New research finds a protein of the virus is not the ion channel previously thought, but it could play a role in making SARS-CoV-2 more virulent.
When Alexandria Miller arrived at Janelia as a postdoc in the Clapham Lab in September 2018, she was prepared to start a project examining ion channels in organelles.
But as the COVID-19 pandemic started to intensify, Miller began reading more about SARS-CoV-2 and its proteins.
One of these proteins, Orf3a, caught her eye. Orf3a had previously been reported to be a viroporin, a type of virus protein that forms ion channels. These channels enable chemical signals to pass between host cells and help a virus to proliferate by defeating the host cells’ defenses. But no one had fully deciphered Orf3a’s structure to better understand the protein’s role.
Instead of pursuing her original project, Miller pivoted. She decided to investigate Orf3a, drawing on her training as a biochemist and structural biologist and the Clapham Lab’s expertise in ion channels.
Two and a half years later, Miller and a team of collaborators report in a new preprint on bioRxiv that Orf3a in fact is not a viroporin, as previously thought, but it could play a different role in making SARS-CoV-2 more virulent.
Building knowledge
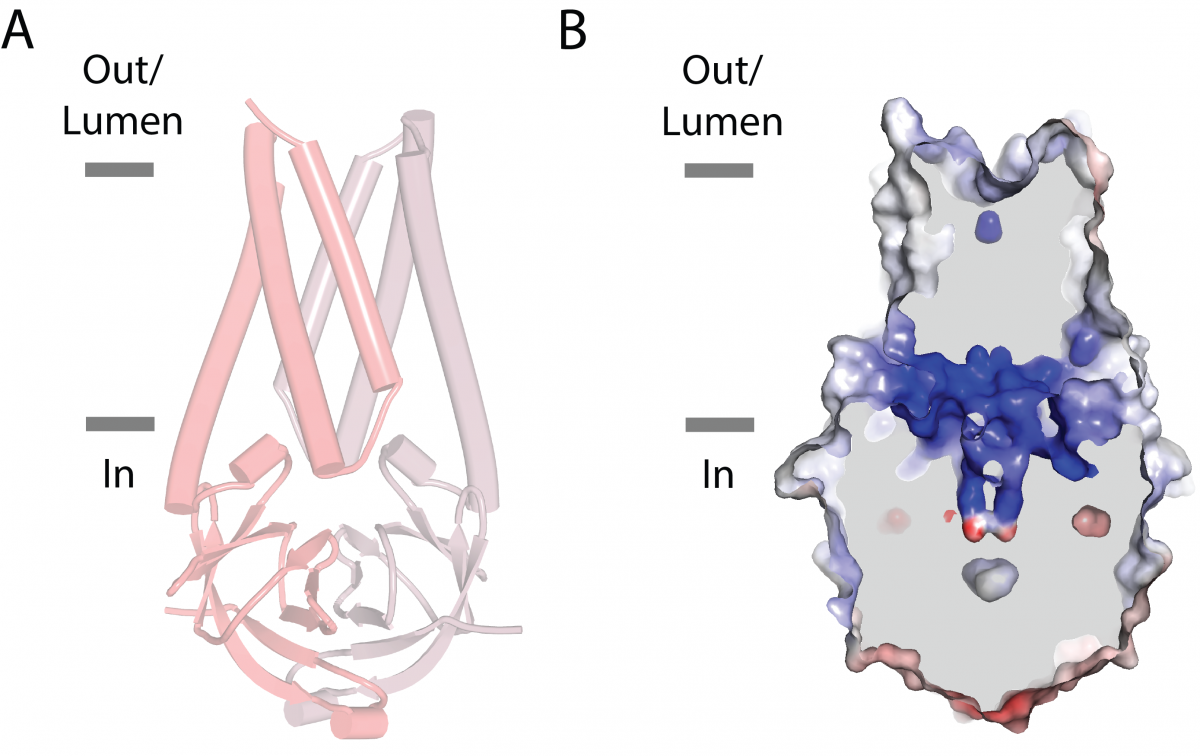
To look for signs that Orf3a could be an ion channel, the researchers combined information about the protein’s structure obtained from high-resolution electron microscopy with functional data acquired from electrophysiology. But neither the structural nor the functional data showed any indication that the protein is an ion channel, leading the researchers to their conclusion that Orf3a is not a viroporin.
“It turned out to be disappointing from one angle, but it is good to know on the other side,” says Senior Group Leader David Clapham. “All kind of effort gets made, when something is misidentified, to try to make something to block it, and that would have been a waste of time. It is good to clarify what it is so people can focus on understanding it in different ways.”
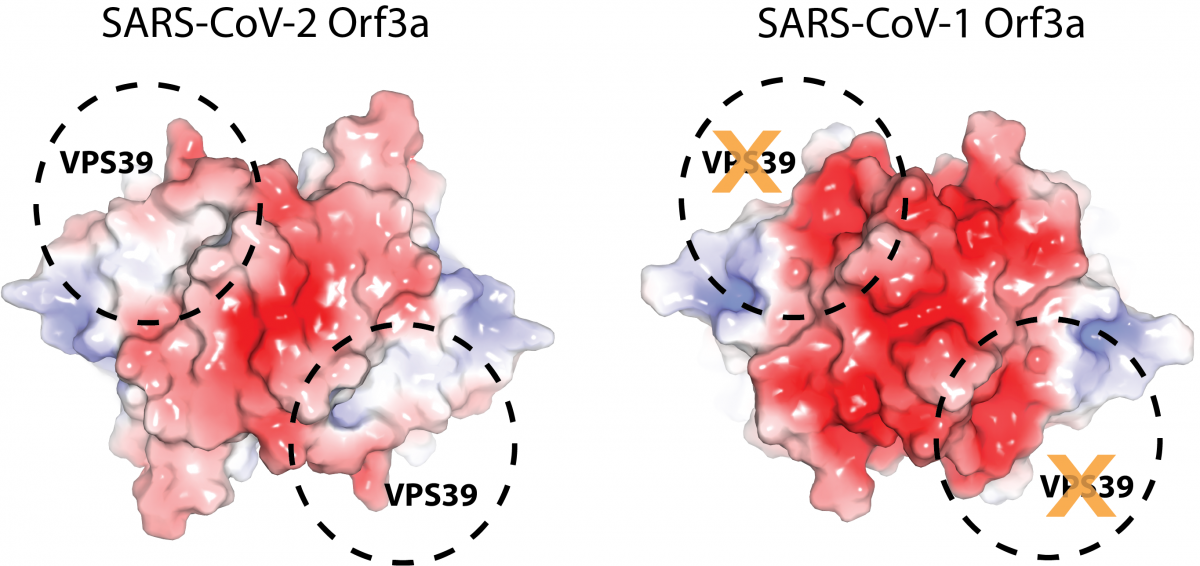
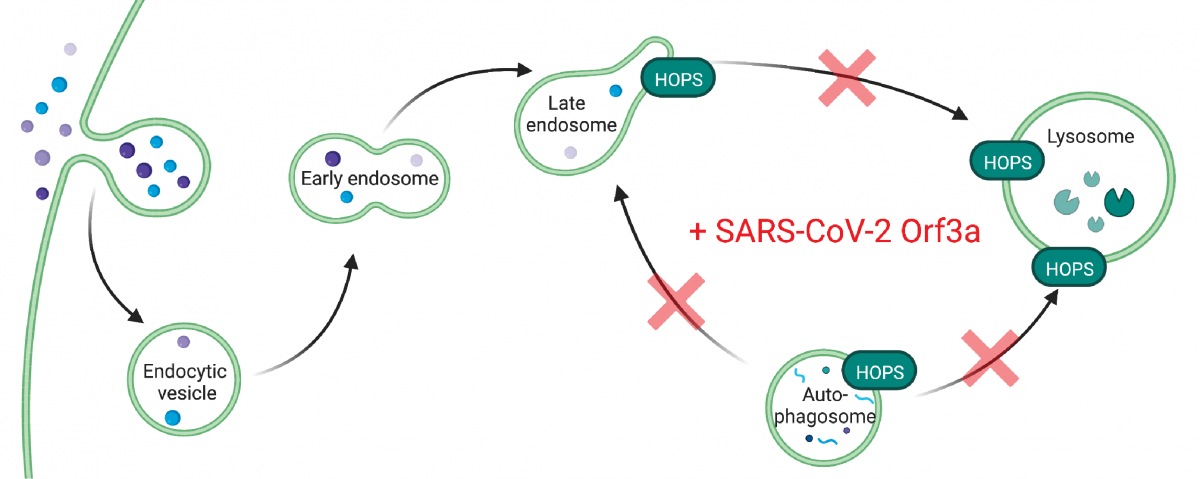
The new study also indicates Orf3a may have a different role than expected in helping SARS-CoV-2 evade the immune system. Using additional structural information, the researchers found that, in SARS-CoV-2, a loop on the Orf3a protein that is not present in its predecessor, SARS-CoV-1, interacts with a protein in the host cell. This interaction could disrupt a process the host cell uses to get rid of the virus. It could also play a role in blocking the degradation of foreign material in the cell.
Both hypotheses suggest Orf3a could enhance SARS-CoV-2’s ability to take over a healthy cell and proliferate, hinting at why the virus is more virulent than SARS-CoV-1. Understanding the protein’s role and its interaction with the host cell could potentially help researchers develop drugs that disrupt the interaction.
“In the end, what will beat COVID-19 is vaccines. But it is good to keep building knowledge about how viruses and infectious agents work because eventually we will need other tools to defeat them. The more we know about them, the better,” Clapham says.
The next step for Miller and the team is to figure out exactly what the Orf3a protein is doing when it interacts with proteins in the host cell and what advantage the protein gives the virus.
COVID’s impact
COVID-19 led Miller down a different research path than the one she anticipated when she started at Janelia. But Miller says understanding Orf3a’s function seemed like a timely and important question to answer and could contribute to scientists’ overall understanding of the physiology of the SARS-CoV-2 virus. And that, in turn, could inform the development of drugs to combat infection.
“It just felt like the right thing to do,” Miller says. “If I have this ability, I might as well contribute to the greater good.”
And, she adds, that’s just how science works. “I think it is important to pursue what you are interested in, and interesting questions. Where you are today is not necessarily where you are going to be five years from now.”
The research also highlighted for her the importance of collaboration. During the early days of the pandemic, when there were limitations on in-person lab work, Miller had to rely on the help and expertise of others to get the work done. This included other researchers in the Clapham Lab, but also other Janelia labs and support teams, such as the Cryo-Electron Microscopy Facility. In addition, Miller participated in weekly discussions comprising a group of Janelia scientists also working on COVID-19-related projects.
“I was very fortunate to have support from leadership, as well as resources at Janelia to do the work,” Miller says. “To be able to just walk in and start doing these experiments is a luxury, and not everyone gets the opportunity to do that. They encouraged me and let me do it, so I am really very grateful.”
###
Citation:
Alexandria N. Miller, Patrick R. Houlihan, Ella Matamala, Deny Cabezas-Bratesco, Gi Young Lee, Ben Cristofori-Armstrong, Tanya L. Dilan, Silvia Sanchez-Martinez, Doreen Matthies, Rui Yan, Zhiheng Yu, Dejian Ren, Sebastian E. Brauchi, David E. Clapham. “The SARS-CoV-2 accessory protein Orf3a is not an ion channel, but does interact with trafficking proteins.” bioRxiv, posted September 3, 2022. DOI: 10.1101/2022.09.02.506428
Media Contacts
Nanci Bompey