Filter
Associated Lab
- Aguilera Castrejon Lab (15) Apply Aguilera Castrejon Lab filter
- Ahrens Lab (11) Apply Ahrens Lab filter
- Baker Lab (19) Apply Baker Lab filter
- Betzig Lab (12) Apply Betzig Lab filter
- Beyene Lab (5) Apply Beyene Lab filter
- Bock Lab (3) Apply Bock Lab filter
- Branson Lab (3) Apply Branson Lab filter
- Card Lab (6) Apply Card Lab filter
- Cardona Lab (19) Apply Cardona Lab filter
- Chklovskii Lab (3) Apply Chklovskii Lab filter
- Clapham Lab (1) Apply Clapham Lab filter
- Darshan Lab (4) Apply Darshan Lab filter
- Dennis Lab (1) Apply Dennis Lab filter
- Dickson Lab (14) Apply Dickson Lab filter
- Druckmann Lab (4) Apply Druckmann Lab filter
- Dudman Lab (12) Apply Dudman Lab filter
- Egnor Lab (7) Apply Egnor Lab filter
- Espinosa Medina Lab (4) Apply Espinosa Medina Lab filter
- Fetter Lab (10) Apply Fetter Lab filter
- Fitzgerald Lab (13) Apply Fitzgerald Lab filter
- Gonen Lab (32) Apply Gonen Lab filter
- Grigorieff Lab (28) Apply Grigorieff Lab filter
- Harris Lab (10) Apply Harris Lab filter
- Heberlein Lab (81) Apply Heberlein Lab filter
- Hermundstad Lab (4) Apply Hermundstad Lab filter
- Hess Lab (3) Apply Hess Lab filter
- Jayaraman Lab (4) Apply Jayaraman Lab filter
- Johnson Lab (5) Apply Johnson Lab filter
- Kainmueller Lab (19) Apply Kainmueller Lab filter
- Karpova Lab (1) Apply Karpova Lab filter
- Keleman Lab (5) Apply Keleman Lab filter
- Keller Lab (15) Apply Keller Lab filter
- Koay Lab (16) Apply Koay Lab filter
- Lavis Lab (12) Apply Lavis Lab filter
- Lee (Albert) Lab (5) Apply Lee (Albert) Lab filter
- Leonardo Lab (4) Apply Leonardo Lab filter
- Li Lab (24) Apply Li Lab filter
- Lippincott-Schwartz Lab (72) Apply Lippincott-Schwartz Lab filter
- Liu (Yin) Lab (5) Apply Liu (Yin) Lab filter
- Liu (Zhe) Lab (5) Apply Liu (Zhe) Lab filter
- Looger Lab (1) Apply Looger Lab filter
- Magee Lab (18) Apply Magee Lab filter
- Menon Lab (6) Apply Menon Lab filter
- Murphy Lab (7) Apply Murphy Lab filter
- O'Shea Lab (1) Apply O'Shea Lab filter
- Otopalik Lab (12) Apply Otopalik Lab filter
- Pachitariu Lab (12) Apply Pachitariu Lab filter
- Pastalkova Lab (13) Apply Pastalkova Lab filter
- Pavlopoulos Lab (12) Apply Pavlopoulos Lab filter
- Pedram Lab (11) Apply Pedram Lab filter
- Reiser Lab (6) Apply Reiser Lab filter
- Riddiford Lab (24) Apply Riddiford Lab filter
- Romani Lab (12) Apply Romani Lab filter
- Rubin Lab (38) Apply Rubin Lab filter
- Saalfeld Lab (17) Apply Saalfeld Lab filter
- Satou Lab (15) Apply Satou Lab filter
- Schreiter Lab (17) Apply Schreiter Lab filter
- Sgro Lab (20) Apply Sgro Lab filter
- Simpson Lab (5) Apply Simpson Lab filter
- Singer Lab (43) Apply Singer Lab filter
- Spruston Lab (36) Apply Spruston Lab filter
- Stern Lab (83) Apply Stern Lab filter
- Sternson Lab (7) Apply Sternson Lab filter
- Stringer Lab (3) Apply Stringer Lab filter
- Svoboda Lab (4) Apply Svoboda Lab filter
- Tebo Lab (24) Apply Tebo Lab filter
- Tillberg Lab (3) Apply Tillberg Lab filter
- Tjian Lab (47) Apply Tjian Lab filter
- Truman Lab (30) Apply Truman Lab filter
- Turaga Lab (12) Apply Turaga Lab filter
- Turner Lab (11) Apply Turner Lab filter
- Wang (Shaohe) Lab (19) Apply Wang (Shaohe) Lab filter
- Wu Lab (1) Apply Wu Lab filter
- Zlatic Lab (2) Apply Zlatic Lab filter
- Zuker Lab (20) Apply Zuker Lab filter
Associated Project Team
Publication Date
- 2024 (1) Apply 2024 filter
- 2023 (1) Apply 2023 filter
- 2022 (26) Apply 2022 filter
- 2021 (19) Apply 2021 filter
- 2020 (19) Apply 2020 filter
- 2019 (25) Apply 2019 filter
- 2018 (26) Apply 2018 filter
- 2017 (31) Apply 2017 filter
- 2016 (18) Apply 2016 filter
- 2015 (57) Apply 2015 filter
- 2014 (46) Apply 2014 filter
- 2013 (58) Apply 2013 filter
- 2012 (78) Apply 2012 filter
- 2011 (92) Apply 2011 filter
- 2010 (100) Apply 2010 filter
- 2009 (102) Apply 2009 filter
- 2008 (100) Apply 2008 filter
- 2007 (85) Apply 2007 filter
- 2006 (89) Apply 2006 filter
- 2005 (67) Apply 2005 filter
- 2004 (57) Apply 2004 filter
- 2003 (58) Apply 2003 filter
- 2002 (39) Apply 2002 filter
- 2001 (28) Apply 2001 filter
- 2000 (29) Apply 2000 filter
- 1999 (14) Apply 1999 filter
- 1998 (18) Apply 1998 filter
- 1997 (16) Apply 1997 filter
- 1996 (10) Apply 1996 filter
- 1995 (18) Apply 1995 filter
- 1994 (12) Apply 1994 filter
- 1993 (10) Apply 1993 filter
- 1992 (6) Apply 1992 filter
- 1991 (11) Apply 1991 filter
- 1990 (11) Apply 1990 filter
- 1989 (6) Apply 1989 filter
- 1988 (1) Apply 1988 filter
- 1987 (7) Apply 1987 filter
- 1986 (4) Apply 1986 filter
- 1985 (5) Apply 1985 filter
- 1984 (2) Apply 1984 filter
- 1983 (2) Apply 1983 filter
- 1982 (3) Apply 1982 filter
- 1981 (3) Apply 1981 filter
- 1980 (1) Apply 1980 filter
- 1979 (1) Apply 1979 filter
- 1976 (2) Apply 1976 filter
- 1973 (1) Apply 1973 filter
- 1970 (1) Apply 1970 filter
- 1967 (1) Apply 1967 filter
Type of Publication
- Remove Non-Janelia filter Non-Janelia
1417 Publications
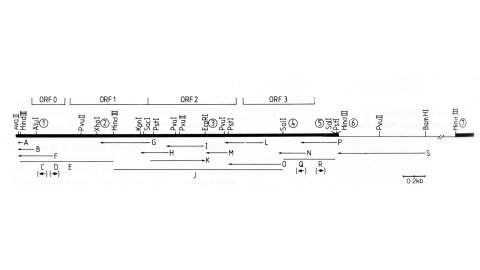
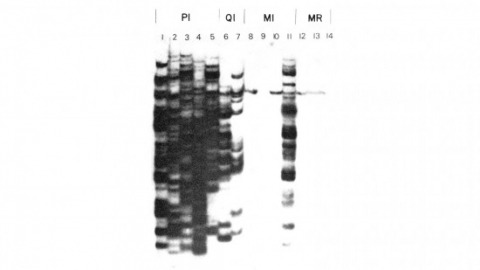
Showing 1401-1410 of 1417 resultsWe have made a P-element derivative called Pc[ry], which carries the selectable marker gene rosy, but which acts like a nondefective, intact P element. It transposes autonomously into the germline chromosomes of an M-strain Drosophila embryo and it mobilizes in trans the defective P elements of the singed-weak allele. Frameshift mutations introduced into any of the four major open reading frames of the P sequence were each sufficient to eliminate the transposase activity, but none affected signals required in cis for transposition of the element. Complementation tests between pairs of mutant elements suggest that a single polypeptide comprises the transposase. We have examined transcripts of P elements both from natural P strains and from lines containing only nondefective Pc[ry] elements, and have identified two RNA species that appear to be specific for autonomous elements.
The major site of in vivo transcriptional initiation for both heavy and light strands of human mitochondrial DNA is the displacement-loop region. Transcripts synthesized in vitro by human mitochondrial RNA polymerase were mapped to the nucleotide level and have identical 5’ end map positions to those reported for in vivo primary transcripts. An ordered series of deletion clones, whose template sequences were truncated at either the 5’ or 3’ end, was used to identify the precise mitochondrial DNA sequence required for initiation of transcription. The data provide a definitive assignment of the promoter for heavy-strand transcription occurring within -16 to +7 of the transcriptional start site 16 nucleotides upstream of the 5’ end of the gene for tRNAPhe and of the promoter for light-strand transcription occurring within -28 to +16 of the transcriptional start site at the 5’ end of "7S RNA." Within each control sequence is a candidate promoter whose consensus sequence is 5’-CANACC(G)CC(A)AAAGAPyA-3’ and in both cases transcriptional initiation occurs within six to eight nucleotides of the 3’ end of this sequence. The transcriptional start site is an integral part of each promoter and each promoter can function in the absence of the other.
Thirty-six isogenic D. melanogaster strains that differed only in the chromosomal location of a 7.2 or an 8.1 kb DNA segment containing the (autosomal) rosy gene were constructed by P-element-mediated gene transfer. Since the flies were homozygous for a rosy- allele, rosy gene function in these indicated the influence of flanking sequences on gene expression. The tissue distribution of XDH activity in all the strains was normal. Each line exhibited a characteristic level of adult XDH-specific activity. The majority of these values were close to wild-type levels; however, the total variation in specific activity among the lines was nearly fivefold. Thus position effects influence expression of the rosy gene quantitatively but do not detectably alter tissue specificity. X-linked rosy insertions were expressed on average 1.6 times more activity in males than in females. Hence the gene acquires at least partial dosage compensation upon insertion into the X chromosome.
Exogenous DNA sequences were introduced into the Drosophila germ line. A rosy transposon (ry1), constructed by inserting a chromosomal DNA fragment containing the wild-type rosy gene into a P transposable element, transformed germ line cells in 20 to 50 percent of the injected rosy mutant embryos. Transformants contained one or two copies of chromosomally integrated, intact ry1 that were stably inherited in subsequent generations. These transformed flies had wild-type eye color indicating that the visible genetic defect in the host strain could be fully and permanently corrected by the transferred gene. To demonstrate the generality of this approach, a DNA segment that does not confer a recognizable phenotype on recipients was also transferred into germ line chromosomes.
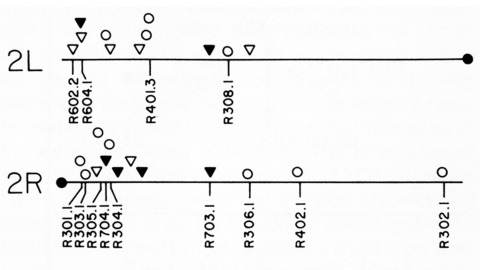
We have shown previously that four of five white mutant alleles arising in P-M dysgenic hybrids result from the insertion of strongly homologous DNA sequence elements. We have named these P elements. We report that P elements are present in 30-50 copies per haploid genome in all P strains examined and apparently are missing entirely from all M strains examined, with one exception. Furthermore, members of the P family apparently transpose frequently in P-M dysgenic hybrids; chromosomes descendant from P-M dysgenic hybrids frequently show newly acquired P elements. Finally, the strain-specific breakpoint hotspots for the rearrangement of the pi 2 P X chromosome occurring in P-M dysgenic hybrids are apparently sites of residence of P elements. These observations strongly support the P factor hypothesis for the mechanistic basis of P-M hybrid dysgenesis.
The transformer-2 (tra-2) locus is one of a set of regulatory loci that control sex determination in Drosophila melanogaster. Temperature-shift experiments with temperature-sensitive tra-2 mutants demonstrate that within single cell lineages tra-2+ function is required at several times, and probably continuously, during development for the occurrence of a series of determinative decisions necessary for female sexual differentiation. Analysis of the effects of tra-2 in the genital disc demonstrates that the tra-2+ function is necessary in females both to prevent male sexual differentiation and to permit female differentiation. These and other results support the model that the tra-2+ and tra+ loci act to control the expression of the bifunctional doublesex (dsx) locus.
The complete sequence of the 16,295 bp mouse L cell mitochondrial DNA genome has been determined. Genes for the 12S and 16S ribosomal RNAs; 22 tRNAs; cytochrome c oxidase subunits I, II and III; ATPase subunit 6; cytochrome b; and eight unidentified proteins have been located. The genome displays exceptional economy of organization, with tRNA genes interspersed between rRNA and protein-coding genes with zero or few noncoding nucleotides between coding sequences. Only two significant portions of the genome, the 879 nucleotide displacement-loop region containing the origin of heavy-strand replication and the 32 nucleotide origin of light-strand replication, do not encode a functional RNA species. All of the remaining nucleotide sequence serves as a defined coding function, with the exception of 32 nucleotides, of which 18 occur at the 5’ ends of open reading frames. Mouse mitochondrial DNA is unique in that the translational start codon is AUN, with any of the four nucleotides in the third position, whereas the only translational stop codon is the orthodox UAA. The mouse mitochondrial DNA genome is highly homologous in overall sequence and in gene organization to human mitochondrial DNA, with the descending order of conserved regions being tRNA genes; origin of light-strand replication; rRNA genes; known protein-coding genes; unidentified protein-coding genes; displacement-loop region.
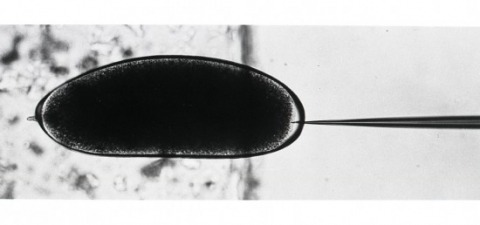
The antennae of male silk moths are extremely sensitive to the female sex pheromone such that a male moth can find a female up to 4.5 km away. This remarkable sensitivity is due to both the morphological and biochemical design of these antennae. Along the branches of the plumose antennae are the sensilla trichodea, each consisting of a hollow cuticular hair containing two unbranched dendrites bathed in a fluid, the receptor lymph ,3. The dendrites and receptor lymph are isolated from the haemolymph by a barrier of epidermal cells which secreted the cuticular hair. Pheromone molecules are thought to diffuse down 100 A-wide pore tubules through the cuticular wall and across the receptor lymph space to receptors located in the dendritic membrane. To prevent the accumulation of residual stimulant and hence sensory adaptation, the pheromone molecules are subsequently inactivated in an apparent two-step process of rapid ’early inactivation’ followed by much slower enzymatic degradation. The biochemistry involved in this sequence of events is largely unknown. We report here the identification of three proteins which interact with the pheromone of the wild silk moth Antheraea polyphemus: a pheromone-binding protein and a pheromone-degrading esterase, both uniquely located in the pheromone-sensitive sensilla; and a second esterase common to all cuticular tissues except the sensilla.
We describe the isolation of a cloned DNA segment carrying unique sequences from the white locus of Drosophila melanogaster. Sequences within the cloned segment are shown to hybridize in situ to the white locus region on the polytene chromosomes of both wild-type strains and strains carrying chromosomal rearrangements whose breakpoints bracket the white locus. We further show that two small deficiency mutations, deleting white locus genetic elements but not those of complementation groups contiguous to white, delete the genomic sequences corresponding to a portion of the cloned segment. The strategy we have employed to isolate this cloned segment exploits the existence of an allele at the white locus containing a copy of a previously cloned transposable, reiterated DNA sequence element. We describe a simple, rapid method for retrieving cloned segments carrying a copy of the transposable element together with contiguous sequences corresponding to this allele. The strategy described is potentially general and we discuss its application to the cloning of the DNA sequences of other genes in Drosophila, including those identified only by genetic analysis and for which no RNA product is known.