Filter
Associated Lab
- Ahrens Lab (1) Apply Ahrens Lab filter
- Aso Lab (1) Apply Aso Lab filter
- Baker Lab (3) Apply Baker Lab filter
- Betzig Lab (7) Apply Betzig Lab filter
- Bock Lab (2) Apply Bock Lab filter
- Branson Lab (1) Apply Branson Lab filter
- Cardona Lab (1) Apply Cardona Lab filter
- Cui Lab (3) Apply Cui Lab filter
- Dickson Lab (2) Apply Dickson Lab filter
- Druckmann Lab (1) Apply Druckmann Lab filter
- Dudman Lab (1) Apply Dudman Lab filter
- Eddy/Rivas Lab (5) Apply Eddy/Rivas Lab filter
- Fetter Lab (4) Apply Fetter Lab filter
- Fitzgerald Lab (1) Apply Fitzgerald Lab filter
- Gonen Lab (5) Apply Gonen Lab filter
- Grigorieff Lab (6) Apply Grigorieff Lab filter
- Heberlein Lab (12) Apply Heberlein Lab filter
- Hermundstad Lab (1) Apply Hermundstad Lab filter
- Hess Lab (2) Apply Hess Lab filter
- Jayaraman Lab (2) Apply Jayaraman Lab filter
- Ji Lab (2) Apply Ji Lab filter
- Kainmueller Lab (1) Apply Kainmueller Lab filter
- Keller Lab (4) Apply Keller Lab filter
- Lavis Lab (5) Apply Lavis Lab filter
- Lee (Albert) Lab (1) Apply Lee (Albert) Lab filter
- Leonardo Lab (2) Apply Leonardo Lab filter
- Lippincott-Schwartz Lab (18) Apply Lippincott-Schwartz Lab filter
- Liu (Zhe) Lab (1) Apply Liu (Zhe) Lab filter
- Looger Lab (7) Apply Looger Lab filter
- Magee Lab (1) Apply Magee Lab filter
- Menon Lab (3) Apply Menon Lab filter
- Murphy Lab (1) Apply Murphy Lab filter
- Pastalkova Lab (1) Apply Pastalkova Lab filter
- Pavlopoulos Lab (2) Apply Pavlopoulos Lab filter
- Reiser Lab (2) Apply Reiser Lab filter
- Riddiford Lab (1) Apply Riddiford Lab filter
- Romani Lab (1) Apply Romani Lab filter
- Rubin Lab (4) Apply Rubin Lab filter
- Satou Lab (3) Apply Satou Lab filter
- Scheffer Lab (2) Apply Scheffer Lab filter
- Schreiter Lab (3) Apply Schreiter Lab filter
- Sgro Lab (2) Apply Sgro Lab filter
- Simpson Lab (3) Apply Simpson Lab filter
- Singer Lab (10) Apply Singer Lab filter
- Spruston Lab (1) Apply Spruston Lab filter
- Stern Lab (4) Apply Stern Lab filter
- Sternson Lab (6) Apply Sternson Lab filter
- Svoboda Lab (7) Apply Svoboda Lab filter
- Tjian Lab (4) Apply Tjian Lab filter
- Truman Lab (1) Apply Truman Lab filter
- Turaga Lab (1) Apply Turaga Lab filter
- Turner Lab (2) Apply Turner Lab filter
- Zlatic Lab (1) Apply Zlatic Lab filter
- Zuker Lab (3) Apply Zuker Lab filter
Associated Project Team
Publication Date
- December 2011 (22) Apply December 2011 filter
- November 2011 (15) Apply November 2011 filter
- October 2011 (14) Apply October 2011 filter
- September 2011 (17) Apply September 2011 filter
- August 2011 (14) Apply August 2011 filter
- July 2011 (10) Apply July 2011 filter
- June 2011 (17) Apply June 2011 filter
- May 2011 (13) Apply May 2011 filter
- April 2011 (11) Apply April 2011 filter
- March 2011 (14) Apply March 2011 filter
- February 2011 (16) Apply February 2011 filter
- January 2011 (27) Apply January 2011 filter
- Remove 2011 filter 2011
Type of Publication
190 Publications
Showing 141-150 of 190 resultsSpinal muscular atrophy (SMA) results from reduced levels of the survival of motor neuron (SMN) protein, which has a well characterized function in spliceosomal small nuclear ribonucleoprotein assembly. Currently, it is not understood how deficiency of a housekeeping protein leads to the selective degeneration of spinal cord motor neurons. Numerous studies have shown that SMN is present in neuronal processes and has many interaction partners, including mRNA-binding proteins, suggesting a potential noncanonical role in axonal mRNA metabolism. In this study, we have established a novel technological approach using bimolecular fluorescence complementation (BiFC) and quantitative image analysis to characterize SMN-protein interactions in primary motor neurons. Consistent with biochemical studies on the SMN complex, BiFC analysis revealed that SMN dimerizes and interacts with Gemin2 in nuclear gems and axonal granules. In addition, using pull down assays, immunofluorescence, cell transfection, and BiFC, we characterized a novel interaction between SMN and the neuronal mRNA-binding protein HuD, which was dependent on the Tudor domain of SMN. A missense mutation in the SMN Tudor domain, which is known to cause SMA, impaired the interaction with HuD, but did not affect SMN axonal localization or self-association. Furthermore, time-lapse microscopy revealed SMN cotransport with HuD in live motor neurons. Importantly, SMN knockdown in primary motor neurons resulted in a specific reduction of both HuD protein and poly(A) mRNA levels in the axonal compartment. These findings reveal a noncanonical role for SMN whereby its interaction with mRNA-binding proteins may facilitate the localization of associated poly(A) mRNAs into axons.
Recent studies of several key developmental transitions have brought into question the long held view of the basal transcriptional apparatus as ubiquitous and invariant. In an effort to better understand the role of core promoter recognition and coactivator complex switching in cellular differentiation, we have examined changes in transcription factor IID (TFIID) and cofactor required for Sp1 activation/Mediator during mouse liver development. Here we show that the differentiation of fetal liver progenitors to adult hepatocytes involves a wholesale depletion of canonical cofactor required for Sp1 activation/Mediator and TFIID complexes at both the RNA and protein level, and that this alteration likely involves silencing of transcription factor promoters as well as protein degradation. It will be intriguing for future studies to determine if a novel and as yet unknown core promoter recognition complex takes the place of TFIID in adult hepatocytes and to uncover the mechanisms that down-regulate TFIID during this critical developmental transition.
Automated tracking of animal movement allows analyses that would not otherwise be possible by providing great quantities of data. The additional capability of tracking in real time–with minimal latency–opens up the experimental possibility of manipulating sensory feedback, thus allowing detailed explorations of the neural basis for control of behaviour. Here, we describe a system capable of tracking the three-dimensional position and body orientation of animals such as flies and birds. The system operates with less than 40 ms latency and can track multiple animals simultaneously. To achieve these results, a multi-target tracking algorithm was developed based on the extended Kalman filter and the nearest neighbour standard filter data association algorithm. In one implementation, an 11-camera system is capable of tracking three flies simultaneously at 60 frames per second using a gigabit network of nine standard Intel Pentium 4 and Core 2 Duo computers. This manuscript presents the rationale and details of the algorithms employed and shows three implementations of the system. An experiment was performed using the tracking system to measure the effect of visual contrast on the flight speed of Drosophila melanogaster. At low contrasts, speed is more variable and faster on average than at high contrasts. Thus, the system is already a useful tool to study the neurobiology and behaviour of freely flying animals. If combined with other techniques, such as ’virtual reality’-type computer graphics or genetic manipulation, the tracking system would offer a powerful new way to investigate the biology of flying animals.
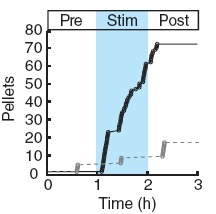
Two intermingled hypothalamic neuron populations specified by expression of agouti-related peptide (AGRP) or pro-opiomelanocortin (POMC) positively and negatively influence feeding behavior, respectively, possibly by reciprocally regulating downstream melanocortin receptors. However, the sufficiency of these neurons to control behavior and the relationship of their activity to the magnitude and dynamics of feeding are unknown. To measure this, we used channelrhodopsin-2 for cell type-specific photostimulation. Activation of only 800 AGRP neurons in mice evoked voracious feeding within minutes. The behavioral response increased with photoexcitable neuron number, photostimulation frequency and stimulus duration. Conversely, POMC neuron stimulation reduced food intake and body weight, which required melanocortin receptor signaling. However, AGRP neuron-mediated feeding was not dependent on suppressing this melanocortin pathway, indicating that AGRP neurons directly engage feeding circuits. Furthermore, feeding was evoked selectively over drinking without training or prior photostimulus exposure, which suggests that AGRP neurons serve a dedicated role coordinating this complex behavior.
We developed a multicolor neuron labeling technique in Drosophila melanogaster that combines the power to specifically target different neural populations with the label diversity provided by stochastic color choice. This adaptation of vertebrate Brainbow uses recombination to select one of three epitope-tagged proteins detectable by immunofluorescence. Two copies of this construct yield six bright, separable colors. We used Drosophila Brainbow to study the innervation patterns of multiple antennal lobe projection neuron lineages in the same preparation and to observe the relative trajectories of individual aminergic neurons. Nerve bundles, and even individual neurites hundreds of micrometers long, can be followed with definitive color labeling. We traced motor neurons in the subesophageal ganglion and correlated them to neuromuscular junctions to identify their specific proboscis muscle targets. The ability to independently visualize multiple lineage or neuron projections in the same preparation greatly advances the goal of mapping how neurons connect into circuits.
The pattern of spikes recorded from place cells in the rodent hippocampus is strongly modulated by both the spatial location in the environment and the theta rhythm. The phases of the spikes in the theta cycle advance during movement through the place field. Recently intracellular recordings from hippocampal neurons (Harvey, Collman, Dombeck, & Tank, 2009 ) showed an increase in the amplitude of membrane potential oscillations inside the place field, which was interpreted as evidence that an intracellular mechanism caused phase precession. Here we show that an existing network model of the hippocampus (Tsodyks, Skaggs, Sejnowski, & McNaughton, 1996 ) can equally reproduce this and other aspects of the intracellular recordings, which suggests that new experiments are needed to distinguish the contributions of intracellular and network mechanisms to phase precession.
Most neurons of the central complex belong to 10 secondary (larvally produced) lineages. In the late larva, undifferentiated axon tracts of these lineages form a primordium in which all of the compartments of the central complex can be recognized as discrete entities. Four posterior lineages (DPMm1, DPMpm1, DPMpm2, and CM4) generate the classes of small-field neurons that interconnect the protocerebral bridge, fan-shaped body, noduli, and ellipsoid body. Three lineages located in the anterior brain, DALv2, BAmv1, and DALcl2, form the large-field neurons of the ellipsoid body and fan-shaped body, respectively. These lineages provide an input channel from the optic tubercle and connect the central complex with adjacent anterior brain compartments. Three lineages in the posterior cortex, CM3, CP2, and DPMpl2, connect the posterior brain neuropil with specific layers of the fan-shaped body. Even though all of the compartments of the central complex are prefigured in the late larval brain by the axon tracts of the above-mentioned lineages, the neuropil differentiates during the first 2 days of the pupal period when terminal branches and synapses of secondary neurons are formed. During this phase the initially straight horizontal layers of the central complex bend in the frontal plane, which produces the characteristic shape of the fan-shaped and ellipsoid body. Our analysis provides a comprehensive picture of the lineages that form the central complex, and will facilitate future studies that address the structure or function of the central complex at the single cell level.
Manipulation of hematopoietic stem/progenitor cells (HSPCs) ex vivo is of clinical importance for stem cell expansion and gene therapy applications. However, most cultured HSPCs are actively cycling, and show a homing and engraftment defect compared with the predominantly quiescent noncultured HSPCs. We previously showed that HSPCs make contact with osteoblasts in vitro via a polarized membrane domain enriched in adhesion molecules such as tetraspanins. Here we show that increased cell cycling during ex vivo culture of HSPCs resulted in disruption of this membrane domain, as evidenced by disruption of polarity of the tetraspanin CD82. Chemical disruption or antibody-mediated blocking of CD82 on noncultured HSPCs resulted in decreased stromal cell adhesion, homing, and engraftment in nonobese diabetic/severe combined immunodeficiency IL-2γ(null) (NSG) mice compared with HSPCs with an intact domain. Most leukemic blasts were actively cycling and correspondingly displayed a loss of domain polarity and decreased homing in NSG mice compared with normal HSPCs. We conclude that quiescent cells, unlike actively cycling cells, display a polarized membrane domain enriched in tetraspanins that mediates homing and engraftment, providing a mechanistic explanation for the homing/engraftment defect of cycling cells and a potential new therapeutic target to enhance engraftment.
Hippocampal neurons can display reliable and long-lasting sequences of transient firing patterns, even in the absence of changing external stimuli. We suggest that time-keeping is an important function of these sequences, and propose a network mechanism for their generation. We show that sequences of neuronal assemblies recorded from rat hippocampal CA1 pyramidal cells can reliably predict elapsed time (15-20 s) during wheel running with a precision of 0.5 s. In addition, we demonstrate the generation of multiple reliable, long-lasting sequences in a recurrent network model. These sequences are generated in the presence of noisy, unstructured inputs to the network, mimicking stationary sensory input. Identical initial conditions generate similar sequences, whereas different initial conditions give rise to distinct sequences. The key ingredients responsible for sequence generation in the model are threshold-adaptation and a Mexican-hat-like pattern of connectivity among pyramidal cells. This pattern may arise from recurrent systems such as the hippocampal CA3 region or the entorhinal cortex. We hypothesize that mechanisms that evolved for spatial navigation also support tracking of elapsed time in behaviorally relevant contexts.
Decoding the wiring diagram of the retina requires simultaneous observation of activity in identified neuron populations. Available recording methods are limited in their scope: electrodes can access only a small fraction of neurons at once, whereas synthetic fluorescent indicator dyes label tissue indiscriminately. Here, we describe a method for studying retinal circuitry at cellular and subcellular levels combining two-photon microscopy and a genetically encoded calcium indicator. Using specific viral and promoter constructs to drive expression of GCaMP3, we labeled all five major neuron classes in the adult mouse retina. Stimulus-evoked GCaMP3 responses as imaged by two-photon microscopy permitted functional cell type annotation. Fluorescence responses were similar to those measured with the small molecule dye OGB-1. Fluorescence intensity correlated linearly with spike rates >10 spikes/s, and a significant change in fluorescence always reflected a significant change in spike firing rate. GCaMP3 expression had no apparent effect on neuronal function. Imaging at subcellular resolution showed compartment-specific calcium dynamics in multiple identified cell types.