setting, into a complete heading representation. Our results provide mechanistic insight into the memory-related computations that are essential for flexible navigation in varied surroundings.
Interindividual differences in neuronal wiring may contribute to behavioral individuality and affect susceptibility to neurological disorders. To investigate the causes and potential consequences of wiring variation in Drosophila melanogaster, we focused on a hemilineage of ventral nerve cord interneurons that exhibits morphological variability. We find that late-born subclasses of the 12A hemilineage are highly sensitive to genetic and environmental variation. Neurons in the second thoracic segment are particularly variable with regard to two developmental decisions, whereas its segmental homologs are more robust. This variability "hotspot" depends on Ultrabithorax expression in the 12A neurons, indicating variability is cell-intrinsic and under genetic control. 12A development is more variable and sensitive to temperature in long-established laboratory strains than in strains recently derived from the wild. Strains with a high frequency of one of the 12A variants also showed a high frequency of animals with delayed spontaneous flight initiation, whereas other wing-related behaviors did not show such a correlation and were thus not overtly affected by 12A variation. These results show that neurodevelopmental robustness is variable and under genetic control in Drosophila and suggest that the fly may serve as a model for identifying conserved gene pathways that stabilize wiring in stressful developmental environments. Moreover, some neuronal lineages are variation hotspots and thus may be more amenable to evolutionary change.
Genetically hard-wired neural mechanisms must enforce behavioral reproductive isolation because interspecies courtship is rare even in sexually na{\"ıve animals of most species. We find that the chemoreceptor Gr32a inhibits male D. melanogaster from courting diverse fruit fly species. Gr32a recognizes nonvolatile aversive cues present on these reproductively dead-end targets, and activity of Gr32a neurons is necessary and sufficient to inhibit interspecies courtship. Male-specific Fruitless (Fru(M)), a master regulator of courtship, also inhibits interspecies courtship. Gr32a and Fru(M) are not coexpressed, but Fru(M) neurons contact Gr32a neurons, suggesting that these genes influence a shared neural circuit that inhibits interspecies courtship. Gr32a and Fru(M) also suppress within-species intermale courtship, but we show that distinct mechanisms preclude sexual displays toward conspecific males and other species. Although this chemosensory pathway does not inhibit interspecies mating in D. melanogaster females, similar mechanisms appear to inhibit this behavior in many other male drosophilids.
Animals execute one particular behavior among many others in a context-dependent manner, yet the mechanisms underlying such behavioral choice remain poorly understood. Here we studied how two fundamental behaviors, sex and sleep, interact at genetic and neuronal levels in Drosophila. We show that an increased need for sleep inhibits male sexual behavior by decreasing the activity of the male-specific P1 neurons that coexpress the sex determination genes fru (M) and dsx, but does not affect female sexual behavior. Further, we delineate a sex-specific neuronal circuit wherein the P1 neurons encoding increased courtship drive suppressed male sleep by forming mutually excitatory connections with the fru (M) -positive sleep-controlling DN1 neurons. In addition, we find that FRU(M) regulates male courtship and sleep through distinct neural substrates. These studies reveal the genetic and neuronal basis underlying the sex-specific interaction between sleep and sexual behaviors in Drosophila, and provide insights into how competing behaviors are co-regulated.Genes and circuits involved in sleep and sexual arousal have been extensively studied in Drosophila. Here the authors identify the sex determination genes fruitless and doublesex, and a sex-specific P1-DN1 neuronal feedback that governs the interaction between these competing behaviors.
Species of the Drosophila melanogaster species subgroup, including the species D. simulans, D. mauritiana, D. yakuba, and D. santomea, have long served as model systems for studying evolution. Studies in these species have been limited, however, by a paucity of genetic and transgenic reagents. Here we describe a collection of transgenic and genetic strains generated to facilitate genetic studies within and between these species. We have generated many strains of each species containing mapped piggyBac transposons including an enhanced yellow fluorescent protein gene expressed in the eyes and a phiC31 attP site-specific integration site. We have tested a subset of these lines for integration efficiency and reporter gene expression levels. We have also generated a smaller collection of other lines expressing other genetically encoded fluorescent molecules in the eyes and a number of other transgenic reagents that will be useful for functional studies in these species. In addition, we have mapped the insertion locations of 58 transposable elements in D. virilis that will be useful for genetic mapping studies.
Male sexual characters are often among the first traits to diverge between closely related species and identifying the genetic basis of such changes can contribute to our understanding of their evolutionary history. However, little is known about the genetic architecture or the specific genes underlying the evolution of male genitalia. The morphology of the claspers, posterior lobes and anal plates exhibit striking differences between Drosophila mauritiana and Drosophila simulans. Using QTL and introgression-based high-resolution mapping, we identified several small regions on chromosome arms 3L and 3R that contribute to differences in these traits. However, we found that the loci underlying the evolution of clasper differences between these two species are independent from those that contribute to posterior lobe and anal plate divergence. Furthermore, while most of the loci affect each trait in the same direction and act additively, we also found evidence for epistasis between loci for clasper bristle number. In addition, we conducted an RNAi screen in D. melanogaster to investigate if positional and expression candidate genes located on chromosome 3L, are also involved in genital development. We found that six of these genes, including components of Wnt signaling and male-specific lethal 3 (msl3), regulate the development of genital traits consistent with the effects of the introgressed regions where they are located and that thus represent promising candidate genes for the evolution these traits.
Different memory components are forgotten through distinct molecular mechanisms. In , the activation of 2 Rho GTPases (Rac1 and Cdc42), respectively, underlies the forgetting of an early labile memory (anesthesia-sensitive memory, ASM) and a form of consolidated memory (anesthesia-resistant memory, ARM). Here, we dissected the molecular mechanisms that tie Rac1 and Cdc42 to the different types of memory forgetting. We found that 2 WASP family proteins, SCAR/WAVE and WASp, act downstream of Rac1 and Cdc42 separately to regulate ASM and ARM forgetting in mushroom body neurons. Arp2/3 complex, which organizes branched actin polymerization, is a canonical downstream effector of WASP family proteins. However, we found that Arp2/3 complex is required in Cdc42/WASp-mediated ARM forgetting but not in Rac1/SCAR-mediated ASM forgetting. Instead, we identified that Rac1/SCAR may function with formin Diaphanous (Dia), a nucleator that facilitates linear actin polymerization, in ASM forgetting. The present study, complementing the previously identified Rac1/cofilin pathway that regulates actin depolymerization, suggests that Rho GTPases regulate forgetting by recruiting both actin polymerization and depolymerization pathways. Moreover, Rac1 and Cdc42 may regulate different types of memory forgetting by tapping into different actin polymerization mechanisms.
Understanding the principles of information processing in neural circuits requires systematic characterization of the participating cell types and their connections, and the ability to measure and perturb their activity. Genetic approaches promise to bring experimental access to complex neural systems, including genetic stalwarts such as the fly and mouse, but also to nongenetic systems such as primates. Together with anatomical and physiological methods, cell-type-specific expression of protein markers and sensors and transducers will be critical to construct circuit diagrams and to measure the activity of genetically defined neurons. Inactivation and activation of genetically defined cell types will establish causal relationships between activity in specific groups of neurons, circuit function, and animal behavior. Genetic analysis thus promises to reveal the logic of the neural circuits in complex brains that guide behaviors. Here we review progress in the genetic analysis of neural circuits and discuss directions for future research and development.
Tremendous progress has been made since Neuron published our Primer on genetic dissection of neural circuits 10 years ago. Since then, cell-type-specific anatomical, neurophysiological, and perturbation studies have been carried out in a multitude of invertebrate and vertebrate organisms, linking neurons and circuits to behavioral functions. New methods allow systematic classification of cell types and provide genetic access to diverse neuronal types for studies of connectivity and neural coding during behavior. Here we evaluate key advances over the past decade and discuss future directions.
Wild-type D. melanogaster males innately possess the ability to perform a multistep courtship ritual to conspecific females. The potential for this behavior is specified by the male-specific products of the fruitless (fru(M)) gene; males without fru(M) do not court females when held in isolation. We show that such fru(M) null males acquire the potential for courtship when grouped with other flies; they apparently learn to court flies with which they were grouped, irrespective of sex or species and retain this behavior for at least a week. The male-specific product of the doublesex gene (dsx(M)) is necessary and sufficient for the acquisition of the potential for such experience-dependent courtship. These results reveal a process that builds, via dsx(M) and social experience, the potential for a more flexible sexual behavior, which could be evolutionarily conserved as dsx-related genes that function in sexual development are found throughout the animal kingdom.