Researchers combine high-resolution microscopy and high-performance computing for a new way to map – and manipulate – brain cells during experiments
Neural circuits spanning far-flung regions of the brain are now easier to decipher – and manipulate. A new tool that merges light-sheet microscopy, high-performance computing, and optogenetics lets researchers turn on or off neurons while observing the entire brain’s response in behaving zebrafish.
The work helps solves a key problem in neuroscience: how to identify the complete set of neurons that underlie any given animal behavior.
“This technique makes it possible to analyze brain activity on the fly, as an experiment is running, for the whole brain,” says Misha Ahrens, a group leader at the Howard Hughes Medical Institute’s Janelia Research Campus. In collaboration with Janelia group leader Philipp Keller and his team, Ahrens’s team reports their new technique on November 30, 2018, in the journal Nature Methods.
Neurons scattered across multiple brain areas often control actions like swimming or talking. Yet conventional tools often limit researchers to studying isolated brain regions that they’ve already identified as important for certain behaviors. This means brain cells or circuits that contribute to behaviors can be overlooked.
The collaborative Janelia team’s technique is unique because it combines high-resolution neuron imaging, manipulation, and computational analysis over an animal’s entire brain. That lets researchers observe the neurons activated when an animal performs a behavior – and then selectively turn them off.
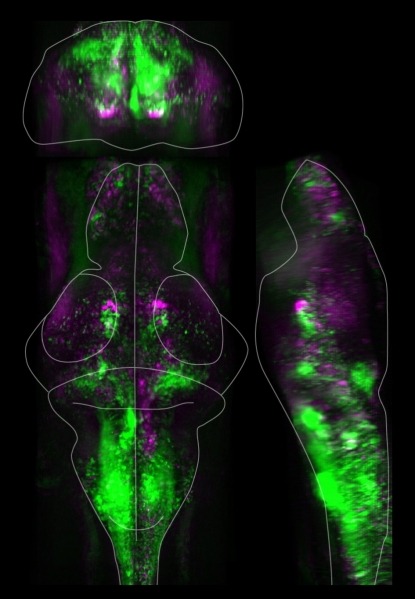 When paired with high performance computing, three-dimensional ‘brain-activity maps’ of larval zebrafish allow researchers to manipulate neurons while monitoring changes in animal behavior and brain activity. Green regions indicate a decrease in neural activity after turning off brain cells in discrete regions (white ovals).First, the researchers image the brain with a light-sheet microscope, which uses very thin laser beams to make three-dimensional measurements of brain activity. The team pairs that imaging setup with a cluster of linked computers, which lets them rapidly analyze neuronal activity data and create “brain activity maps.”
When paired with high performance computing, three-dimensional ‘brain-activity maps’ of larval zebrafish allow researchers to manipulate neurons while monitoring changes in animal behavior and brain activity. Green regions indicate a decrease in neural activity after turning off brain cells in discrete regions (white ovals).First, the researchers image the brain with a light-sheet microscope, which uses very thin laser beams to make three-dimensional measurements of brain activity. The team pairs that imaging setup with a cluster of linked computers, which lets them rapidly analyze neuronal activity data and create “brain activity maps.”
The next step is using those maps to select which neurons to test. Ahrens’s and Keller’s teams looked at the neurons underlying a swimming response in zebrafish, and then used optogenetic techniques to switch those neurons on or off.
For nearly any brain region that the researchers manipulated, the activity maps they generated also showed responses in faraway neurons, says Keller. These responses in distant neurons would be difficult to record – or manipulate – using conventional techniques.
This integration of observation, manipulation, and experimentation in the whole brain of an individual animal provides new research opportunities, Ahrens says. For instance, researchers can now generate hypotheses about neurons’ roles and test those hypotheses within the same experiment, not only in zebrafish but also in other model organisms. The work will make it easier to determine each part widely distributed neurons play in specific animal behaviors.
###
Citation
Nikita Vladimirov, Chen Wang, Burkhard Hoeckendorf, Avinash Pujala, Masashi Tanimoto, Yu Mu, Chao-Tsung Yang, Jason D. Wittenbach, Jeremy Freeman, Stephan Preibisch, Minoru Koyama, Philipp J. Keller, and Misha B Ahrens. “Brain-wide circuit interrogation at the cellular level guided by online analysis of neuronal function.” Nature Methods. Published online November 30, 2018. doi: 10.1038/s41592-018-0221-x
Media Contacts
Meghan Rosen
301-215-8859
rosenm2@hhmi.org




