Filter
Result Type
- Apply filter
- Apply filter
- Apply filter
- Apply filter
- Apply filter
- Apply filter
- Apply filter
- Apply filter
- Apply filter
- Area Landing Page (10) Apply Area Landing Page filter
- Collaborations (2) Apply Collaborations filter
- Conferences (256) Apply Conferences filter
- Janelia Archives (19) Apply Janelia Archives filter
- Janelia Archives Landing (1) Apply Janelia Archives Landing filter
- Lab (58) Apply Lab filter
- News Stories (285) Apply News Stories filter
- Other (533) Apply Other filter
- People (683) Apply People filter
- Project Team (15) Apply Project Team filter
- Publications (2852) Apply Publications filter
- Support Team (21) Apply Support Team filter
- Theory Fellow Landing Page (3) Apply Theory Fellow Landing Page filter
- Tool (132) Apply Tool filter
Associated Lab
- Aguilera Castrejon Lab (9) Apply Aguilera Castrejon Lab filter
- Ahrens Lab (81) Apply Ahrens Lab filter
- Aso Lab (49) Apply Aso Lab filter
- Baker Lab (20) Apply Baker Lab filter
- Betzig Lab (114) Apply Betzig Lab filter
- Beyene Lab (22) Apply Beyene Lab filter
- Bock Lab (15) Apply Bock Lab filter
- Branson Lab (65) Apply Branson Lab filter
- Card Lab (38) Apply Card Lab filter
- Cardona Lab (45) Apply Cardona Lab filter
- Chklovskii Lab (10) Apply Chklovskii Lab filter
- Clapham Lab (17) Apply Clapham Lab filter
- Cui Lab (20) Apply Cui Lab filter
- Darshan Lab (8) Apply Darshan Lab filter
- Dennis Lab (10) Apply Dennis Lab filter
- Dickson Lab (34) Apply Dickson Lab filter
- Druckmann Lab (21) Apply Druckmann Lab filter
- Dudman Lab (55) Apply Dudman Lab filter
- Eddy/Rivas Lab (30) Apply Eddy/Rivas Lab filter
- Egnor Lab (5) Apply Egnor Lab filter
- Espinosa Medina Lab (32) Apply Espinosa Medina Lab filter
- Feliciano Lab (19) Apply Feliciano Lab filter
- Fetter Lab (31) Apply Fetter Lab filter
- FIB-SEM Technology (1) Apply FIB-SEM Technology filter
- Fitzgerald Lab (17) Apply Fitzgerald Lab filter
- Freeman Lab (16) Apply Freeman Lab filter
- Funke Lab (48) Apply Funke Lab filter
- Gonen Lab (60) Apply Gonen Lab filter
- Grigorieff Lab (34) Apply Grigorieff Lab filter
- Harris Lab (66) Apply Harris Lab filter
- Heberlein Lab (15) Apply Heberlein Lab filter
- Hermundstad Lab (33) Apply Hermundstad Lab filter
- Hess Lab (88) Apply Hess Lab filter
- Ilanges Lab (10) Apply Ilanges Lab filter
- Jayaraman Lab (59) Apply Jayaraman Lab filter
- Ji Lab (34) Apply Ji Lab filter
- Johnson Lab (7) Apply Johnson Lab filter
- Kainmueller Lab (1) Apply Kainmueller Lab filter
- Karpova Lab (24) Apply Karpova Lab filter
- Keleman Lab (8) Apply Keleman Lab filter
- Keller Lab (80) Apply Keller Lab filter
- Koay Lab (10) Apply Koay Lab filter
- Lavis Lab (169) Apply Lavis Lab filter
- Lee (Albert) Lab (32) Apply Lee (Albert) Lab filter
- Leonardo Lab (19) Apply Leonardo Lab filter
- Li Lab (16) Apply Li Lab filter
- Lippincott-Schwartz Lab (122) Apply Lippincott-Schwartz Lab filter
- Liu (Yin) Lab (9) Apply Liu (Yin) Lab filter
- Liu (Zhe) Lab (68) Apply Liu (Zhe) Lab filter
- Looger Lab (144) Apply Looger Lab filter
- Magee Lab (31) Apply Magee Lab filter
- Menon Lab (12) Apply Menon Lab filter
- Murphy Lab (7) Apply Murphy Lab filter
- O'Shea Lab (14) Apply O'Shea Lab filter
- Otopalik Lab (7) Apply Otopalik Lab filter
- Pachitariu Lab (51) Apply Pachitariu Lab filter
- Pastalkova Lab (7) Apply Pastalkova Lab filter
- Pavlopoulos Lab (7) Apply Pavlopoulos Lab filter
- Pedram Lab (11) Apply Pedram Lab filter
- Podgorski Lab (18) Apply Podgorski Lab filter
- Reiser Lab (67) Apply Reiser Lab filter
- Riddiford Lab (21) Apply Riddiford Lab filter
- Romani Lab (54) Apply Romani Lab filter
- Rubin Lab (127) Apply Rubin Lab filter
- Ryan Lab (1) Apply Ryan Lab filter
- Saalfeld Lab (57) Apply Saalfeld Lab filter
- Satou Lab (9) Apply Satou Lab filter
- Scheffer Lab (39) Apply Scheffer Lab filter
- Schreiter Lab (67) Apply Schreiter Lab filter
- Schulze Lab (4) Apply Schulze Lab filter
- Sgro Lab (17) Apply Sgro Lab filter
- Shroff Lab (44) Apply Shroff Lab filter
- Simpson Lab (18) Apply Simpson Lab filter
- Singer Lab (39) Apply Singer Lab filter
- Spruston Lab (78) Apply Spruston Lab filter
- Stern Lab (86) Apply Stern Lab filter
- Sternson Lab (52) Apply Sternson Lab filter
- Stringer Lab (48) Apply Stringer Lab filter
- Svoboda Lab (146) Apply Svoboda Lab filter
- Tavakoli Lab (3) Apply Tavakoli Lab filter
- Tebo Lab (22) Apply Tebo Lab filter
- Tervo Lab (16) Apply Tervo Lab filter
- Tillberg Lab (23) Apply Tillberg Lab filter
- Tjian Lab (19) Apply Tjian Lab filter
- Truman Lab (59) Apply Truman Lab filter
- Turaga Lab (62) Apply Turaga Lab filter
- Turner Lab (33) Apply Turner Lab filter
- Vale Lab (10) Apply Vale Lab filter
- Voigts Lab (10) Apply Voigts Lab filter
- Wang (Meng) Lab (46) Apply Wang (Meng) Lab filter
- Wang (Shaohe) Lab (14) Apply Wang (Shaohe) Lab filter
- Wong-Campos Lab (6) Apply Wong-Campos Lab filter
- Wu Lab (9) Apply Wu Lab filter
- Zlatic Lab (26) Apply Zlatic Lab filter
- Zuker Lab (5) Apply Zuker Lab filter
Associated Project Team
- CellMap (30) Apply CellMap filter
- COSEM (3) Apply COSEM filter
- FIB-SEM Technology (11) Apply FIB-SEM Technology filter
- Fly Descending Interneuron (14) Apply Fly Descending Interneuron filter
- Fly Functional Connectome (15) Apply Fly Functional Connectome filter
- Fly Olympiad (5) Apply Fly Olympiad filter
- FlyEM (66) Apply FlyEM filter
- FlyLight (59) Apply FlyLight filter
- FuncEWOrm (9) Apply FuncEWOrm filter
- GENIE (68) Apply GENIE filter
- Integrative Imaging (9) Apply Integrative Imaging filter
- Larval Olympiad (2) Apply Larval Olympiad filter
- MouseLight (26) Apply MouseLight filter
- NeuroSeq (2) Apply NeuroSeq filter
- ThalamoSeq (1) Apply ThalamoSeq filter
- Tool Translation Team (T3) (38) Apply Tool Translation Team (T3) filter
- Transcription Imaging (48) Apply Transcription Imaging filter
Associated Support Team
- Project Pipeline Support (39) Apply Project Pipeline Support filter
- Anatomy and Histology (25) Apply Anatomy and Histology filter
- Cryo-Electron Microscopy (56) Apply Cryo-Electron Microscopy filter
- Electron Microscopy (24) Apply Electron Microscopy filter
- Flow Cytometry (4) Apply Flow Cytometry filter
- Gene Targeting and Transgenics (20) Apply Gene Targeting and Transgenics filter
- High Performance Computing (14) Apply High Performance Computing filter
- Immortalized Cell Line Culture (6) Apply Immortalized Cell Line Culture filter
- Integrative Imaging (39) Apply Integrative Imaging filter
- Invertebrate Shared Resource (50) Apply Invertebrate Shared Resource filter
- Janelia Experimental Technology (105) Apply Janelia Experimental Technology filter
- Management Team (1) Apply Management Team filter
- Mass Spectrometry (5) Apply Mass Spectrometry filter
- Media Facil\ (7) Apply Media Facil\ filter
- Molecular Genomics (20) Apply Molecular Genomics filter
- Project Technical Resources (71) Apply Project Technical Resources filter
- Quantitative Genomics (27) Apply Quantitative Genomics filter
- Scientific Computing (144) Apply Scientific Computing filter
- Stem Cell & Primary Culture (25) Apply Stem Cell & Primary Culture filter
- Viral Tools (22) Apply Viral Tools filter
- Vivarium (10) Apply Vivarium filter
Publication Date
- 2026 (77) Apply 2026 filter
- 2025 (247) Apply 2025 filter
- 2024 (239) Apply 2024 filter
- 2023 (187) Apply 2023 filter
- 2022 (192) Apply 2022 filter
- 2021 (187) Apply 2021 filter
- 2020 (194) Apply 2020 filter
- 2019 (201) Apply 2019 filter
- 2018 (221) Apply 2018 filter
- 2017 (202) Apply 2017 filter
- 2016 (207) Apply 2016 filter
- 2015 (222) Apply 2015 filter
- 2014 (216) Apply 2014 filter
- 2013 (152) Apply 2013 filter
- 2012 (112) Apply 2012 filter
- 2011 (98) Apply 2011 filter
- 2010 (61) Apply 2010 filter
- 2009 (56) Apply 2009 filter
- 2008 (40) Apply 2008 filter
- 2007 (21) Apply 2007 filter
- 2006 (3) Apply 2006 filter
Tool Types
- Data (9) Apply Data filter
- Data Application (7) Apply Data Application filter
- Figshare (1) Apply Figshare filter
- Human Health (2) Apply Human Health filter
- Imaging Instrumentation (11) Apply Imaging Instrumentation filter
- Laboratory Hardware (3) Apply Laboratory Hardware filter
- Laboratory Tool (6) Apply Laboratory Tool filter
- Laboratory Tools (51) Apply Laboratory Tools filter
- Medical Technology (1) Apply Medical Technology filter
- Model Organisms (9) Apply Model Organisms filter
- Reagents (29) Apply Reagents filter
- Software (20) Apply Software filter
5017 Results
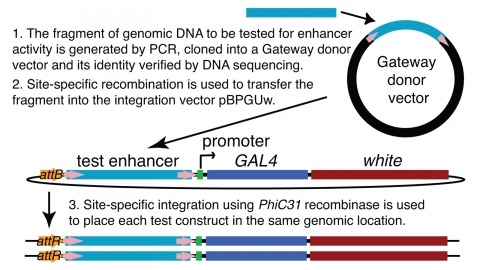
Showing 4681-4690 of 5017 resultsWe demonstrate the feasibility of generating thousands of transgenic Drosophila melanogaster lines in which the expression of an exogenous gene is reproducibly directed to distinct small subsets of cells in the adult brain. We expect the expression patterns produced by the collection of 5,000 lines that we are currently generating to encompass all neurons in the brain in a variety of intersecting patterns. Overlapping 3-kb DNA fragments from the flanking noncoding and intronic regions of genes thought to have patterned expression in the adult brain were inserted into a defined genomic location by site-specific recombination. These fragments were then assayed for their ability to function as transcriptional enhancers in conjunction with a synthetic core promoter designed to work with a wide variety of enhancer types. An analysis of 44 fragments from four genes found that >80% drive expression patterns in the brain; the observed patterns were, on average, comprised of <100 cells. Our results suggest that the D. melanogaster genome contains >50,000 enhancers and that multiple enhancers drive distinct subsets of expression of a gene in each tissue and developmental stage. We expect that these lines will be valuable tools for neuroanatomy as well as for the elucidation of neuronal circuits and information flow in the fly brain.
Sparse manipulation of neuron excitability during free behavior is critical for identifying neural substrates of behavior. Genetic tools for precise neuronal manipulation exist in the fruit fly, Drosophila melanogaster, but behavioral tools are still lacking to identify potentially subtle phenotypes only detectible using high-throughput and high spatiotemporal resolution. We developed three assay components that can be used modularly to study natural and optogenetically induced behaviors. FlyGate automatically releases flies one at a time into an assay. FlyDetect tracks flies in real time, is robust to severe occlusions, and can be used to track appendages, such as the head. GlobeDisplay is a spherical projection system covering the fly's visual receptive field with a single projector. We demonstrate the utility of these components in an integrated system, FlyPEZ, by comprehensively modeling the input-output function for directional looming-evoked escape takeoffs and describing a millisecond-timescale phenotype from genetic silencing of a single visual projection neuron type.
The striatum shows general topographic organization and regional differences in behavioral functions. How corticostriatal topography differs across cortical areas and cell types to support these distinct functions is unclear. This study contrasted corticostriatal projections from two layer 5 cell types, intratelencephalic (IT-type) and pyramidal tract (PT-type) neurons, using viral vectors expressing fluorescent reporters in Cre-driver mice. Corticostriatal projections from sensory and motor cortex are somatotopic, with a decreasing topographic specificity as injection sites move from sensory to motor and frontal areas. Topographic organization differs between IT-type and PT-type neurons, including injections in the same site, with IT-type neurons having higher topographic stereotypy than PT-type neurons. Furthermore, IT-type projections from interconnected cortical areas have stronger correlations in corticostriatal targeting than PT-type projections do. As predicted by a longstanding model, corticostriatal projections of interconnected cortical areas form parallel circuits in the basal ganglia.
Insects and mammals share similarities of neural organization underlying the perception of odors, taste, vision, sound, and gravity. We observed that insect somatosensation also corresponds to that of mammals. In Drosophila, the projections of all the somatosensory neuron types to the insect's equivalent of the spinal cord segregated into modality-specific layers comparable to those in mammals. Some sensory neurons innervate the ventral brain directly to form modality-specific and topological somatosensory maps. Ascending interneurons with dendrites in matching layers of the nerve cord send axons that converge to respective brain regions. Pathways arising from leg somatosensory neurons encode distinct qualities of leg movement information and play different roles in ground detection. Establishment of the ground pattern and genetic tools for neuronal manipulation should provide the basis for elucidating the mechanisms underlying somatosensation.
The medial entorhinal cortex is part of a neural system for mapping the position of an individual within a physical environment. Grid cells, a key component of this system, fire in a characteristic hexagonal pattern of locations, and are organized in modules that collectively form a population code for the animal's allocentric position. The invariance of the correlation structure of this population code across environments and behavioural states, independent of specific sensory inputs, has pointed to intrinsic, recurrently connected continuous attractor networks (CANs) as a possible substrate of the grid pattern. However, whether grid cell networks show continuous attractor dynamics, and how they interface with inputs from the environment, has remained unclear owing to the small samples of cells obtained so far. Here, using simultaneous recordings from many hundreds of grid cells and subsequent topological data analysis, we show that the joint activity of grid cells from an individual module resides on a toroidal manifold, as expected in a two-dimensional CAN. Positions on the torus correspond to positions of the moving animal in the environment. Individual cells are preferentially active at singular positions on the torus. Their positions are maintained between environments and from wakefulness to sleep, as predicted by CAN models for grid cells but not by alternative feedforward models. This demonstration of network dynamics on a toroidal manifold provides a population-level visualization of CAN dynamics in grid cells.