Filter
Associated Lab
- Ahrens Lab (1) Apply Ahrens Lab filter
- Aso Lab (2) Apply Aso Lab filter
- Baker Lab (1) Apply Baker Lab filter
- Betzig Lab (2) Apply Betzig Lab filter
- Branson Lab (1) Apply Branson Lab filter
- Cui Lab (1) Apply Cui Lab filter
- Freeman Lab (1) Apply Freeman Lab filter
- Harris Lab (1) Apply Harris Lab filter
- Heberlein Lab (1) Apply Heberlein Lab filter
- Keller Lab (1) Apply Keller Lab filter
- Lavis Lab (2) Apply Lavis Lab filter
- Leonardo Lab (2) Apply Leonardo Lab filter
- Lippincott-Schwartz Lab (2) Apply Lippincott-Schwartz Lab filter
- Liu (Zhe) Lab (1) Apply Liu (Zhe) Lab filter
- Looger Lab (1) Apply Looger Lab filter
- Riddiford Lab (1) Apply Riddiford Lab filter
- Rubin Lab (2) Apply Rubin Lab filter
- Scheffer Lab (1) Apply Scheffer Lab filter
- Singer Lab (1) Apply Singer Lab filter
- Spruston Lab (1) Apply Spruston Lab filter
- Stern Lab (3) Apply Stern Lab filter
- Sternson Lab (3) Apply Sternson Lab filter
- Tjian Lab (1) Apply Tjian Lab filter
Associated Project Team
Publication Date
- December 30, 2014 (1) Apply December 30, 2014 filter
- December 28, 2014 (1) Apply December 28, 2014 filter
- December 24, 2014 (1) Apply December 24, 2014 filter
- December 23, 2014 (4) Apply December 23, 2014 filter
- December 22, 2014 (6) Apply December 22, 2014 filter
- December 19, 2014 (1) Apply December 19, 2014 filter
- December 16, 2014 (2) Apply December 16, 2014 filter
- December 15, 2014 (1) Apply December 15, 2014 filter
- December 13, 2014 (2) Apply December 13, 2014 filter
- December 12, 2014 (3) Apply December 12, 2014 filter
- December 11, 2014 (1) Apply December 11, 2014 filter
- December 10, 2014 (2) Apply December 10, 2014 filter
- December 6, 2014 (1) Apply December 6, 2014 filter
- December 4, 2014 (4) Apply December 4, 2014 filter
- December 3, 2014 (1) Apply December 3, 2014 filter
- December 1, 2014 (4) Apply December 1, 2014 filter
- Remove December 2014 filter December 2014
Type of Publication
35 Publications
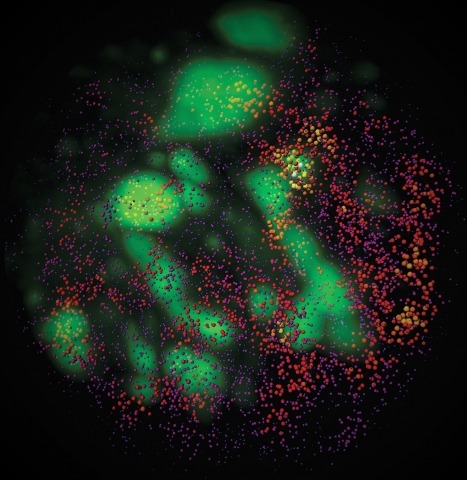
Showing 1-10 of 35 resultsCombinatorial cis-regulatory networks encoded in animal genomes represent the foundational gene expression mechanism for directing cell-fate commitment and maintenance of cell identity by transcription factors (TFs). However, the 3D spatial organization of cis-elements and how such sub-nuclear structures influence TF activity remain poorly understood. Here, we combine lattice light-sheet imaging, single-molecule tracking, numerical simulations, and ChIP-exo mapping to localize and functionally probe Sox2 enhancer-organization in living embryonic stem cells. Sox2 enhancers form 3D-clusters that are segregated from heterochromatin but overlap with a subset of Pol II enriched regions. Sox2 searches for specific binding targets via a 3D-diffusion dominant mode when shuttling long-distances between clusters while chromatin-bound states predominate within individual clusters. Thus, enhancer clustering may reduce global search efficiency but enables rapid local fine-tuning of TF search parameters. Our results suggest an integrated model linking cis-element 3D spatial distribution to local-versus-global target search modalities essential for regulating eukaryotic gene transcription.
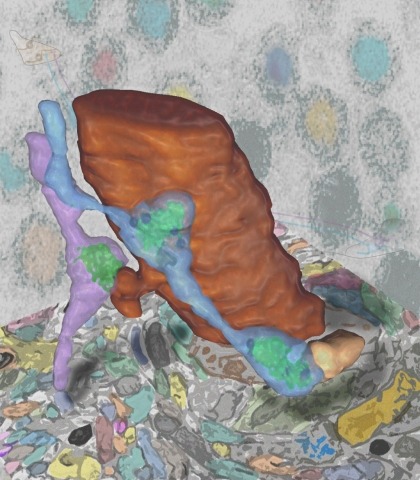
Synaptic connectivity and molecular composition provide a blueprint for information processing in neural circuits. Detailed structural analysis of neural circuits requires nanometer resolution, which can be obtained with serial-section electron microscopy. However, this technique remains challenging for reconstructing molecularly defined synapses. We used a genetically encoded synaptic marker for electron microscopy (GESEM) based on intra-vesicular generation of electron-dense labeling in axonal boutons. This approach allowed the identification of synapses from Cre recombinase-expressing or GAL4-expressing neurons in the mouse and fly with excellent preservation of ultrastructure. We applied this tool to visualize long-range connectivity of AGRP and POMC neurons in the mouse, two molecularly defined hypothalamic populations that are important for feeding behavior. Combining selective ultrastructural reconstruction of neuropil with functional and viral circuit mapping, we characterized some basic features of circuit organization for axon projections of these cell types. Our findings demonstrate that GESEM labeling enables long-range connectomics with molecularly defined cell types.
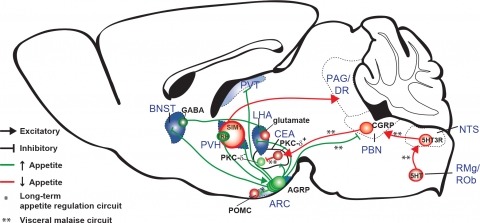
New tools for mapping and manipulating molecularly defined neural circuits have improved understanding of how the central nervous system regulates appetite. Studies focused on AGRP neurons, a starvation-sensitive hypothalamic population, have identified multiple circuit elements that can elicit or suppress feeding behavior. Distinct axon projections of this neuron population point to different circuits that regulate long-term appetite, short-term feeding, or visceral malaise-mediated anorexia. Here, we review recent studies examining these neural circuits that control food intake. © 2014 S. Karger AG, Basel.
Aphids evolved novel cells, called bacteriocytes, that differentiate specifically to harbour the obligatory mutualistic endosymbiotic bacteria Buchnera aphidicola. The genome of the host aphid Acyrthosiphon pisum contains many orphan genes that display no similarity with genes found in other sequenced organisms, prompting us to hypothesize that some of these orphan genes are related to lineage-specific traits, such as symbiosis. We conducted deep sequencing of bacteriocytes mRNA followed by whole mount in situ hybridizations of over-represented transcripts encoding aphid-specific orphan proteins. We identified a novel class of genes that encode small proteins with signal peptides, which are often cysteine-rich, that are over-represented in bacteriocytes. These genes are first expressed at a developmental time point coincident with the incorporation of symbionts strictly in the cells that contribute to the bacteriocyte and this bacteriocyte-specific expression is maintained throughout the aphid's life. The expression pattern suggests that recently evolved secretion proteins act within bacteriocytes, perhaps to mediate the symbiosis with beneficial bacterial partners, which is reminiscent of the evolution of novel cysteine-rich secreted proteins of leguminous plants that regulate nitrogen-fixing endosymbionts.
The way the hippocampus processes information and encodes memories in the form of "cell assemblies" is likely determined in part by how its circuits are wired up during development. In this issue, Xu et al. now provide new insight into how neurons arising from a single common precursor migrate to their final destination and form functionally synchronous ensembles.
Muscular hydrostats (such as mollusks), and fluid-filled animals (such as annelids), can exploit their constant-volume tissues to transfer forces and displacements in predictable ways, much as articulated animals use hinges and levers. Although larval insects contain pressurized fluids, they also have internal air tubes that are compressible and, as a result, they have more uncontrolled degrees of freedom. Therefore, the mechanisms by which larval insects control their movements are expected to reveal useful strategies for designing soft biomimetic robots. Using caterpillars as a tractable model system, it is now possible to identify the biomechanical and neural strategies for controlling movements in such highly deformable animals. For example, the tobacco hornworm, Manduca sexta, can stiffen its body by increasing muscular tension (and therefore body pressure) but the internal cavity (hemocoel) is not iso-barometric, nor is pressure used to directly control the movements of its limbs. Instead, fluid and tissues flow within the hemocoel and the body is soft and flexible to conform to the substrate. Even the gut contributes to the biomechanics of locomotion; it is decoupled from the movements of the body wall and slides forward within the body cavity at the start of each step. During crawling the body is kept in tension for part of the stride and compressive forces are exerted on the substrate along the axis of the caterpillar, thereby using the environment as a skeleton. The timing of muscular activity suggests that crawling is coordinated by proleg-retractor motoneurons and that the large segmental muscles produce anterograde waves of lifting that do not require precise timing. This strategy produces a robust form of locomotion in which the kinematics changes little with orientation. In different species of caterpillar, the presence of prolegs on particular body segments is related to alternative kinematics such as "inching." This suggests a mechanism for the evolution of different gaits through changes in the usage of prolegs, rather than, through extensive alterations in the motor program controlling the body wall. Some of these findings are being used to design and test novel control-strategies for highly deformable robots. These "softworm" devices are providing new insights into the challenges faced by any soft animal navigating in a terrestrial environment.
The hemoglobinopathies, such as β-thalassemia and sickle cell anemia (SCA), are characterized by mutations of the β-globin gene resulting in either decreased or functionally abnormal hemoglobin (Hb) production. As bone marrow transplant is the only curative option for these patients, there is a strong need for new therapeutic approaches. Both β-thalassemia and SCA represent ideal targets for gene therapy since introduction of a normal β-globin gene can ameliorate the phenotype, as we and others have shown previously. Overcoming the developmental silencing of the fetal γ-globin gene represents an additional approach for the treatment of hemoglobinopathies. Here, we directly compare a recently established approach to activate the γ-globin gene using forced chromatin looping with pharmacologic approaches to raise γ-globin expression. The β-type globin genes are activated through dynamic interactions with a distal upstream enhancer, the locus control region (LCR). The LCR physically contacts the developmental stage appropriate globin gene via chromatin looping, a process partially dependent on the protein Ldb1. Previously, we have shown that tethering Ldb1 to the murine β-globin promoter with a custom designed zinc finger protein (ZF-Ldb1) can induce loop formation and β-globin transcription in an erythroid cell line (Deng et al., 2012). Further work showed that forced chromatin looping can be exploited to potently reactivate fetal globin gene expression in adult human erythroid cells (Deng et al., 2014). Here we compared the efficacy and toxicity of ZF-Ldb1 to pharmacologic compounds that induce HbF in cultured hematopoietic stem progenitor cell-derived erythroid cultures from normal and SCA donors. ZF-Ldb1 increased HbF synthesis in SCA erythroid cells (N=8) up to 86% and, concurrently, reduced sickle Hb (HbS) below 15%, consistent with previous studies of erythroid cells from normal probands. Preliminary results obtained from treating SCA specimens (N=3) show that the induction of HbF in cells treated with ZF-Ldb1 is twice as high (+35.55% ± 8.34%, at a dose of ~ one ZF-Ldb1 transgene copy per cell) as that observed using pomalidomide (+16.50% ± 14.57%, 20μM) and decitabine (+15.60% ± 12.36%, 0.5μM). Tranylcypromine and hydroxyurea showed the lowest HbF increase (+9.67% ± 3.26% and +5.06 ± 2.82%, 1.5μM and 150μM respectively). Importantly, decitabine and pomalidomide treatment lowered cell viability to 39% and 26%, respectively, while ZF-Ldb1 expressing cells retained normal viability similar to control populations. In related experiments, we are comparing the expression of a battery of genes known to regulate HbF levels (BCL11A, SOX6, KLF1 and C-Myb) in normal and SCA derived erythroid cells treated with ZF-Ldb1 or HbF inducers and compared to controls. Preliminary analyses indicate altered expression of KLF1 in SCA versus normal cells, consistent with a superior response of SCA cells to HbF induction. In conclusion, lentiviral-mediated ZF-Ldb1 gene transfer appears superior to pharmacologic compounds in terms of efficacy and cell viability further supporting suitability for the reactivation of HbF in SCA erythroid cells.
Similar morphological, physiological, and behavioral features have evolved independently in different species, a pattern known as convergence. It is known that morphological convergence can occur through changes in orthologous genes. In some cases of convergence, cis-regulatory changes generate parallel modifications in the expression patterns of orthologous genes. Our understanding of how changes in cis-regulatory regions contribute to convergence is hampered, usually, by a limited understanding of the global cis-regulatory structure of the evolving genes. Here we examine the genetic causes of a case of precise phenotypic convergence between Drosophila sechellia and Drosophila ezoana, species that diverged ~40 Mya. Previous studies revealed that changes in multiple transcriptional enhancers of shavenbaby (svb, a transcript of the ovo locus) caused phenotypic evolution in the D. sechellia lineage. It has also been shown that the convergent phenotype of D. ezoana was likely caused by cis-regulatory evolution of svb. Here we show that the large-scale cis-regulatory architecture of svb is conserved between these Drosophila species. Furthermore, we show that the D. ezoana orthologs of the evolved D. sechellia enhancers have also evolved expression patterns that correlate precisely with the changes in the phenotype. Our results suggest that phenotypic convergence resulted from multiple noncoding changes that occurred in parallel in the D. sechellia and D. ezoana lineages.
Pheromones, chemical signals that convey social information, mediate many insect social behaviors, including navigation and aggregation. Several studies have suggested that behavior during the immature larval stages of Drosophila development is influenced by pheromones, but none of these compounds or the pheromone-receptor neurons that sense them have been identified. Here we report a larval pheromone-signaling pathway. We found that larvae produce two novel long-chain fatty acids that are attractive to other larvae. We identified a single larval chemosensory neuron that detects these molecules. Two members of the pickpocket family of DEG/ENaC channel subunits (ppk23 and ppk29) are required to respond to these pheromones. This pheromone system is evolving quickly, since the larval exudates of D. simulans, the sister species of D. melanogaster, are not attractive to other larvae. Our results define a new pheromone signaling system in Drosophila that shares characteristics with pheromone systems in a wide diversity of insects.