Filter
Associated Lab
- Ahrens Lab (1) Apply Ahrens Lab filter
- Aso Lab (1) Apply Aso Lab filter
- Baker Lab (3) Apply Baker Lab filter
- Betzig Lab (7) Apply Betzig Lab filter
- Bock Lab (2) Apply Bock Lab filter
- Branson Lab (1) Apply Branson Lab filter
- Cardona Lab (1) Apply Cardona Lab filter
- Cui Lab (3) Apply Cui Lab filter
- Dickson Lab (2) Apply Dickson Lab filter
- Druckmann Lab (1) Apply Druckmann Lab filter
- Dudman Lab (1) Apply Dudman Lab filter
- Eddy/Rivas Lab (5) Apply Eddy/Rivas Lab filter
- Fetter Lab (4) Apply Fetter Lab filter
- Fitzgerald Lab (1) Apply Fitzgerald Lab filter
- Gonen Lab (5) Apply Gonen Lab filter
- Grigorieff Lab (6) Apply Grigorieff Lab filter
- Heberlein Lab (12) Apply Heberlein Lab filter
- Hermundstad Lab (1) Apply Hermundstad Lab filter
- Hess Lab (2) Apply Hess Lab filter
- Jayaraman Lab (2) Apply Jayaraman Lab filter
- Ji Lab (2) Apply Ji Lab filter
- Kainmueller Lab (1) Apply Kainmueller Lab filter
- Keller Lab (4) Apply Keller Lab filter
- Lavis Lab (5) Apply Lavis Lab filter
- Lee (Albert) Lab (1) Apply Lee (Albert) Lab filter
- Leonardo Lab (2) Apply Leonardo Lab filter
- Lippincott-Schwartz Lab (18) Apply Lippincott-Schwartz Lab filter
- Liu (Zhe) Lab (1) Apply Liu (Zhe) Lab filter
- Looger Lab (7) Apply Looger Lab filter
- Magee Lab (1) Apply Magee Lab filter
- Menon Lab (3) Apply Menon Lab filter
- Murphy Lab (1) Apply Murphy Lab filter
- Pastalkova Lab (1) Apply Pastalkova Lab filter
- Pavlopoulos Lab (2) Apply Pavlopoulos Lab filter
- Reiser Lab (2) Apply Reiser Lab filter
- Riddiford Lab (1) Apply Riddiford Lab filter
- Romani Lab (1) Apply Romani Lab filter
- Rubin Lab (4) Apply Rubin Lab filter
- Satou Lab (3) Apply Satou Lab filter
- Scheffer Lab (2) Apply Scheffer Lab filter
- Schreiter Lab (3) Apply Schreiter Lab filter
- Sgro Lab (2) Apply Sgro Lab filter
- Simpson Lab (3) Apply Simpson Lab filter
- Singer Lab (10) Apply Singer Lab filter
- Spruston Lab (1) Apply Spruston Lab filter
- Stern Lab (4) Apply Stern Lab filter
- Sternson Lab (6) Apply Sternson Lab filter
- Svoboda Lab (7) Apply Svoboda Lab filter
- Tjian Lab (4) Apply Tjian Lab filter
- Truman Lab (1) Apply Truman Lab filter
- Turaga Lab (1) Apply Turaga Lab filter
- Turner Lab (2) Apply Turner Lab filter
- Zlatic Lab (1) Apply Zlatic Lab filter
- Zuker Lab (3) Apply Zuker Lab filter
Associated Project Team
Publication Date
- December 2011 (22) Apply December 2011 filter
- November 2011 (15) Apply November 2011 filter
- October 2011 (14) Apply October 2011 filter
- September 2011 (17) Apply September 2011 filter
- August 2011 (14) Apply August 2011 filter
- July 2011 (10) Apply July 2011 filter
- June 2011 (17) Apply June 2011 filter
- May 2011 (13) Apply May 2011 filter
- April 2011 (11) Apply April 2011 filter
- March 2011 (14) Apply March 2011 filter
- February 2011 (16) Apply February 2011 filter
- January 2011 (27) Apply January 2011 filter
- Remove 2011 filter 2011
Type of Publication
190 Publications
Showing 91-100 of 190 resultsHMMER is a software suite for protein sequence similarity searches using probabilistic methods. Previously, HMMER has mainly been available only as a computationally intensive UNIX command-line tool, restricting its use. Recent advances in the software, HMMER3, have resulted in a 100-fold speed gain relative to previous versions. It is now feasible to make efficient profile hidden Markov model (profile HMM) searches via the web. A HMMER web server (http://hmmer.janelia.org) has been designed and implemented such that most protein database searches return within a few seconds. Methods are available for searching either a single protein sequence, multiple protein sequence alignment or profile HMM against a target sequence database, and for searching a protein sequence against Pfam. The web server is designed to cater to a range of different user expertise and accepts batch uploading of multiple queries at once. All search methods are also available as RESTful web services, thereby allowing them to be readily integrated as remotely executed tasks in locally scripted workflows. We have focused on minimizing search times and the ability to rapidly display tabular results, regardless of the number of matches found, developing graphical summaries of the search results to provide quick, intuitive appraisement of them.
The relationship between alcohol consumption, sensitivity, and tolerance is an important question that has been addressed in humans and rodent models. Studies have shown that alcohol consumption and risk of abuse may correlate with (1) increased sensitivity to the stimulant effects of alcohol, (2) decreased sensitivity to the depressant effects of alcohol, and (3) increased alcohol tolerance. However, many conflicting results have been observed. To complement these studies, we utilized a different organism and approach to analyze the relationship between ethanol consumption and other ethanol responses. Using a set of 20 Drosophila melanogaster mutants that were isolated for altered ethanol sensitivity, we measured ethanol-induced hyperactivity, ethanol sedation, sedation tolerance, and ethanol consumption preference. Ethanol preference showed a strong positive correlation with ethanol tolerance, consistent with some rodent and human studies, but not with ethanol hyperactivity or sedation. No pairwise correlations were observed between ethanol hyperactivity, sedation, and tolerance. The evolutionary conservation of the relationship between tolerance and ethanol consumption in flies, rodents, and humans indicates that there are fundamental biological mechanisms linking specific ethanol responses.
The performance of information processing systems, from artificial neural networks to natural neuronal ensembles, depends heavily on the underlying system architecture. In this study, we compare the performance of parallel and layered network architectures during sequential tasks that require both acquisition and retention of information, thereby identifying tradeoffs between learning and memory processes. During the task of supervised, sequential function approximation, networks produce and adapt representations of external information. Performance is evaluated by statistically analyzing the error in these representations while varying the initial network state, the structure of the external information, and the time given to learn the information. We link performance to complexity in network architecture by characterizing local error landscape curvature. We find that variations in error landscape structure give rise to tradeoffs in performance; these include the ability of the network to maximize accuracy versus minimize inaccuracy and produce specific versus generalizable representations of information. Parallel networks generate smooth error landscapes with deep, narrow minima, enabling them to find highly specific representations given sufficient time. While accurate, however, these representations are difficult to generalize. In contrast, layered networks generate rough error landscapes with a variety of local minima, allowing them to quickly find coarse representations. Although less accurate, these representations are easily adaptable. The presence of measurable performance tradeoffs in both layered and parallel networks has implications for understanding the behavior of a wide variety of natural and artificial learning systems.
Morphology evolves often through changes in developmental genes, but the causal mutations, and their effects, remain largely unknown. The evolution of naked cuticle on larvae of Drosophila sechellia resulted from changes in five transcriptional enhancers of shavenbaby (svb), a transcript of the ovo locus that encodes a transcription factor that governs morphogenesis of microtrichiae, hereafter called ’trichomes’. Here we show that the function of one of these enhancers evolved through multiple single-nucleotide substitutions that altered both the timing and level of svb expression. The consequences of these nucleotide substitutions on larval morphology were quantified with a novel functional assay. We found that each substitution had a relatively small phenotypic effect, and that many nucleotide changes account for this large morphological difference. In addition, we observed that the substitutions had non-additive effects. These data provide unprecedented resolution of the phenotypic effects of substitutions and show how individual nucleotide changes in a transcriptional enhancer have caused morphological evolution.
Mammalian genomes contain numerous regulatory DNA sites with unknown target genes. We used mice with an extra β-globin locus control region (LCR) to investigate how a regulator searches the genome for target genes. We find that the LCR samples a restricted nuclear subvolume, wherein it preferentially contacts genes controlled by shared transcription factors. No contacted gene is detectably upregulated except for endogenous β-globin genes located on another chromosome. This demonstrates genetically that mammalian trans activation is possible, but suggests that it will be rare. Trans activation occurs not pan-cellularly, but in 'jackpot' cells enriched for the interchromosomal interaction. Therefore, cell-specific long-range DNA contacts can cause variegated expression.
The male-specific Fruitless proteins (Fru(M)) act to establish the potential for male courtship behavior in Drosophila melanogaster and are expressed in small groups of neurons throughout the nervous system. We screened 1000 GAL4 lines, using assays for general courtship, male-male interactions, and male fertility to determine the phenotypes resulting from the GAL4 driven inhibition of Fru(M) expression in subsets of these neurons. A battery of secondary assays showed that the phenotypic classes of GAL4 lines could be divided into subgroups based on additional neurobiological and behavioral criteria. For example, in some lines restoration of Fru(M) expression in cholinergic neurons restores fertility or reduces male-male courtship. Persistent chains of males courting each other in some lines results from males courting both sexes indiscriminately whereas in other lines this phenotype result from apparent habituation deficits. Inhibition of ectopic Fru(M) expression in females, in populations of neurons where Fru(M) is necessary for male fertility, can rescue female infertility. To identify the neurons responsible for some of the observed behavioral alterations, we determined the overlap between the identified GAL4 lines and endogenous Fru(M) expression in lines with fertility defects. The GAL4 lines causing fertility defects generally had widespread overlap with Fru(M) expression in many regions of the nervous system suggesting likely redundant Fru(M)-expressing neuronal pathways capable of conferring male fertility. From associations between the screened behaviors, we propose a functional model for courtship initiation.
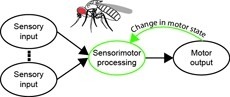
Sensorimotor integration is a field rich in theory backed by a large body of psychophysical evidence. Relating the underlying neural circuitry to these theories has, however, been more challenging. With a wide array of complex behaviors coordinated by their small brains, insects provide powerful model systems to study key features of sensorimotor integration at a mechanistic level. Insect neural circuits perform both hard-wired and learned sensorimotor transformations. They modulate their neural processing based on both internal variables, such as the animal’s behavioral state, and external ones, such as the time of day. Here we present some studies using insect model systems that have produced insights, at the level of individual neurons, about sensorimotor integration and the various ways in which it can be modified by context.
We found that glia secrete myoglianin, a TGF-β ligand, to instruct developmental neural remodeling in Drosophila. Glial myoglianin upregulated neuronal expression of an ecdysone nuclear receptor that triggered neurite remodeling following the late-larval ecdysone peak. Thus glia orchestrate developmental neural remodeling not only by engulfment of unwanted neurites but also by enabling neuron remodeling.
Aversive olfactory memory is formed in the mushroom bodies in Drosophila melanogaster. Memory retrieval requires mushroom body output, but the manner in which a memory trace in the mushroom body drives conditioned avoidance of a learned odor remains unknown. To identify neurons that are involved in olfactory memory retrieval, we performed an anatomical and functional screen of defined sets of mushroom body output neurons. We found that MB-V2 neurons were essential for retrieval of both short- and long-lasting memory, but not for memory formation or memory consolidation. MB-V2 neurons are cholinergic efferent neurons that project from the mushroom body vertical lobes to the middle superiormedial protocerebrum and the lateral horn. Notably, the odor response of MB-V2 neurons was modified after conditioning. As the lateral horn has been implicated in innate responses to repellent odorants, we propose that MB-V2 neurons recruit the olfactory pathway involved in innate odor avoidance during memory retrieval.
Generating microislands of culture substrate on coverslips by spray application of poly-d lysine is a commonly used method for culturing isolated neurons that form self (autaptic) synapses. This preparation has multiple advantages for studying synaptic transmission in isolation; however, generating microislands by spraying produces islands of non-uniform size and thus cultures vary widely in the number of islands containing single neurons. To address these problems, we developed a high-throughput method for reliably generating uniformly shaped microislands of culture substrate. Stamp molds formed of poly(dimethylsiloxane) (PDMS) were fabricated with arrays of circles and used to generate stamps made of 9.2% agarose. The agarose stamps were capable of loading sufficient poly D-lysine and collagen dissolved in acetic acid to rapidly generate coverslips containing at least 64 microislands per coverslip. When hippocampal neurons were cultured on these coverslips, there were significantly more single-neuron islands per coverslip. We noted that single neurons tended to form one of three distinct neurite-arbor morphologies, which varied with island size and the location of the cell body on the island. To our surprise, the number of synapses per autaptic neuron did not correlate with arbor shape or island size, suggesting that other factors regulate the number of synapses formed by isolated neurons. The stamping method we report can be used to increase the number of single-neuron islands per culture and aid in the rapid visualization of microislands.