Filter
Associated Lab
- Ahrens Lab (3) Apply Ahrens Lab filter
- Aso Lab (3) Apply Aso Lab filter
- Baker Lab (5) Apply Baker Lab filter
- Betzig Lab (8) Apply Betzig Lab filter
- Branson Lab (6) Apply Branson Lab filter
- Card Lab (1) Apply Card Lab filter
- Cardona Lab (1) Apply Cardona Lab filter
- Chklovskii Lab (2) Apply Chklovskii Lab filter
- Cui Lab (2) Apply Cui Lab filter
- Darshan Lab (1) Apply Darshan Lab filter
- Dickson Lab (6) Apply Dickson Lab filter
- Druckmann Lab (1) Apply Druckmann Lab filter
- Dudman Lab (4) Apply Dudman Lab filter
- Eddy/Rivas Lab (3) Apply Eddy/Rivas Lab filter
- Egnor Lab (1) Apply Egnor Lab filter
- Fetter Lab (1) Apply Fetter Lab filter
- Fitzgerald Lab (1) Apply Fitzgerald Lab filter
- Freeman Lab (3) Apply Freeman Lab filter
- Gonen Lab (10) Apply Gonen Lab filter
- Grigorieff Lab (3) Apply Grigorieff Lab filter
- Harris Lab (2) Apply Harris Lab filter
- Heberlein Lab (2) Apply Heberlein Lab filter
- Hermundstad Lab (2) Apply Hermundstad Lab filter
- Hess Lab (5) Apply Hess Lab filter
- Jayaraman Lab (1) Apply Jayaraman Lab filter
- Ji Lab (3) Apply Ji Lab filter
- Johnson Lab (1) Apply Johnson Lab filter
- Kainmueller Lab (2) Apply Kainmueller Lab filter
- Karpova Lab (1) Apply Karpova Lab filter
- Keller Lab (8) Apply Keller Lab filter
- Lavis Lab (7) Apply Lavis Lab filter
- Lee (Albert) Lab (4) Apply Lee (Albert) Lab filter
- Leonardo Lab (4) Apply Leonardo Lab filter
- Li Lab (1) Apply Li Lab filter
- Lippincott-Schwartz Lab (12) Apply Lippincott-Schwartz Lab filter
- Liu (Zhe) Lab (4) Apply Liu (Zhe) Lab filter
- Looger Lab (11) Apply Looger Lab filter
- Magee Lab (1) Apply Magee Lab filter
- Menon Lab (3) Apply Menon Lab filter
- Murphy Lab (1) Apply Murphy Lab filter
- Pavlopoulos Lab (2) Apply Pavlopoulos Lab filter
- Reiser Lab (2) Apply Reiser Lab filter
- Riddiford Lab (4) Apply Riddiford Lab filter
- Romani Lab (2) Apply Romani Lab filter
- Rubin Lab (9) Apply Rubin Lab filter
- Saalfeld Lab (1) Apply Saalfeld Lab filter
- Scheffer Lab (7) Apply Scheffer Lab filter
- Sgro Lab (1) Apply Sgro Lab filter
- Simpson Lab (2) Apply Simpson Lab filter
- Singer Lab (10) Apply Singer Lab filter
- Spruston Lab (1) Apply Spruston Lab filter
- Stern Lab (6) Apply Stern Lab filter
- Sternson Lab (5) Apply Sternson Lab filter
- Stringer Lab (1) Apply Stringer Lab filter
- Svoboda Lab (7) Apply Svoboda Lab filter
- Tebo Lab (2) Apply Tebo Lab filter
- Tervo Lab (1) Apply Tervo Lab filter
- Tillberg Lab (1) Apply Tillberg Lab filter
- Tjian Lab (4) Apply Tjian Lab filter
- Truman Lab (1) Apply Truman Lab filter
- Turaga Lab (1) Apply Turaga Lab filter
- Turner Lab (1) Apply Turner Lab filter
- Wang (Shaohe) Lab (1) Apply Wang (Shaohe) Lab filter
- Wu Lab (2) Apply Wu Lab filter
- Zlatic Lab (1) Apply Zlatic Lab filter
- Zuker Lab (1) Apply Zuker Lab filter
Associated Project Team
Publication Date
- December 2014 (35) Apply December 2014 filter
- November 2014 (14) Apply November 2014 filter
- October 2014 (15) Apply October 2014 filter
- September 2014 (17) Apply September 2014 filter
- August 2014 (14) Apply August 2014 filter
- July 2014 (26) Apply July 2014 filter
- June 2014 (14) Apply June 2014 filter
- May 2014 (14) Apply May 2014 filter
- April 2014 (20) Apply April 2014 filter
- March 2014 (18) Apply March 2014 filter
- February 2014 (15) Apply February 2014 filter
- January 2014 (34) Apply January 2014 filter
- Remove 2014 filter 2014
Type of Publication
236 Publications
Showing 11-20 of 236 resultsMitochondrial fractions isolated from tissue culture cells or tissue such as liver after differential centrifugation can be purified further by density gradient centrifugation. Here we describe the use of sucrose for this purpose because it is commonly used and inexpensive and the resulting mitochondria preparations are useful for many purposes.
BACKGROUND: The popular view on insect sociality is that of a harmonious division of labor among two morphologically distinct and functionally non-overlapping castes. But this is a highly derived state and not a prerequisite for a functional society. Rather, caste-flexibility is a central feature in many eusocial wasps, where adult females have the potential to become queens or workers, depending on the social environment. In non-swarming paper wasps (e.g., Polistes), prospective queens fight one another to assert their dominance, with losers becoming workers if they remain on the nest. This aggression is fueled by juvenile hormone (JH) and ecdysteroids, major factors involved in caste differentiation in most eusocial insects. We tested whether these hormones have conserved aggression-promoting functions in Synoeca surinama, a caste-flexible swarm-founding wasp (Epiponini) where reproductive competition is high and aggressive displays are common. RESULTS: We observed the behavioral interactions of S. surinama females in field nests before and after we had removed the egg-laying queen(s). We measured the ovarian reproductive status, hemolymph JH and ecdysteroid titers, ovarian ecdysteroid content, and analyzed the cuticular hydrocarbon (CHC) composition of females engaged in competitive interactions in both queenright and queenless contexts. These data, in combination with hormone manipulation experiments, revealed that neither JH nor ecdysteroids are necessary for the expression of dominance behaviors in S. surinama. Instead, we show that JH likely functions as a gonadotropin and directly modifies the cuticular hydrocarbon blend of young workers to match that of a reproductive. Hemolymph ecdysteroids, in contrast, are not different between queens and workers despite great differences in ovarian ecdysteroid content. CONCLUSIONS: The endocrine profile of S. surinama shows surprising differences from those of other caste-flexible wasps, although a rise in JH titers in replacement queens is a common theme. Extensive remodeling of hormone functions is also evident in the highly eusocial bees, which has been attributed to the evolution of morphologically defined castes. Our results show that hormones which regulate caste-plasticity can lose these roles even while caste-plasticity is preserved.
Single molecule-based superresolution imaging has become an essential tool in modern cell biology. Because of the limited depth of field of optical imaging systems, one of the major challenges in superresolution imaging resides in capturing the 3D nanoscale morphology of the whole cell. Despite many previous attempts to extend the application of photo-activated localization microscopy (PALM) and stochastic optical reconstruction microscopy (STORM) techniques into three dimensions, effective localization depths do not typically exceed 1.2 µm. Thus, 3D imaging of whole cells (or even large organelles) still demands sequential acquisition at different axial positions and, therefore, suffers from the combined effects of out-of-focus molecule activation (increased background) and bleaching (loss of detections). Here, we present the use of multifocus microscopy for volumetric multicolor superresolution imaging. By simultaneously imaging nine different focal planes, the multifocus microscope instantaneously captures the distribution of single molecules (either fluorescent proteins or synthetic dyes) throughout an ∼4-µm-deep volume, with lateral and axial localization precisions of ∼20 and 50 nm, respectively. The capabilities of multifocus microscopy to rapidly image the 3D organization of intracellular structures are illustrated by superresolution imaging of the mammalian mitochondrial network and yeast microtubules during cell division.
Aphids evolved novel cells, called bacteriocytes, that differentiate specifically to harbour the obligatory mutualistic endosymbiotic bacteria Buchnera aphidicola. The genome of the host aphid Acyrthosiphon pisum contains many orphan genes that display no similarity with genes found in other sequenced organisms, prompting us to hypothesize that some of these orphan genes are related to lineage-specific traits, such as symbiosis. We conducted deep sequencing of bacteriocytes mRNA followed by whole mount in situ hybridizations of over-represented transcripts encoding aphid-specific orphan proteins. We identified a novel class of genes that encode small proteins with signal peptides, which are often cysteine-rich, that are over-represented in bacteriocytes. These genes are first expressed at a developmental time point coincident with the incorporation of symbionts strictly in the cells that contribute to the bacteriocyte and this bacteriocyte-specific expression is maintained throughout the aphid's life. The expression pattern suggests that recently evolved secretion proteins act within bacteriocytes, perhaps to mediate the symbiosis with beneficial bacterial partners, which is reminiscent of the evolution of novel cysteine-rich secreted proteins of leguminous plants that regulate nitrogen-fixing endosymbionts.
Similar morphological, physiological, and behavioral features have evolved independently in different species, a pattern known as convergence. It is known that morphological convergence can occur through changes in orthologous genes. In some cases of convergence, cis-regulatory changes generate parallel modifications in the expression patterns of orthologous genes. Our understanding of how changes in cis-regulatory regions contribute to convergence is hampered, usually, by a limited understanding of the global cis-regulatory structure of the evolving genes. Here we examine the genetic causes of a case of precise phenotypic convergence between Drosophila sechellia and Drosophila ezoana, species that diverged ~40 Mya. Previous studies revealed that changes in multiple transcriptional enhancers of shavenbaby (svb, a transcript of the ovo locus) caused phenotypic evolution in the D. sechellia lineage. It has also been shown that the convergent phenotype of D. ezoana was likely caused by cis-regulatory evolution of svb. Here we show that the large-scale cis-regulatory architecture of svb is conserved between these Drosophila species. Furthermore, we show that the D. ezoana orthologs of the evolved D. sechellia enhancers have also evolved expression patterns that correlate precisely with the changes in the phenotype. Our results suggest that phenotypic convergence resulted from multiple noncoding changes that occurred in parallel in the D. sechellia and D. ezoana lineages.
Visualization of specific molecules and their interactions in real time and space is essential to delineate how cellular dynamics and the signaling circuit are orchestrated. Spatial regulation of conformational dynamics and structural plasticity of protein interactions is required to rewire signaling circuitry in response to extracellular cues. We introduce a method for optically imaging intracellular protein interactions at nanometer spatial resolution in live cells, using photoactivatable complementary fluorescent (PACF) proteins. Subsets of complementary fluorescent protein molecules were activated, localized, and then bleached; this was followed by the assembly of superresolution images from aggregate position of sum interactive molecules. Using PACF, we obtained precise localization of dynamic microtubule plus-end hub protein EB1 dimers and their distinct distributions at the leading edges and in the cell bodies of migrating cells. We further delineated the structure-function relationship of EB1 by generating EB1-PACF dimers (EB1(wt):EB1(wt), EB1(wt):EB1(mt), and EB1(mt):EB1(mt)) and imaging their precise localizations in culture cells. Surprisingly, our analyses revealed critical role of a previously uncharacterized EB1 linker region in tracking microtubule plus ends in live cells. Thus PACF provides a unique approach to delineating spatial dynamics of homo- or heterodimerized proteins at the nanometer scale and establishes a platform to report the precise regulation of protein interactions in space and time in live cells.
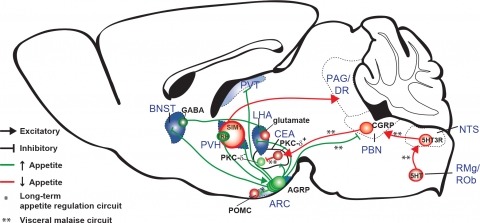
New tools for mapping and manipulating molecularly defined neural circuits have improved understanding of how the central nervous system regulates appetite. Studies focused on AGRP neurons, a starvation-sensitive hypothalamic population, have identified multiple circuit elements that can elicit or suppress feeding behavior. Distinct axon projections of this neuron population point to different circuits that regulate long-term appetite, short-term feeding, or visceral malaise-mediated anorexia. Here, we review recent studies examining these neural circuits that control food intake. © 2014 S. Karger AG, Basel.
Successful neurological rehabilitation depends on accurate diagnosis, effective treatment, and quantitative evaluation. Neural coding, a technology for interpretation of functional and structural information of the nervous system, has contributed to the advancements in neuroimaging, brain-machine interface (BMI), and design of training devices for rehabilitation purposes. In this review, we summarized the latest breakthroughs in neuroimaging from microscale to macroscale levels with potential diagnostic applications for rehabilitation. We also reviewed the achievements in electrocorticography (ECoG) coding with both animal models and human beings for BMI design, electromyography (EMG) interpretation for interaction with external robotic systems, and robot-assisted quantitative evaluation on the progress of rehabilitation programs. Future rehabilitation would be more home-based, automatic, and self-served by patients. Further investigations and breakthroughs are mainly needed in aspects of improving the computational efficiency in neuroimaging and multichannel ECoG by selection of localized neuroinformatics, validation of the effectiveness in BMI guided rehabilitation programs, and simplification of the system operation in training devices.
Pheromones, chemical signals that convey social information, mediate many insect social behaviors, including navigation and aggregation. Several studies have suggested that behavior during the immature larval stages of Drosophila development is influenced by pheromones, but none of these compounds or the pheromone-receptor neurons that sense them have been identified. Here we report a larval pheromone-signaling pathway. We found that larvae produce two novel long-chain fatty acids that are attractive to other larvae. We identified a single larval chemosensory neuron that detects these molecules. Two members of the pickpocket family of DEG/ENaC channel subunits (ppk23 and ppk29) are required to respond to these pheromones. This pheromone system is evolving quickly, since the larval exudates of D. simulans, the sister species of D. melanogaster, are not attractive to other larvae. Our results define a new pheromone signaling system in Drosophila that shares characteristics with pheromone systems in a wide diversity of insects.