Filter
Associated Lab
- Betzig Lab (1) Apply Betzig Lab filter
- Harris Lab (1) Apply Harris Lab filter
- Heberlein Lab (2) Apply Heberlein Lab filter
- Jayaraman Lab (1) Apply Jayaraman Lab filter
- Ji Lab (1) Apply Ji Lab filter
- Lee (Albert) Lab (1) Apply Lee (Albert) Lab filter
- Lippincott-Schwartz Lab (1) Apply Lippincott-Schwartz Lab filter
- Looger Lab (1) Apply Looger Lab filter
- Pachitariu Lab (3) Apply Pachitariu Lab filter
- Spruston Lab (2) Apply Spruston Lab filter
- Stringer Lab (2) Apply Stringer Lab filter
Associated Project Team
Associated Support Team
Publication Date
- April 29, 2019 (2) Apply April 29, 2019 filter
- April 26, 2019 (2) Apply April 26, 2019 filter
- April 25, 2019 (2) Apply April 25, 2019 filter
- April 19, 2019 (1) Apply April 19, 2019 filter
- April 15, 2019 (1) Apply April 15, 2019 filter
- April 12, 2019 (1) Apply April 12, 2019 filter
- April 11, 2019 (1) Apply April 11, 2019 filter
- April 10, 2019 (1) Apply April 10, 2019 filter
- April 4, 2019 (2) Apply April 4, 2019 filter
- April 1, 2019 (3) Apply April 1, 2019 filter
- Remove April 2019 filter April 2019
- Remove 2019 filter 2019
16 Janelia Publications
Showing 1-10 of 16 resultsThe three-dimensional genome structure plays a fundamental role in gene regulation and cellular functions. Recent studies in genomics based on sequencing technologies inferred the very basic functional chromatin folding structures of the genome known as chromatin loops, the long-range chromatin interactions that are often mediated by protein factors. To visualize the looping structure of chromatin we applied super-resolution microscopy iPALM to image a specific chromatin loop in GM12878 cells. Totally, we have generated six images of the target chromatin region at the single molecule resolution. To infer the chromatin structures from the captured images, we modeled them as looping conformations using different computational algorithms and then evaluated the models by comparing with Hi-C data to examine the concordance. The results showed a good correlation between the imaging data and sequencing data, suggesting the visualization of higher-order chromatin structures for the very short genomic segments can be realized by microscopic imaging.
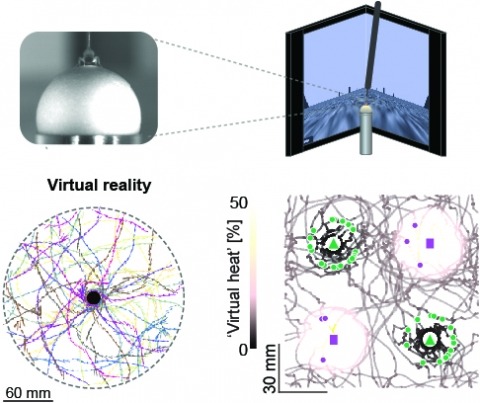
Studying the intertwined roles of sensation, experience, and directed action in navigation has been facilitated by the development of virtual reality (VR) environments for head-fixed animals, allowing for quantitative measurements of behavior in well-controlled conditions. VR has long featured in studies of Drosophila melanogaster, but these experiments have typically allowed the fly to change only its heading in a visual scene and not its position. Here we explore how flies move in two dimensions (2D) using a visual VR environment that more closely captures an animal's experience during free behavior. We show that flies' 2D interaction with landmarks cannot be automatically derived from their orienting behavior under simpler one-dimensional (1D) conditions. Using novel paradigms, we then demonstrate that flies in 2D VR adapt their behavior in response to optogenetically delivered appetitive and aversive stimuli. Much like free-walking flies after encounters with food, head-fixed flies exploring a 2D VR respond to optogenetic activation of sugar-sensing neurons by initiating a local search, which appears not to rely on visual landmarks. Visual landmarks can, however, help flies to avoid areas in VR where they experience an aversive, optogenetically generated heat stimulus. By coupling aversive virtual heat to the flies' presence near visual landmarks of specific shapes, we elicit selective learned avoidance of those landmarks. Thus, we demonstrate that head-fixed flies adaptively navigate in 2D virtual environments, but their reliance on visual landmarks is context dependent. These behavioral paradigms set the stage for interrogation of the fly brain circuitry underlying flexible navigation in complex multisensory environments.
Female behavior changes profoundly after mating. In Drosophila, the mechanisms underlying the long-term changes led by seminal products have been extensively studied. However, the effect of the sensory component of copulation on the female's internal state and behavior remains elusive. We pursued this question by dissociating the effect of coital sensory inputs from those of male ejaculate. We found that the sensory inputs of copulation cause a reduction of post-coital receptivity in females, referred to as the "copulation effect." We identified three layers of a neural circuit underlying this phenomenon. Abdominal neurons expressing the mechanosensory channel Piezo convey the signal of copulation to female-specific ascending neurons, LSANs, in the ventral nerve cord. LSANs relay this information to neurons expressing myoinhibitory peptides in the brain. We hereby provide a neural mechanism by which the experience of copulation facilitates females encoding their mating status, thus adjusting behavior to optimize reproduction.
Cells in the brain act as components of extended networks. Therefore, to understand neurobiological processes in a physiological context, it is essential to study them in vivo. Super-resolution microscopy has spatial resolution beyond the diffraction limit, thus promising to provide structural and functional insights that are not accessible with conventional microscopy. However, to apply it to in vivo brain imaging, we must address the challenges of 3D imaging in an optically heterogeneous tissue that is constantly in motion. We optimized image acquisition and reconstruction to combat sample motion and applied adaptive optics to correcting sample-induced optical aberrations in super-resolution structured illumination microscopy (SIM) in vivo. We imaged the brains of live zebrafish larvae and mice and observed the dynamics of dendrites and dendritic spines at nanoscale resolution.
Human leukocyte antigen (HLA) is a key genetic factor conferring risk of systemic lupus erythematosus (SLE), but precise independent localization of HLA effects is extremely challenging. As a result, the contribution of specific HLA alleles and amino-acid residues to the overall risk of SLE and to risk of specific autoantibodies are far from completely understood. Here, we dissected (a) overall SLE association signals across HLA, (b) HLA-peptide interaction, and (c) residue-autoantibody association. Classical alleles, SNPs, and amino-acid residues of eight HLA genes were imputed across 4,915 SLE cases and 13,513 controls from Eastern Asia. We performed association followed by conditional analysis across HLA, assessing both overall SLE risk and risk of autoantibody production. DR15 alleles HLA-DRB1*15:01 (P = 1.4x10-27, odds ratio (OR) = 1.57) and HLA-DQB1*06:02 (P = 7.4x10-23, OR = 1.55) formed the most significant haplotype (OR = 2.33). Conditioned protein-residue signals were stronger than allele signals and mapped predominantly to HLA-DRB1 residue 13 (P = 2.2x10-75) and its proxy position 11 (P = 1.1x10-67), followed by HLA-DRB1-37 (P = 4.5x10-24). After conditioning on HLA-DRB1, novel associations at HLA-A-70 (P = 1.4x10-8), HLA-DPB1-35 (P = 9.0x10-16), HLA-DQB1-37 (P = 2.7x10-14), and HLA-B-9 (P = 6.5x10-15) emerged. Together, these seven residues increased the proportion of explained heritability due to HLA to 2.6%. Risk residues for both overall disease and hallmark autoantibodies (i.e., nRNP: DRB1-11, P = 2.0x10-14; DRB1-13, P = 2.9x10-13; DRB1-30, P = 3.9x10-14) localized to the peptide-binding groove of HLA-DRB1. Enrichment for specific amino-acid characteristics in the peptide-binding groove correlated with overall SLE risk and with autoantibody presence. Risk residues were in primarily negatively charged side-chains, in contrast with rheumatoid arthritis. We identified novel SLE signals in HLA Class I loci (HLA-A, HLA-B), and localized primary Class II signals to five residues in HLA-DRB1, HLA-DPB1, and HLA-DQB1. These findings provide insights about the mechanisms by which the risk residues interact with each other to produce autoantibodies and are involved in SLE pathophysiology.
Sensory cortices are active in the absence of external sensory stimuli. To understand the nature of this ongoing activity, we used two-photon calcium imaging to record from over 10,000 neurons in the visual cortex of mice awake in darkness while monitoring their behavior videographically. Ongoing population activity was multidimensional, exhibiting at least 100 significant dimensions, some of which were related to the spontaneous behaviors of the mice. The largest single dimension was correlated with the running speed and pupil area, while a 16-dimensional summary of orofacial behaviors could predict ~45% of the explainable neural variance. Electrophysiological recordings with 8 simultaneous Neuropixels probes revealed a similar encoding of high-dimensional orofacial behaviors across multiple forebrain regions. Representation of motor variables continued uninterrupted during visual stimulus presentation, occupying dimensions nearly orthogonal to the stimulus responses. Our results show that a multidimensional representation of motor state is encoded across the forebrain, and is integrated with visual input by neuronal populations in primary visual cortex.
Ascidian species of the Phallusia and Ciona genera are distantly related, their last common ancestor dating several hundred million years ago. Although their genome sequences have extensively diverged since this radiation, Phallusia and Ciona species share almost identical early morphogenesis and stereotyped cell lineages. Here, we explored the evolution of transcriptional control between P. mammillata and C. robusta. We combined genome-wide mapping of open chromatin regions in both species with a comparative analysis of the regulatory sequences of a test set of 10 pairs of orthologous early regulatory genes with conserved expression patterns. We find that ascidian chromatin accessibility landscapes obey similar rules as in other metazoa. Open-chromatin regions are short, highly conserved within each genus and cluster around regulatory genes. The dynamics of chromatin accessibility and closest-gene expression are strongly correlated during early embryogenesis. Open-chromatin regions are highly enriched in cis-regulatory elements: 73% of 49 open chromatin regions around our test genes behaved as either distal enhancers or proximal enhancer/promoters following electroporation in Phallusia eggs. Analysis of this datasets suggests a pervasive use in ascidians of "shadow" enhancers with partially overlapping activities. Cross-species electroporations point to a deep conservation of both the trans-regulatory logic between these distantly-related ascidians and the cis-regulatory activities of individual enhancers. Finally, we found that the relative order and approximate distance to the transcription start site of open chromatin regions can be conserved between Ciona and Phallusia species despite extensive sequence divergence, a property that can be used to identify orthologous enhancers, whose regulatory activity can partially diverge.
Understanding the principles governing neuronal diversity is a fundamental goal for neuroscience. Here we provide an anatomical and transcriptomic database of nearly 200 genetically identified cell populations. By separately analyzing the robustness and pattern of expression differences across these cell populations, we identify two gene classes contributing distinctly to neuronal diversity. Short homeobox transcription factors distinguish neuronal populations combinatorially, and exhibit extremely low transcriptional noise, enabling highly robust expression differences. Long neuronal effector genes, such as channels and cell adhesion molecules, contribute disproportionately to neuronal diversity, based on their patterns rather than robustness of expression differences. By linking transcriptional identity to genetic strains and anatomical atlases we provide an extensive resource for further investigation of mouse neuronal cell types.
Reconstruction of neural circuitry at single-synapse resolution is an attractive target for improving understanding of the nervous system in health and disease. Serial section transmission electron microscopy (ssTEM) is among the most prolific imaging methods employed in pursuit of such reconstructions. We demonstrate how Flood-Filling Networks (FFNs) can be used to computationally segment a forty-teravoxel whole-brain Drosophila ssTEM volume. To compensate for data irregularities and imperfect global alignment, FFNs were combined with procedures that locally re-align serial sections and dynamically adjust image content. The proposed approach produced a largely merger-free segmentation of the entire ssTEM Drosophila brain, which we make freely available. As compared to manual tracing using an efficient skeletonization strategy, the segmentation enabled circuit reconstruction and analysis workflows that were an order of magnitude faster.