![]() Serial section transmission electron microscopy (ssTEM) produces terabytes of image data (millions of files) that have to be registered and processed in a post-acquisition step to reconstruct the sample volume as a whole. Such reconstruction requires precise management of individual image transformation types, transformation parameters and image metadata in a dedicated database, at the core of the processing workflow. In conjunction with Stephan Saalfeld's group, we provide and maintain a set of web services built around this database to facilitate critical workflow components such as image viewing, registration, filtering and persistence to disk. The web services faciliate management of large scale transformation data sets and provide a RESTful HTTP interface for generating transformed images. They support client integration with the web services and include utilities to prepare image data for downstream processing.
Serial section transmission electron microscopy (ssTEM) produces terabytes of image data (millions of files) that have to be registered and processed in a post-acquisition step to reconstruct the sample volume as a whole. Such reconstruction requires precise management of individual image transformation types, transformation parameters and image metadata in a dedicated database, at the core of the processing workflow. In conjunction with Stephan Saalfeld's group, we provide and maintain a set of web services built around this database to facilitate critical workflow components such as image viewing, registration, filtering and persistence to disk. The web services faciliate management of large scale transformation data sets and provide a RESTful HTTP interface for generating transformed images. They support client integration with the web services and include utilities to prepare image data for downstream processing.
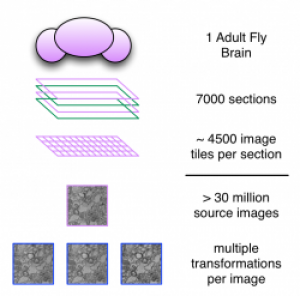
Currently, the services and tools are being used in the assembly of an adult fly brain volume comprised of over 30 million transmission electron microscopy images.
During assembly, the services facilitate iterative reconstruction refinement by maintaining separate transformation data sets that can be compared with and related to each other. One key benefit of maintaining separate data sets is that traces made against one data set can be mapped into another data set. This allows researchers to trace on less refined data while the reconstruction process is still being improved. Later when new refinements are introduced, traces are simply "mapped forward".
The core web services are Java components built around Stephan Saalfeld's transformation and rendering libraries (https://github.com/axtimwalde/mpicbg). The services are deployed on Jetty web servers and utilize mongoDB to persist tile transformation data. Java and Matlab client libraries have been developed to access the services.
The source code can be found on GitHub: https://github.com/saalfeldlab/render.