Filter
Associated Lab
- Aso Lab (2) Apply Aso Lab filter
- Bock Lab (1) Apply Bock Lab filter
- Card Lab (1) Apply Card Lab filter
- Cardona Lab (1) Apply Cardona Lab filter
- Fetter Lab (4) Apply Fetter Lab filter
- Funke Lab (1) Apply Funke Lab filter
- Harris Lab (1) Apply Harris Lab filter
- Hess Lab (10) Apply Hess Lab filter
- Jayaraman Lab (1) Apply Jayaraman Lab filter
- Rubin Lab (7) Apply Rubin Lab filter
- Saalfeld Lab (2) Apply Saalfeld Lab filter
- Scheffer Lab (36) Apply Scheffer Lab filter
- Sternson Lab (1) Apply Sternson Lab filter
- Turner Lab (1) Apply Turner Lab filter
Associated Project Team
Publication Date
- 2023 (2) Apply 2023 filter
- 2022 (1) Apply 2022 filter
- 2021 (2) Apply 2021 filter
- 2020 (1) Apply 2020 filter
- 2019 (2) Apply 2019 filter
- 2018 (4) Apply 2018 filter
- 2017 (4) Apply 2017 filter
- 2015 (2) Apply 2015 filter
- 2014 (7) Apply 2014 filter
- 2013 (3) Apply 2013 filter
- 2012 (3) Apply 2012 filter
- 2011 (2) Apply 2011 filter
- 2010 (3) Apply 2010 filter
Type of Publication
36 Publications
Showing 11-20 of 36 resultsNew reconstruction techniques are generating connectomes of unprecedented size. These must be analyzed to generate human comprehensible results. The analyses being used fall into three general categories. The first is interactive tools used during reconstruction, to help guide the effort, look for possible errors, identify potential cell classes, and answer other preliminary questions. The second type of analysis is support for formal documents such as papers and theses. Scientific norms here require that the data be archived and accessible, and the analysis reproducible. In contrast to some other "omic" fields such as genomics, where a few specific analyses dominate usage, connectomics is rapidly evolving and the analyses used are often specific to the connectome being analyzed. These analyses are typically performed in a variety of conventional programming language, such as Matlab, R, Python, or C++, and read the connectomic data either from a file or through database queries, neither of which are standardized. In the short term we see no alternative to the use of specific analyses, so the best that can be done is to publish the analysis code, and the interface by which it reads connectomic data. A similar situation exists for archiving connectome data. Each group independently makes their data available, but there is no standardized format and long-term accessibility is neither enforced nor funded. In the long term, as connectomics becomes more common, a natural evolution would be a central facility for storing and querying connectomic data, playing a role similar to the National Center for Biotechnology Information for genomes. The final form of analysis is the import of connectome data into downstream tools such as neural simulation or machine learning. In this process, there are two main problems that need to be addressed. First, the reconstructed circuits contain huge amounts of detail, which must be intelligently reduced to a form the downstream tools can use. Second, much of the data needed for these downstream operations must be obtained by other methods (such as genetic or optical) and must be merged with the extracted connectome.
Fruit flies (Drosophila melanogaster) are small insects, with correspondingly small power budgets. Despite this, they perform sophisticated neural computations in real time. Careful study of these insects is revealing how some of these circuits work. Insights from these systems might be helpful in designing other low power circuits.
Connectomes derived from volume EM imaging of the brain can generate detailed physical models of every neuron, and simulators such as NEURON or GENESIS are designed to work with such models. In principal, combining these technologies, plus transmitter and channel models, should allow detailed and accurate simulation of real neural circuits. Here we experiment with this combination, using a well-studied system (motion detection in Drosophila. Since simulation requires both the physical geometry (which we have) and the models of the synapses (which are not currently available), we built approximate synapses corresponding to their known and estimated function. Once we did so, we reproduced direction selectivity in T4 cells, one of the main functions of this neural circuit. This verified the basic functionality of both extraction and simulations, and provided a biologically relevant computation we could use in further experiments. We then compared models with different degrees of physical realism, from full detailed models down to models consisting of a single node, to examine the tradeoff of simulation resources required versus accuracy achieved. Our results show that much simpler models may be adequate, at least in the case of medulla neurons in Drosophila. Such models can be easily derived from fully detailed models, and result in simulations that are much smaller, much faster, and accurate enough for many purposes. Biologically, we show that a lumped neuron model reproduces the main motion detector operation, confirming the result of Gruntman, that dendritic compution is not required for this function.
Understanding memory formation, storage and retrieval requires knowledge of the underlying neuronal circuits. In Drosophila, the mushroom body (MB) is the major site of associative learning. We reconstructed the morphologies and synaptic connections of all 983 neurons within the three functional units, or compartments, that compose the adult MB’s α lobe, using a dataset of isotropic 8-nm voxels collected by focused ion-beam milling scanning electron microscopy. We found that Kenyon cells (KCs), whose sparse activity encodes sensory information, each make multiple en passant synapses to MB output neurons (MBONs) in each compartment. Some MBONs have inputs from all KCs, while others differentially sample sensory modalities. Only six percent of KC>MBON synapses receive a direct synapse from a dopaminergic neuron (DAN). We identified two unanticipated classes of synapses, KC>DAN and DAN>MBON. DAN activation produces a slow depolarization of the MBON in these DAN>MBON synapses and can weaken memory recall.
Analysing computations in neural circuits often uses simplified models because the actual neuronal implementation is not known. For example, a problem in vision, how the eye detects image motion, has long been analysed using Hassenstein-Reichardt (HR) detector or Barlow-Levick (BL) models. These both simulate motion detection well, but the exact neuronal circuits undertaking these tasks remain elusive. We reconstructed a comprehensive connectome of the circuits of Drosophila's motion-sensing T4 cells using a novel EM technique. We uncover complex T4 inputs and reveal that putative excitatory inputs cluster at T4's dendrite shafts, while inhibitory inputs localize to the bases. Consistent with our previous study, we reveal that Mi1 and Tm3 cells provide most synaptic contacts onto T4. We are, however, unable to reproduce the spatial offset between these cells reported previously. Our comprehensive connectome reveals complex circuits that include candidate anatomical substrates for both HR and BL types of motion detectors.
We reconstructed the synaptic circuits of seven columns in the second neuropil or medulla behind the fly's compound eye. These neurons embody some of the most stereotyped circuits in one of the most miniaturized of animal brains. The reconstructions allow us, for the first time to our knowledge, to study variations between circuits in the medulla's neighboring columns. This variation in the number of synapses and the types of their synaptic partners has previously been little addressed because methods that visualize multiple circuits have not resolved detailed connections, and existing connectomic studies, which can see such connections, have not so far examined multiple reconstructions of the same circuit. Here, we address the omission by comparing the circuits common to all seven columns to assess variation in their connection strengths and the resultant rates of several different and distinct types of connection error. Error rates reveal that, overall, <1% of contacts are not part of a consensus circuit, and we classify those contacts that supplement (E+) or are missing from it (E-). Autapses, in which the same cell is both presynaptic and postsynaptic at the same synapse, are occasionally seen; two cells in particular, Dm9 and Mi1, form ≥20-fold more autapses than do other neurons. These results delimit the accuracy of developmental events that establish and normally maintain synaptic circuits with such precision, and thereby address the operation of such circuits. They also establish a precedent for error rates that will be required in the new science of connectomics.
Synaptic circuits for identified behaviors in the Drosophila brain have typically been considered from either a developmental or functional perspective without reference to how the circuits might have been inherited from ancestral forms. For example, two candidate pathways for ON- and OFF-edge motion detection in the visual system act via circuits that use respectively either T4 or T5, two cell types of the fourth neuropil, or lobula plate (LOP), that exhibit narrow-field direction-selective responses and provide input to wide-field tangential neurons. T4 or T5 both have four subtypes that terminate one each in the four strata of the LOP. Representatives are reported in a wide range of Diptera, and both cell types exhibit various similarities in: (1) the morphology of their dendritic arbors; (2) their four morphological and functional subtypes; (3) their cholinergic profile in Drosophila; (4) their input from the pathways of L3 cells in the first neuropil, or lamina (LA), and by one of a pair of LA cells, L1 (to the T4 pathway) and L2 (to the T5 pathway); and (5) their innervation by a single, wide-field contralateral tangential neuron from the central brain. Progenitors of both also express the gene atonal early in their proliferation from the inner anlage of the developing optic lobe, being alone among many other cell type progeny to do so. Yet T4 receives input in the second neuropil, or medulla (ME), and T5 in the third neuropil or lobula (LO). Here we suggest that these two cell types were originally one, that their ancestral cell population duplicated and split to innervate separate ME and LO neuropils, and that a fiber crossing-the internal chiasma-arose between the two neuropils. The split most plausibly occurred, we suggest, with the formation of the LO as a new neuropil that formed when it separated from its ancestral neuropil to leave the ME, suggesting additionally that ME input neurons to T4 and T5 may also have had a common origin.
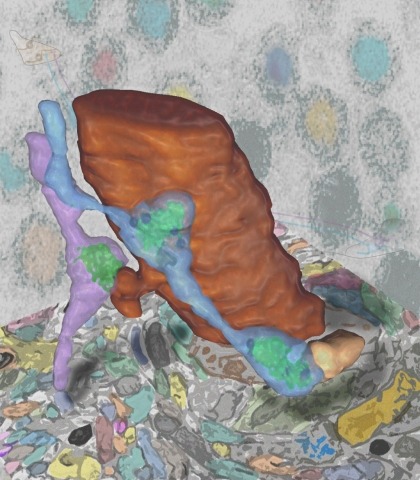
Synaptic connectivity and molecular composition provide a blueprint for information processing in neural circuits. Detailed structural analysis of neural circuits requires nanometer resolution, which can be obtained with serial-section electron microscopy. However, this technique remains challenging for reconstructing molecularly defined synapses. We used a genetically encoded synaptic marker for electron microscopy (GESEM) based on intra-vesicular generation of electron-dense labeling in axonal boutons. This approach allowed the identification of synapses from Cre recombinase-expressing or GAL4-expressing neurons in the mouse and fly with excellent preservation of ultrastructure. We applied this tool to visualize long-range connectivity of AGRP and POMC neurons in the mouse, two molecularly defined hypothalamic populations that are important for feeding behavior. Combining selective ultrastructural reconstruction of neuropil with functional and viral circuit mapping, we characterized some basic features of circuit organization for axon projections of these cell types. Our findings demonstrate that GESEM labeling enables long-range connectomics with molecularly defined cell types.
Reconstructing neuronal circuits at the level of synapses is a central problem in neuroscience and becoming a focus of the emerging field of connectomics. To date, electron microscopy (EM) is the most proven technique for identifying and quantifying synaptic connections. As advances in EM make acquiring larger datasets possible, subsequent manual synapse identification ({\em i.e.}, proofreading) for deciphering a connectome becomes a major time bottleneck. Here we introduce a large-scale, high-throughput, and semi-automated methodology to efficiently identify synapses. We successfully applied our methodology to the Drosophila medulla optic lobe, annotating many more synapses than previous connectome efforts. Our approaches are extensible and will make the often complicated process of synapse identification accessible to a wider-community of potential proofreaders.