Filter
Associated Lab
- Aguilera Castrejon Lab (19) Apply Aguilera Castrejon Lab filter
- Ahrens Lab (73) Apply Ahrens Lab filter
- Aso Lab (42) Apply Aso Lab filter
- Baker Lab (38) Apply Baker Lab filter
- Betzig Lab (116) Apply Betzig Lab filter
- Beyene Lab (15) Apply Beyene Lab filter
- Bock Lab (17) Apply Bock Lab filter
- Branson Lab (55) Apply Branson Lab filter
- Card Lab (43) Apply Card Lab filter
- Cardona Lab (64) Apply Cardona Lab filter
- Chklovskii Lab (13) Apply Chklovskii Lab filter
- Clapham Lab (16) Apply Clapham Lab filter
- Cui Lab (19) Apply Cui Lab filter
- Darshan Lab (12) Apply Darshan Lab filter
- Dennis Lab (3) Apply Dennis Lab filter
- Dickson Lab (46) Apply Dickson Lab filter
- Druckmann Lab (25) Apply Druckmann Lab filter
- Dudman Lab (56) Apply Dudman Lab filter
- Eddy/Rivas Lab (30) Apply Eddy/Rivas Lab filter
- Egnor Lab (11) Apply Egnor Lab filter
- Espinosa Medina Lab (23) Apply Espinosa Medina Lab filter
- Feliciano Lab (12) Apply Feliciano Lab filter
- Fetter Lab (41) Apply Fetter Lab filter
- FIB-SEM Technology (1) Apply FIB-SEM Technology filter
- Fitzgerald Lab (30) Apply Fitzgerald Lab filter
- Freeman Lab (15) Apply Freeman Lab filter
- Funke Lab (46) Apply Funke Lab filter
- Gonen Lab (91) Apply Gonen Lab filter
- Grigorieff Lab (62) Apply Grigorieff Lab filter
- Harris Lab (65) Apply Harris Lab filter
- Heberlein Lab (94) Apply Heberlein Lab filter
- Hermundstad Lab (30) Apply Hermundstad Lab filter
- Hess Lab (80) Apply Hess Lab filter
- Ilanges Lab (4) Apply Ilanges Lab filter
- Jayaraman Lab (48) Apply Jayaraman Lab filter
- Ji Lab (33) Apply Ji Lab filter
- Johnson Lab (6) Apply Johnson Lab filter
- Kainmueller Lab (19) Apply Kainmueller Lab filter
- Karpova Lab (15) Apply Karpova Lab filter
- Keleman Lab (13) Apply Keleman Lab filter
- Keller Lab (76) Apply Keller Lab filter
- Koay Lab (20) Apply Koay Lab filter
- Lavis Lab (158) Apply Lavis Lab filter
- Lee (Albert) Lab (34) Apply Lee (Albert) Lab filter
- Leonardo Lab (23) Apply Leonardo Lab filter
- Li Lab (32) Apply Li Lab filter
- Lippincott-Schwartz Lab (180) Apply Lippincott-Schwartz Lab filter
- Liu (Yin) Lab (8) Apply Liu (Yin) Lab filter
- Liu (Zhe) Lab (65) Apply Liu (Zhe) Lab filter
- Looger Lab (138) Apply Looger Lab filter
- Magee Lab (49) Apply Magee Lab filter
- Menon Lab (18) Apply Menon Lab filter
- Murphy Lab (13) Apply Murphy Lab filter
- O'Shea Lab (8) Apply O'Shea Lab filter
- Otopalik Lab (13) Apply Otopalik Lab filter
- Pachitariu Lab (54) Apply Pachitariu Lab filter
- Pastalkova Lab (19) Apply Pastalkova Lab filter
- Pavlopoulos Lab (19) Apply Pavlopoulos Lab filter
- Pedram Lab (15) Apply Pedram Lab filter
- Podgorski Lab (16) Apply Podgorski Lab filter
- Reiser Lab (55) Apply Reiser Lab filter
- Riddiford Lab (44) Apply Riddiford Lab filter
- Romani Lab (51) Apply Romani Lab filter
- Rubin Lab (149) Apply Rubin Lab filter
- Saalfeld Lab (64) Apply Saalfeld Lab filter
- Satou Lab (18) Apply Satou Lab filter
- Scheffer Lab (38) Apply Scheffer Lab filter
- Schreiter Lab (70) Apply Schreiter Lab filter
- Schulze Lab (1) Apply Schulze Lab filter
- Sgro Lab (23) Apply Sgro Lab filter
- Shroff Lab (31) Apply Shroff Lab filter
- Simpson Lab (23) Apply Simpson Lab filter
- Singer Lab (80) Apply Singer Lab filter
- Spruston Lab (98) Apply Spruston Lab filter
- Stern Lab (160) Apply Stern Lab filter
- Sternson Lab (54) Apply Sternson Lab filter
- Stringer Lab (43) Apply Stringer Lab filter
- Svoboda Lab (136) Apply Svoboda Lab filter
- Tebo Lab (35) Apply Tebo Lab filter
- Tervo Lab (10) Apply Tervo Lab filter
- Tillberg Lab (22) Apply Tillberg Lab filter
- Tjian Lab (64) Apply Tjian Lab filter
- Truman Lab (88) Apply Truman Lab filter
- Turaga Lab (53) Apply Turaga Lab filter
- Turner Lab (38) Apply Turner Lab filter
- Vale Lab (8) Apply Vale Lab filter
- Voigts Lab (4) Apply Voigts Lab filter
- Wang (Meng) Lab (29) Apply Wang (Meng) Lab filter
- Wang (Shaohe) Lab (25) Apply Wang (Shaohe) Lab filter
- Wong-Campos Lab (1) Apply Wong-Campos Lab filter
- Wu Lab (9) Apply Wu Lab filter
- Zlatic Lab (28) Apply Zlatic Lab filter
- Zuker Lab (25) Apply Zuker Lab filter
Associated Project Team
- CellMap (13) Apply CellMap filter
- COSEM (3) Apply COSEM filter
- FIB-SEM Technology (5) Apply FIB-SEM Technology filter
- Fly Descending Interneuron (12) Apply Fly Descending Interneuron filter
- Fly Functional Connectome (14) Apply Fly Functional Connectome filter
- Fly Olympiad (5) Apply Fly Olympiad filter
- FlyEM (56) Apply FlyEM filter
- FlyLight (50) Apply FlyLight filter
- GENIE (47) Apply GENIE filter
- Integrative Imaging (9) Apply Integrative Imaging filter
- Larval Olympiad (2) Apply Larval Olympiad filter
- MouseLight (18) Apply MouseLight filter
- NeuroSeq (1) Apply NeuroSeq filter
- ThalamoSeq (1) Apply ThalamoSeq filter
- Tool Translation Team (T3) (29) Apply Tool Translation Team (T3) filter
- Transcription Imaging (49) Apply Transcription Imaging filter
Publication Date
- 2026 (74) Apply 2026 filter
- 2025 (222) Apply 2025 filter
- 2024 (210) Apply 2024 filter
- 2023 (158) Apply 2023 filter
- 2022 (192) Apply 2022 filter
- 2021 (194) Apply 2021 filter
- 2020 (196) Apply 2020 filter
- 2019 (202) Apply 2019 filter
- 2018 (232) Apply 2018 filter
- 2017 (217) Apply 2017 filter
- 2016 (209) Apply 2016 filter
- 2015 (252) Apply 2015 filter
- 2014 (236) Apply 2014 filter
- 2013 (194) Apply 2013 filter
- 2012 (190) Apply 2012 filter
- 2011 (190) Apply 2011 filter
- 2010 (161) Apply 2010 filter
- 2009 (158) Apply 2009 filter
- 2008 (140) Apply 2008 filter
- 2007 (106) Apply 2007 filter
- 2006 (92) Apply 2006 filter
- 2005 (67) Apply 2005 filter
- 2004 (57) Apply 2004 filter
- 2003 (58) Apply 2003 filter
- 2002 (39) Apply 2002 filter
- 2001 (28) Apply 2001 filter
- 2000 (29) Apply 2000 filter
- 1999 (14) Apply 1999 filter
- 1998 (18) Apply 1998 filter
- 1997 (16) Apply 1997 filter
- 1996 (10) Apply 1996 filter
- 1995 (18) Apply 1995 filter
- 1994 (12) Apply 1994 filter
- 1993 (10) Apply 1993 filter
- 1992 (6) Apply 1992 filter
- 1991 (11) Apply 1991 filter
- 1990 (11) Apply 1990 filter
- 1989 (6) Apply 1989 filter
- 1988 (1) Apply 1988 filter
- 1987 (7) Apply 1987 filter
- 1986 (4) Apply 1986 filter
- 1985 (5) Apply 1985 filter
- 1984 (2) Apply 1984 filter
- 1983 (2) Apply 1983 filter
- 1982 (3) Apply 1982 filter
- 1981 (3) Apply 1981 filter
- 1980 (1) Apply 1980 filter
- 1979 (1) Apply 1979 filter
- 1976 (2) Apply 1976 filter
- 1973 (1) Apply 1973 filter
- 1970 (1) Apply 1970 filter
- 1967 (1) Apply 1967 filter
Type of Publication
4269 Publications
Showing 3381-3390 of 4269 resultsProbing the architecture, mechanism, and dynamics of genome folding is fundamental to our understanding of genome function in homeostasis and disease. Most chromosome conformation capture studies dissect the genome architecture with population- and time-averaged snapshots and thus have limited capabilities to reveal 3D nuclear organization and dynamics at the single-cell level. Here, we discuss emerging imaging techniques ranging from light microscopy to electron microscopy that enable investigation of genome folding and dynamics at high spatial and temporal resolution. Results from these studies complement genomic data, unveiling principles underlying the spatial arrangement of the genome and its potential functional links to diverse biological activities in the nucleus.
Animal survival requires a functioning nervous system to develop during embryogenesis. Newborn neurons must assemble into circuits producing activity patterns capable of instructing behaviors. Elucidating how this process is coordinated requires new methods that follow maturation and activity of all cells across a developing circuit. We present an imaging method for comprehensively tracking neuron lineages, movements, molecular identities, and activity in the entire developing zebrafish spinal cord, from neurogenesis until the emergence of patterned activity instructing the earliest spontaneous motor behavior. We found that motoneurons are active first and form local patterned ensembles with neighboring neurons. These ensembles merge, synchronize globally after reaching a threshold size, and finally recruit commissural interneurons to orchestrate the left-right alternating patterns important for locomotion in vertebrates. Individual neurons undergo functional maturation stereotypically based on their birth time and anatomical origin. Our study provides a general strategy for reconstructing how functioning circuits emerge during embryogenesis.
Recognition of environmental cues is essential for the survival of all organisms. Transcriptional changes occur to enable the generation and function of the neural circuits underlying sensory perception. To gain insight into these changes, we generated single-cell transcriptomes of olfactory- (ORNs), thermo-, and hygro-sensory neurons at an early developmental and adult stage using single-cell and single-nucleus RNA sequencing. We discovered that ORNs maintain expression of the same olfactory receptors across development. Using receptor expression and computational approaches, we matched transcriptomic clusters corresponding to anatomically and physiologically defined neuron types across multiple developmental stages. We found that cell-type-specific transcriptomes partly reflected axon trajectory choices in development and sensory modality in adults. We uncovered stage-specific genes that could regulate the wiring and sensory responses of distinct ORN types. Collectively, our data reveal transcriptomic features of sensory neuron biology and provide a resource for future studies of their development and physiology.
The regulatory mechanisms by which neurons coordinate their physiology and connectivity are not well understood. The Drosophila olfactory receptor neurons (ORNs) provide an excellent system to investigate this question. Each ORN type expresses a unique olfactory receptor, or a combination thereof, and sends their axons to a stereotyped glomerulus. Using single-cell RNA sequencing, we identified 33 transcriptomic clusters for ORNs and mapped 20 to their glomerular types, demonstrating that transcriptomic clusters correspond well with anatomically and physiologically defined ORN types. Each ORN type expresses hundreds of transcription factors. Transcriptome-instructed genetic analyses revealed that (1) one broadly expressed transcription factor (Acj6) only regulates olfactory receptor expression in one ORN type and only wiring specificity in another type, (2) one type-restricted transcription factor (Forkhead) only regulates receptor expression, and (3) another type-restricted transcription factor (Unplugged) regulates both events. Thus, ORNs utilize diverse strategies and complex regulatory networks to coordinate their physiology and connectivity.
To perform most behaviors, animals must send commands from higher-order processing centers in the brain to premotor circuits that reside in ganglia distinct from the brain, such as the mammalian spinal cord or insect ventral nerve cord. How these circuits are functionally organized to generate the great diversity of animal behavior remains unclear. An important first step in unraveling the organization of premotor circuits is to identify their constituent cell types and create tools to monitor and manipulate these with high specificity to assess their functions. This is possible in the tractable ventral nerve cord of the fly. To generate such a toolkit, we used a combinatorial genetic technique (split-GAL4) to create 195 sparse transgenic driver lines targeting 196 individual cell types in the ventral nerve cord. These included wing and haltere motoneurons, modulatory neurons, and interneurons. Using a combination of behavioral, developmental, and anatomical analyses, we systematically characterized the cell types targeted in our collection. In addition, we identified correspondences between the cells in this collection and a recent connectomic data set of the ventral nerve cord. Taken together, the resources and results presented here form a powerful toolkit for future investigations of neuronal circuits and connectivity of premotor circuits while linking them to behavioral outputs.
Live-cell imaging of mRNA yields important insights into gene expression, but it has generally been limited to the labeling of one RNA species and has never been used to count single mRNAs over time in yeast. We demonstrate a two-color imaging system with single-molecule resolution using MS2 and PP7 RNA labeling. We use this methodology to measure intrinsic noise in mRNA levels and RNA polymerase II kinetics at a single gene.
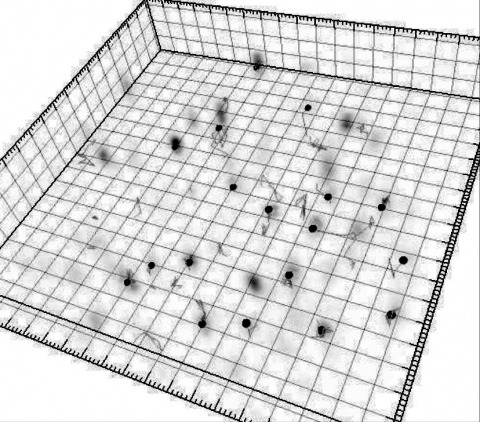
Within dendritic spines, actin is presumed to anchor receptors in the postsynaptic density and play numerous roles regulating synaptic transmission. However, the submicron dimensions of spines have hindered examination of actin dynamics within them and prevented live-cell discrimination of perisynaptic actin filaments. Using photoactivated localization microscopy, we measured movement of individual actin molecules within living spines. Velocity of single actin molecules along filaments, an index of filament polymerization rate, was highly heterogeneous within individual spines. Most strikingly, molecular velocity was elevated in discrete, well-separated foci occurring not principally at the spine tip, but in subdomains throughout the spine, including the neck. Whereas actin velocity on filaments at the synapse was substantially elevated, at the endocytic zone there was no enhanced polymerization activity. We conclude that actin subserves spatially diverse, independently regulated processes throughout spines. Perisynaptic actin forms a uniquely dynamic structure well suited for direct, active regulation of the synapse.
Commentary: A nice application of single particle tracking PALM (sptPALM), showing the flow of actin in the spines of live cultured neurons. Since 2008, the PALM in our lab has largely become a user facility, available to outside users as well as Janelians. Grad student Nick Frost in Tom Blanpied’s group at the U. of Maryland Med School visited on a number of occasions to use the PALM, with training and assistance from Hari.
The full promise of human genomics will be realized only when the genomes of thousands of individuals can be sequenced for comparative analysis. A reference sequence enables the use of short read length. We report an amplification-free method for determining the nucleotide sequence of more than 280,000 individual DNA molecules simultaneously. A DNA polymerase adds labeled nucleotides to surface-immobilized primer-template duplexes in stepwise fashion, and the asynchronous growth of individual DNA molecules was monitored by fluorescence imaging. Read lengths of >25 bases and equivalent phred software program quality scores approaching 30 were achieved. We used this method to sequence the M13 virus to an average depth of >150x and with 100% coverage; thus, we resequenced the M13 genome with high-sensitivity mutation detection. This demonstrates a strategy for high-throughput low-cost resequencing.
Enhancer-binding pluripotency regulators (Sox2 and Oct4) play a seminal role in embryonic stem (ES) cell-specific gene regulation. Here, we combine in vivo and in vitro single-molecule imaging, transcription factor (TF) mutagenesis, and ChIP-exo mapping to determine how TFs dynamically search for and assemble on their cognate DNA target sites. We find that enhanceosome assembly is hierarchically ordered with kinetically favored Sox2 engaging the target DNA first, followed by assisted binding of Oct4. Sox2/Oct4 follow a trial-and-error sampling mechanism involving 84-97 events of 3D diffusion (3.3-3.7 s) interspersed with brief nonspecific collisions (0.75-0.9 s) before acquiring and dwelling at specific target DNA (12.0-14.6 s). Sox2 employs a 3D diffusion-dominated search mode facilitated by 1D sliding along open DNA to efficiently locate targets. Our findings also reveal fundamental aspects of gene and developmental regulation by fine-tuning TF dynamics and influence of the epigenome on target search parameters.
Conserved ATP-dependent chromatin remodelers establish and maintain genome-wide chromatin architectures of regulatory DNA during cellular lifespan, but the temporal interactions between remodelers and chromatin targets have been obscure. We performed live-cell single-molecule tracking for RSC, SWI/SNF, CHD1, ISW1, ISW2, and INO80 remodeling complexes in budding yeast and detected hyperkinetic behaviors for chromatin-bound molecules that frequently transition to the free state for all complexes. Chromatin-bound remodelers display notably higher diffusion than nucleosomal histones, and strikingly fast dissociation kinetics with 4-7 s mean residence times. These enhanced dynamics require ATP binding or hydrolysis by the catalytic ATPase, uncovering an additional function to its established role in nucleosome remodeling. Kinetic simulations show that multiple remodelers can repeatedly occupy the same promoter region on a timescale of minutes, implicating an unending 'tug-of-war' that controls a temporally shifting window of accessibility for the transcription initiation machinery.