Filter
Associated Lab
- Aguilera Castrejon Lab (19) Apply Aguilera Castrejon Lab filter
- Ahrens Lab (73) Apply Ahrens Lab filter
- Aso Lab (42) Apply Aso Lab filter
- Baker Lab (38) Apply Baker Lab filter
- Betzig Lab (116) Apply Betzig Lab filter
- Beyene Lab (15) Apply Beyene Lab filter
- Bock Lab (17) Apply Bock Lab filter
- Branson Lab (55) Apply Branson Lab filter
- Card Lab (43) Apply Card Lab filter
- Cardona Lab (64) Apply Cardona Lab filter
- Chklovskii Lab (13) Apply Chklovskii Lab filter
- Clapham Lab (16) Apply Clapham Lab filter
- Cui Lab (19) Apply Cui Lab filter
- Darshan Lab (12) Apply Darshan Lab filter
- Dennis Lab (3) Apply Dennis Lab filter
- Dickson Lab (46) Apply Dickson Lab filter
- Druckmann Lab (25) Apply Druckmann Lab filter
- Dudman Lab (56) Apply Dudman Lab filter
- Eddy/Rivas Lab (30) Apply Eddy/Rivas Lab filter
- Egnor Lab (11) Apply Egnor Lab filter
- Espinosa Medina Lab (23) Apply Espinosa Medina Lab filter
- Feliciano Lab (12) Apply Feliciano Lab filter
- Fetter Lab (41) Apply Fetter Lab filter
- FIB-SEM Technology (1) Apply FIB-SEM Technology filter
- Fitzgerald Lab (30) Apply Fitzgerald Lab filter
- Freeman Lab (15) Apply Freeman Lab filter
- Funke Lab (46) Apply Funke Lab filter
- Gonen Lab (91) Apply Gonen Lab filter
- Grigorieff Lab (62) Apply Grigorieff Lab filter
- Harris Lab (65) Apply Harris Lab filter
- Heberlein Lab (94) Apply Heberlein Lab filter
- Hermundstad Lab (30) Apply Hermundstad Lab filter
- Hess Lab (80) Apply Hess Lab filter
- Ilanges Lab (4) Apply Ilanges Lab filter
- Jayaraman Lab (48) Apply Jayaraman Lab filter
- Ji Lab (33) Apply Ji Lab filter
- Johnson Lab (6) Apply Johnson Lab filter
- Kainmueller Lab (19) Apply Kainmueller Lab filter
- Karpova Lab (14) Apply Karpova Lab filter
- Keleman Lab (13) Apply Keleman Lab filter
- Keller Lab (76) Apply Keller Lab filter
- Koay Lab (19) Apply Koay Lab filter
- Lavis Lab (158) Apply Lavis Lab filter
- Lee (Albert) Lab (34) Apply Lee (Albert) Lab filter
- Leonardo Lab (23) Apply Leonardo Lab filter
- Li Lab (32) Apply Li Lab filter
- Lippincott-Schwartz Lab (180) Apply Lippincott-Schwartz Lab filter
- Liu (Yin) Lab (8) Apply Liu (Yin) Lab filter
- Liu (Zhe) Lab (65) Apply Liu (Zhe) Lab filter
- Looger Lab (138) Apply Looger Lab filter
- Magee Lab (49) Apply Magee Lab filter
- Menon Lab (18) Apply Menon Lab filter
- Murphy Lab (13) Apply Murphy Lab filter
- O'Shea Lab (8) Apply O'Shea Lab filter
- Otopalik Lab (13) Apply Otopalik Lab filter
- Pachitariu Lab (52) Apply Pachitariu Lab filter
- Pastalkova Lab (19) Apply Pastalkova Lab filter
- Pavlopoulos Lab (19) Apply Pavlopoulos Lab filter
- Pedram Lab (15) Apply Pedram Lab filter
- Podgorski Lab (16) Apply Podgorski Lab filter
- Reiser Lab (55) Apply Reiser Lab filter
- Riddiford Lab (44) Apply Riddiford Lab filter
- Romani Lab (51) Apply Romani Lab filter
- Rubin Lab (149) Apply Rubin Lab filter
- Saalfeld Lab (64) Apply Saalfeld Lab filter
- Satou Lab (18) Apply Satou Lab filter
- Scheffer Lab (38) Apply Scheffer Lab filter
- Schreiter Lab (70) Apply Schreiter Lab filter
- Schulze Lab (1) Apply Schulze Lab filter
- Sgro Lab (23) Apply Sgro Lab filter
- Shroff Lab (31) Apply Shroff Lab filter
- Simpson Lab (23) Apply Simpson Lab filter
- Singer Lab (80) Apply Singer Lab filter
- Spruston Lab (98) Apply Spruston Lab filter
- Stern Lab (160) Apply Stern Lab filter
- Sternson Lab (54) Apply Sternson Lab filter
- Stringer Lab (41) Apply Stringer Lab filter
- Svoboda Lab (136) Apply Svoboda Lab filter
- Tebo Lab (35) Apply Tebo Lab filter
- Tervo Lab (9) Apply Tervo Lab filter
- Tillberg Lab (22) Apply Tillberg Lab filter
- Tjian Lab (64) Apply Tjian Lab filter
- Truman Lab (88) Apply Truman Lab filter
- Turaga Lab (53) Apply Turaga Lab filter
- Turner Lab (38) Apply Turner Lab filter
- Vale Lab (8) Apply Vale Lab filter
- Voigts Lab (4) Apply Voigts Lab filter
- Wang (Meng) Lab (29) Apply Wang (Meng) Lab filter
- Wang (Shaohe) Lab (25) Apply Wang (Shaohe) Lab filter
- Wong-Campos Lab (1) Apply Wong-Campos Lab filter
- Wu Lab (9) Apply Wu Lab filter
- Zlatic Lab (28) Apply Zlatic Lab filter
- Zuker Lab (25) Apply Zuker Lab filter
Associated Project Team
- CellMap (13) Apply CellMap filter
- COSEM (3) Apply COSEM filter
- FIB-SEM Technology (5) Apply FIB-SEM Technology filter
- Fly Descending Interneuron (12) Apply Fly Descending Interneuron filter
- Fly Functional Connectome (14) Apply Fly Functional Connectome filter
- Fly Olympiad (5) Apply Fly Olympiad filter
- FlyEM (56) Apply FlyEM filter
- FlyLight (50) Apply FlyLight filter
- GENIE (47) Apply GENIE filter
- Integrative Imaging (9) Apply Integrative Imaging filter
- Larval Olympiad (2) Apply Larval Olympiad filter
- MouseLight (18) Apply MouseLight filter
- NeuroSeq (1) Apply NeuroSeq filter
- ThalamoSeq (1) Apply ThalamoSeq filter
- Tool Translation Team (T3) (29) Apply Tool Translation Team (T3) filter
- Transcription Imaging (49) Apply Transcription Imaging filter
Publication Date
- 2026 (70) Apply 2026 filter
- 2025 (222) Apply 2025 filter
- 2024 (210) Apply 2024 filter
- 2023 (158) Apply 2023 filter
- 2022 (192) Apply 2022 filter
- 2021 (194) Apply 2021 filter
- 2020 (196) Apply 2020 filter
- 2019 (202) Apply 2019 filter
- 2018 (232) Apply 2018 filter
- 2017 (217) Apply 2017 filter
- 2016 (209) Apply 2016 filter
- 2015 (252) Apply 2015 filter
- 2014 (236) Apply 2014 filter
- 2013 (194) Apply 2013 filter
- 2012 (190) Apply 2012 filter
- 2011 (190) Apply 2011 filter
- 2010 (161) Apply 2010 filter
- 2009 (158) Apply 2009 filter
- 2008 (140) Apply 2008 filter
- 2007 (106) Apply 2007 filter
- 2006 (92) Apply 2006 filter
- 2005 (67) Apply 2005 filter
- 2004 (57) Apply 2004 filter
- 2003 (58) Apply 2003 filter
- 2002 (39) Apply 2002 filter
- 2001 (28) Apply 2001 filter
- 2000 (29) Apply 2000 filter
- 1999 (14) Apply 1999 filter
- 1998 (18) Apply 1998 filter
- 1997 (16) Apply 1997 filter
- 1996 (10) Apply 1996 filter
- 1995 (18) Apply 1995 filter
- 1994 (12) Apply 1994 filter
- 1993 (10) Apply 1993 filter
- 1992 (6) Apply 1992 filter
- 1991 (11) Apply 1991 filter
- 1990 (11) Apply 1990 filter
- 1989 (6) Apply 1989 filter
- 1988 (1) Apply 1988 filter
- 1987 (7) Apply 1987 filter
- 1986 (4) Apply 1986 filter
- 1985 (5) Apply 1985 filter
- 1984 (2) Apply 1984 filter
- 1983 (2) Apply 1983 filter
- 1982 (3) Apply 1982 filter
- 1981 (3) Apply 1981 filter
- 1980 (1) Apply 1980 filter
- 1979 (1) Apply 1979 filter
- 1976 (2) Apply 1976 filter
- 1973 (1) Apply 1973 filter
- 1970 (1) Apply 1970 filter
- 1967 (1) Apply 1967 filter
Type of Publication
4265 Publications
Showing 2611-2620 of 4265 resultsNear-field scanning optical microscopy (NSOM) has been used to image and record domains in thin-film magneto-optic (MO) materials. In the imaging mode, resolution of 30-50 nm has been consistently obtained, whereas in the recording mode, domains down to -60 nm have been written reproducibly. Data densities of -45 Gbits/in.’ have been achieved, well in excess of current magnetic or MO technologies. A brief analysis of speed and other issues indicates that the technique may represent a viable alternative to density data storage needs.
Commentary: The first demonstration of optical recording and playback beyond the diffraction limit, using magneto-optic multilayer films and polarization contrast near-field microscopy. Bits as small as 60 nm were recorded – beyond estimates at the time of the superparamagnetic limit to bit stability. Bit densities of 45 Gbits/in2 were also achieved, well in excess of optical or magnetic recording technologies of the era. In the years following this work, massive resources were spent on the commercialization of near-field data storage, largely for naught.
The near-field optical interaction between a sharp probe and a sample of interest can be exploited to image, spectroscopically probe, or modify surfaces at a resolution (down to approximately 12 nm) inaccessible by traditional far-field techniques. Many of the attractive features of conventional optics are retained, including noninvasiveness, reliability, and low cost. In addition, most optical contrast mechanisms can be extended to the near-field regime, resulting in a technique of considerable versatility. This versatility is demonstrated by several examples, such as the imaging of nanometric-scale features in mammalian tissue sections and the creation of ultrasmall, magneto-optic domains having implications for highdensity data storage. Although the technique may find uses in many diverse fields, two of the most exciting possibilities are localized optical spectroscopy of semiconductors and the fluorescence imaging of living cells.
Commentary: An overview of our work in near-field optics at the time, after our invention of the adiabatically tapered fiber probe and shear force feedback (see below) led to the first practical near-field scanning optical microscope. In this work, superresolution imaging via absorption, reflectivity, fluorescence, spectroscopy, polarization, and refractive index contrast were all demonstrated. Unlike all far-field superresolution fluorescence methods that were to appear a decade later, near-field microscopy remains the only superresolution technique capable of taking advantage of the full panoply of optical contrast mechanisms.
Luminescent centers with sharp (<0.07 millielectron volt), spectrally distinct emission lines were imaged in a GaAs/AIGaAs quantum well by means of low-temperature near-field scanning optical microscopy. Temperature, magnetic field, and linewidth measurements establish that these centers arise from excitons laterally localized at interface fluctuations. For sufficiently narrow wells, virtually all emission originates from such centers. Near-field microscopy/spectroscopy provides a means to access energies and homogeneous line widths for the individual eigenstates of these centers, and thus opens a rich area of physics involving quantum resolved systems.
Luminescent centers with sharp (<0.07 millielectron volt), spectrally distinct emission lines were imaged in a GaAs/AIGaAs quantum well by means of low-temperature near-field scanning optical microscopy. Temperature, magnetic field, and linewidth measurements establish that these centers arise from excitons laterally localized at interface fluctuations. For sufficiently narrow wells, virtually all emission originates from such centers. Near-field microscopy/spectroscopy provides a means to access energies and homogeneous line widths for the individual eigenstates of these centers, and thus opens a rich area of physics involving quantum resolved systems.
Commentary: Harald Hess and I joined forces, combining my near-field optical technology with his cryogenic scanned probe microscope to produce the first paper on high resolution spectroscopy beyond the diffraction limit. We discovered that the broad luminescence spectrum traditionally observed from quantum well heterostructures reflects a resolution-limited ensemble average of emission from numerous discrete sites of exciton recombination occurring at atomic-scale corrugations in the confining interfaces. With the combination of high spatial resolution from near-field excitation and high spectral resolution from cryogenic operation, we were able to isolate these emission sites in a multidimensional space of xy position and wavelength, even though their density was too great to isolate them on the basis of spatial resolution alone. This insight was very influential in the genesis of the concept (see above) that would eventually lead to far-field superresolution by PALM.
Dopamine neuromodulation of neural synapses is a process implicated in a number of critical brain functions and diseases. Development of protocols to visualize this dynamic neurochemical process is essential to understanding how dopamine modulates brain function. We have developed a non-genetically encoded, near-IR (nIR) catecholamine nanosensor (nIRCat) capable of identifying ~2-µm dopamine release hotspots in dorsal striatal brain slices. nIRCat is readily synthesized through sonication of single walled carbon nanotubes with DNA oligos, can be readily introduced into both genetically tractable and intractable organisms and is compatible with a number of dopamine receptor agonists and antagonists. Here we describe the synthesis, characterization and implementation of nIRCat in acute mouse brain slices. We demonstrate how nIRCat can be used to image electrically or optogenetically stimulated dopamine release, and how these procedures can be leveraged to study the effects of dopamine receptor pharmacology. In addition, we provide suggestions for building or adapting wide-field microscopy to be compatible with nIRCat nIR fluorescence imaging. We discuss strategies for analyzing nIR video data to identify dopamine release hotspots and quantify their kinetics. This protocol can be adapted and implemented for imaging other neuromodulators by using probes of this class and can be used in a broad range of species without genetic manipulation. The synthesis and characterization protocols for nIRCat take ~5 h, and the preparation and fluorescence imaging of live brain slices by using nIRCats require ~6 h.
BACKGROUND: The use of genetically-encoded fluorescent reporters is essential for the identification and observation of cells that express transgenic modulatory proteins. Near-infrared (NIR) fluorescent proteins have superior light penetration through biological tissue, but are not yet widely adopted. NEW METHOD: Using the near-infrared fluorescent protein, iRFP713, improves the imaging resolution in thick tissue sections or the intact brain due to the reduced light-scattering at the longer, NIR wavelengths used to image the protein. Additionally, iRFP713 can be used to identify transgenic cells without photobleaching other fluorescent reporters or affecting opsin function. We have generated a set of adeno-associated vectors in which iRFP713 has been fused to optogenetic channels, and can be expressed constitutively or Cre-dependently. RESULTS: iRFP713 is detectable when expressed in neurons both in vitro and in vivo without exogenously supplied chromophore biliverdin. Neuronally-expressed iRFP713 has similar properties to GFP-like fluorescent proteins, including the ability to be translationally fused to channelrhodopsin or halorhodopsin, however, it shows superior photostability compared to EYFP. Furthermore, electrophysiological recordings from iRFP713-labeled cells compared to cells labeled with mCherry suggest that iRFP713 cells are healthier and therefore more stable and reliable in an ex vivo preparation. Lastly, we have generated a transgenic rat that expresses iRFP713 in a Cre-dependent manner. CONCLUSIONS: Overall, we have demonstrated that iRFP713 can be used as a reporter in neurons without the use of exogenous biliverdin, with minimal impact on viability and function thereby making it feasible to extend the capabilities for imaging genetically-tagged neurons in slices and in vivo.
Oxytocin plays a critical role in regulating social behaviors, yet our understanding of its function in both neurological health and disease remains incomplete. Real-time oxytocin imaging probes with spatiotemporal resolution relevant to its endogenous signaling are required to fully elucidate oxytocin's role in the brain. Herein, we describe a near-infrared oxytocin nanosensor (nIROXT), a synthetic probe capable of imaging oxytocin in the brain without interference from its structural analogue, vasopressin. nIROXT leverages the inherent tissue-transparent fluorescence of single-walled carbon nanotubes (SWCNT) and the molecular recognition capacity of an oxytocin receptor peptide fragment to selectively and reversibly image oxytocin. We employ these nanosensors to monitor electrically stimulated oxytocin release in brain tissue, revealing oxytocin release sites with a median size of 3 µm in the paraventricular nucleus of C57BL/6 mice, which putatively represents the spatial diffusion of oxytocin from its point of release. These data demonstrate that covalent SWCNT constructs, such as nIROXT, are powerful optical tools that can be leveraged to measure neuropeptide release in brain tissue.
Imaging approaches based on single molecule localization break the diffraction barrier of conventional fluorescence microscopy, allowing for bioimaging with nanometer resolution. It remains a challenge, however, to precisely localize photon-limited single molecules in 3D. We have developed a new localization-based imaging technique achieving almost isotropic subdiffraction resolution in 3D. A tilted mirror is used to generate a side view in addition to the front view of activated single emitters, allowing their 3D localization to be precisely determined for superresolution imaging. Because both front and side views are in focus, this method is able to efficiently collect emitted photons. The technique is simple to implement on a commercial fluorescence microscope, and especially suitable for biological samples with photon-limited chromophores such as endogenously expressed photoactivatable fluorescent proteins. Moreover, this method is relatively resistant to optical aberration, as it requires only centroid determination for localization analysis. Here we demonstrate the application of this method to 3D imaging of bacterial protein distribution and neuron dendritic morphology with subdiffraction resolution.
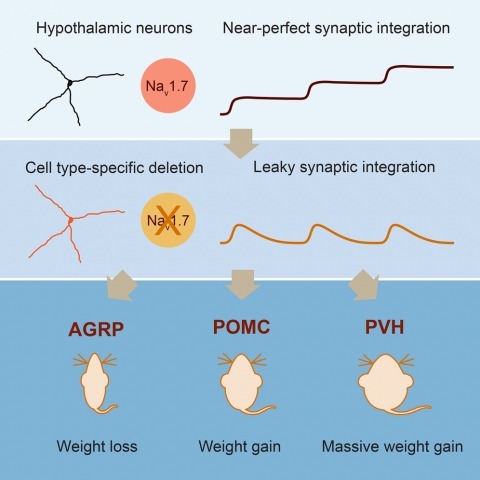
Neurons are well suited for computations on millisecond timescales, but some neuronal circuits set behavioral states over long time periods, such as those involved in energy homeostasis. We found that multiple types of hypothalamic neurons, including those that oppositely regulate body weight, are specialized as near-perfect synaptic integrators that summate inputs over extended timescales. Excitatory postsynaptic potentials (EPSPs) are greatly prolonged, outlasting the neuronal membrane time-constant up to 10-fold. This is due to the voltage-gated sodium channel Nav1.7 (Scn9a), previously associated with pain-sensation but not synaptic integration. Scn9a deletion in AGRP, POMC, or paraventricular hypothalamic neurons reduced EPSP duration, synaptic integration, and altered body weight in mice. In vivo whole-cell recordings in the hypothalamus confirmed near-perfect synaptic integration. These experiments show that integration of synaptic inputs over time by Nav1.7 is critical for body weight regulation and reveal a mechanism for synaptic control of circuits regulating long term homeostatic functions.
Presynaptic, electron-dense, cytoplasmic protrusions such as the T-bar (Drosophila) or ribbon (vertebrates) are believed to facilitate vesicle movement to the active zone (AZ) of synapses throughout the nervous system. The molecular composition of these structures including the T-bar and ribbon are largely unknown, as are the mechanisms that specify their synapse-specific assembly and distribution. In a large-scale, forward genetic screen, we have identified a mutation termed air traffic controller (atc) that causes T-bar-like protein aggregates to form abnormally in motoneuron axons. This mutation disrupts a gene that encodes for a serine-arginine protein kinase (SRPK79D). This mutant phenotype is specific to SRPK79D and is not secondary to impaired kinesin-dependent axonal transport. The srpk79D gene is neuronally expressed, and transgenic rescue experiments are consistent with SRPK79D kinase activity being necessary in neurons. The SRPK79D protein colocalizes with the T-bar-associated protein Bruchpilot (Brp) in both the axon and synapse. We propose that SRPK79D is a novel T-bar-associated protein kinase that represses T-bar assembly in peripheral axons, and that SRPK79D-dependent repression must be relieved to facilitate site-specific AZ assembly. Consistent with this model, overexpression of SRPK79D disrupts AZ-specific Brp organization and significantly impairs presynaptic neurotransmitter release. These data identify a novel AZ-associated protein kinase and reveal a new mechanism of negative regulation involved in AZ assembly. This mechanism could contribute to the speed and specificity with which AZs are assembled throughout the nervous system.