Main Menu (Mobile)- Block
- Overview
-
Support Teams
- Overview
- Anatomy and Histology
- Cryo-Electron Microscopy
- Electron Microscopy
- Flow Cytometry
- Gene Targeting and Transgenics
- High Performance Computing
- Immortalized Cell Line Culture
- Integrative Imaging
- Invertebrate Shared Resource
- Janelia Experimental Technology
- Mass Spectrometry
- Media Prep
- Molecular Genomics
- Stem Cell & Primary Culture
- Project Pipeline Support
- Project Technical Resources
- Quantitative Genomics
- Scientific Computing
- Viral Tools
- Vivarium
- Open Science
- You + Janelia
- About Us
Main Menu - Block
- Overview
- Anatomy and Histology
- Cryo-Electron Microscopy
- Electron Microscopy
- Flow Cytometry
- Gene Targeting and Transgenics
- High Performance Computing
- Immortalized Cell Line Culture
- Integrative Imaging
- Invertebrate Shared Resource
- Janelia Experimental Technology
- Mass Spectrometry
- Media Prep
- Molecular Genomics
- Stem Cell & Primary Culture
- Project Pipeline Support
- Project Technical Resources
- Quantitative Genomics
- Scientific Computing
- Viral Tools
- Vivarium
Efficient processing and analysis of large-scale light-sheet microscopy data
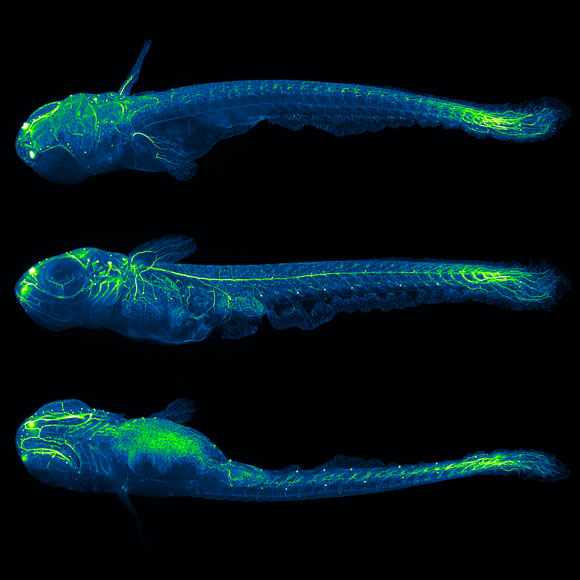
Light-sheet microscopy is a powerful method for imaging the development and function of complex biological systems at high spatiotemporal resolution and over long time scales. Such experiments typically generate terabytes of multidimensional image data, and thus they demand efficient computational solutions for data management, processing, and analysis. We present protocols and software to tackle these steps, focusing on the imaging-based study of animal development. Our protocols facilitate (i) high-speed lossless data compression and content-based multiview image fusion optimized for multicore CPU architectures, reducing image data size 30–500-fold; (ii) automated large-scale cell tracking and segmentation; and (iii) visualization, editing, and annotation of multiterabyte image data and cell-lineage reconstructions with tens of millions of data points. These software modules are open source. They provide high data throughput using a single computer workstation and are readily applicable to a wide spectrum of biological model systems.
The light-sheet microscopy image processing and analysis framework publication is available from the literature section (Amat, Höckendorf, Wan, Lemon, McDole and Keller 2015, Nature Protocols).
By clicking on the links on the right of the page to download the software, you agree to the terms of the Janelia Open-Source Software License linked here. The Github repository is also provided at right.
Publications
Janelia Collaborators
Janelia Open-Source Sofware License Agreement
By downloading this software, you agree to the terms of the Janelia Open-Source Software License linked here.