Filter
Associated Lab
- Ahrens Lab (1) Apply Ahrens Lab filter
- Aso Lab (1) Apply Aso Lab filter
- Baker Lab (3) Apply Baker Lab filter
- Betzig Lab (7) Apply Betzig Lab filter
- Bock Lab (2) Apply Bock Lab filter
- Branson Lab (1) Apply Branson Lab filter
- Cardona Lab (1) Apply Cardona Lab filter
- Cui Lab (3) Apply Cui Lab filter
- Dickson Lab (2) Apply Dickson Lab filter
- Druckmann Lab (1) Apply Druckmann Lab filter
- Dudman Lab (1) Apply Dudman Lab filter
- Eddy/Rivas Lab (5) Apply Eddy/Rivas Lab filter
- Fetter Lab (4) Apply Fetter Lab filter
- Fitzgerald Lab (1) Apply Fitzgerald Lab filter
- Gonen Lab (5) Apply Gonen Lab filter
- Grigorieff Lab (6) Apply Grigorieff Lab filter
- Heberlein Lab (12) Apply Heberlein Lab filter
- Hermundstad Lab (1) Apply Hermundstad Lab filter
- Hess Lab (2) Apply Hess Lab filter
- Jayaraman Lab (2) Apply Jayaraman Lab filter
- Ji Lab (2) Apply Ji Lab filter
- Kainmueller Lab (1) Apply Kainmueller Lab filter
- Keller Lab (4) Apply Keller Lab filter
- Lavis Lab (5) Apply Lavis Lab filter
- Lee (Albert) Lab (1) Apply Lee (Albert) Lab filter
- Leonardo Lab (2) Apply Leonardo Lab filter
- Lippincott-Schwartz Lab (18) Apply Lippincott-Schwartz Lab filter
- Liu (Zhe) Lab (1) Apply Liu (Zhe) Lab filter
- Looger Lab (7) Apply Looger Lab filter
- Magee Lab (1) Apply Magee Lab filter
- Menon Lab (3) Apply Menon Lab filter
- Murphy Lab (1) Apply Murphy Lab filter
- Pastalkova Lab (1) Apply Pastalkova Lab filter
- Pavlopoulos Lab (2) Apply Pavlopoulos Lab filter
- Reiser Lab (2) Apply Reiser Lab filter
- Riddiford Lab (1) Apply Riddiford Lab filter
- Romani Lab (1) Apply Romani Lab filter
- Rubin Lab (4) Apply Rubin Lab filter
- Satou Lab (3) Apply Satou Lab filter
- Scheffer Lab (2) Apply Scheffer Lab filter
- Schreiter Lab (3) Apply Schreiter Lab filter
- Sgro Lab (2) Apply Sgro Lab filter
- Simpson Lab (3) Apply Simpson Lab filter
- Singer Lab (10) Apply Singer Lab filter
- Spruston Lab (1) Apply Spruston Lab filter
- Stern Lab (4) Apply Stern Lab filter
- Sternson Lab (6) Apply Sternson Lab filter
- Svoboda Lab (7) Apply Svoboda Lab filter
- Tjian Lab (4) Apply Tjian Lab filter
- Truman Lab (1) Apply Truman Lab filter
- Turaga Lab (1) Apply Turaga Lab filter
- Turner Lab (2) Apply Turner Lab filter
- Zlatic Lab (1) Apply Zlatic Lab filter
- Zuker Lab (3) Apply Zuker Lab filter
Associated Project Team
Publication Date
- December 2011 (22) Apply December 2011 filter
- November 2011 (15) Apply November 2011 filter
- October 2011 (14) Apply October 2011 filter
- September 2011 (17) Apply September 2011 filter
- August 2011 (14) Apply August 2011 filter
- July 2011 (10) Apply July 2011 filter
- June 2011 (17) Apply June 2011 filter
- May 2011 (13) Apply May 2011 filter
- April 2011 (11) Apply April 2011 filter
- March 2011 (14) Apply March 2011 filter
- February 2011 (16) Apply February 2011 filter
- January 2011 (27) Apply January 2011 filter
- Remove 2011 filter 2011
Type of Publication
190 Publications
Showing 181-190 of 190 resultsStarvation induces a protective process of self-cannibalization called autophagy that is thought to mediate nonselective degradation of cytoplasmic material. We recently reported that mitochondria escape autophagosomal degradation through extensive fusion into mitochondrial networks upon certain starvation conditions. The extent of mitochondrial elongation is dependent on the type of nutrient deprivation, with amino acid depletion having a particularly strong effect. Downregulation of the mitochondrial fission protein Drp1 was determined to be important in bringing about starvation-induced mitochondrial fusion. The formation of mitochondrial networks during nutrient depletion selectively blocked their autophagic degradation, presumably allowing cells to sustain efficient ATP production and thereby survive starvation.
Expression of an individual gene can vary considerably among genetically identical cells because of stochastic fluctuations in transcription. However, proteins comprising essential complexes or pathways have similar abundances and lower variability. It is not known whether coordination in the expression of subunits of essential complexes occurs at the level of transcription, mRNA abundance or protein expression. To directly measure the level of coordination in the expression of genes, we used highly sensitive fluorescence in situ hybridization (FISH) to count individual mRNAs of functionally related and unrelated genes within single Saccharomyces cerevisiae cells. Our results revealed that transcript levels of temporally induced genes are highly correlated in individual cells. In contrast, transcription of constitutive genes encoding essential subunits of complexes is not coordinated because of stochastic fluctuations. The coordination of these functional complexes therefore must occur post-transcriptionally, and likely post-translationally.
Mitochondria are highly dynamic organelles that mediate essential cell functions such as apoptosis and cell-cycle control in addition to their role as efficient ATP generators. Mitochondrial morphology changes are tightly regulated, and their shape can shift between small, fragmented units and larger networks of elongated mitochondria. We demonstrate that mitochondrial elements become significantly elongated and interconnected shortly after nutrient depletion. This mitochondrial morphological shift depends on the type of starvation, with an additive effect observed when multiple nutrients are depleted simultaneously. We further show that starvation-induced mitochondrial elongation is mediated by down-regulation of dynamin-related protein 1 (Drp1) through modulation of two Drp1 phosphorylation sites, leading to unopposed mitochondrial fusion. Finally, we establish that mitochondrial tubulation upon nutrient deprivation protects mitochondria from autophagosomal degradation, which could permit mitochondria to maximize energy production and supply autophagosomal membranes during starvation.
The innate sexual behaviors of Drosophila melanogaster males are an attractive system for elucidating how complex behavior patterns are generated. The potential for male sexual behavior in D. melanogaster is specified by the fruitless (fru) and doublesex (dsx) sex regulatory genes. We used the temperature-sensitive activator dTRPA1 to probe the roles of fru(M)- and dsx-expressing neurons in male courtship behaviors. Almost all steps of courtship, from courtship song to ejaculation, can be induced at very high levels through activation of either all fru(M) or all dsx neurons in solitary males. Detailed characterizations reveal different roles for fru(M) and dsx in male courtship. Surprisingly, the system for mate discrimination still works well when all dsx neurons are activated, but is impaired when all fru(M) neurons are activated. Most strikingly, we provide evidence for a fru(M)-independent courtship pathway that is primarily vision dependent.
Multiphoton imaging (MPI) is widely used for recording activity simultaneously from many neurons in superficial cortical layers in vivo. We combined regenerative amplification multiphoton microscopy (RAMM) with genetically encoded calcium indicators to extend MPI of neuronal population activity into layer 5 (L5) of adult mouse somatosensory cortex. We found that this approach could be used to record and quantify spontaneous and sensory-evoked activity in populations of L5 neuronal somata located as much as 800 μm below the pia. In addition, we found that RAMM could be used to simultaneously image activity from large (80) populations of apical dendrites and follow these dendrites down to their somata of origin.
Mammalian genomes contain numerous regulatory DNA sites with unknown target genes. We used mice with an extra β-globin locus control region (LCR) to investigate how a regulator searches the genome for target genes. We find that the LCR samples a restricted nuclear subvolume, wherein it preferentially contacts genes controlled by shared transcription factors. No contacted gene is detectably upregulated except for endogenous β-globin genes located on another chromosome. This demonstrates genetically that mammalian trans activation is possible, but suggests that it will be rare. Trans activation occurs not pan-cellularly, but in 'jackpot' cells enriched for the interchromosomal interaction. Therefore, cell-specific long-range DNA contacts can cause variegated expression.
The ability of insects to learn and navigate to specific locations in the environment has fascinated naturalists for decades. The impressive navigational abilities of ants, bees, wasps and other insects demonstrate that insects are capable of visual place learning, but little is known about the underlying neural circuits that mediate these behaviours. Drosophila melanogaster (common fruit fly) is a powerful model organism for dissecting the neural circuitry underlying complex behaviours, from sensory perception to learning and memory. Drosophila can identify and remember visual features such as size, colour and contour orientation. However, the extent to which they use vision to recall specific locations remains unclear. Here we describe a visual place learning platform and demonstrate that Drosophila are capable of forming and retaining visual place memories to guide selective navigation. By targeted genetic silencing of small subsets of cells in the Drosophila brain, we show that neurons in the ellipsoid body, but not in the mushroom bodies, are necessary for visual place learning. Together, these studies reveal distinct neuroanatomical substrates for spatial versus non-spatial learning, and establish Drosophila as a powerful model for the study of spatial memories.
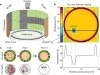
We have developed miniature telemetry systems that capture neural, EMG, and acceleration signals from a freely moving insect or other small animal and transmit the data wirelessly to a remote digital receiver. The systems are based on custom low-power integrated circuits (ICs) that amplify, filter, and digitize four biopotential signals using low-noise circuits. One of the chips also digitizes three acceleration signals from an off-chip microelectromechanical-system accelerometer. All information is transmitted over a wireless ~ 900-MHz telemetry link. The first unit, using a custom chip fabricated in a 0.6- μm BiCMOS process, weighs 0.79 g and runs for two hours on two small batteries. We have used this system to monitor neural and EMG signals in jumping and flying locusts as well as transdermal potentials in weakly swimming electric fish. The second unit, using a custom chip fabricated in a 0.35-μ m complementary metal-oxide semiconductor CMOS process, weighs 0.17 g and runs for five hours on a single 1.5-V battery. This system has been used to monitor neural potentials in untethered perching dragonflies.
Wiring economy has successfully explained the individual placement of neurons in simple nervous systems like that of Caenorhabditis elegans [1-3] and the locations of coarser structures like cortical areas in complex vertebrate brains [4]. However, it remains unclear whether wiring economy can explain the placement of individual neurons in brains larger than that of C. elegans. Indeed, given the greater number of neuronal interconnections in larger brains, simply minimizing the length of connections results in unrealistic configurations, with multiple neurons occupying the same position in space. Avoiding such configurations, or volume exclusion, repels neurons from each other, thus counteracting wiring economy. Here we test whether wiring economy together with volume exclusion can explain the placement of neurons in a module of the Drosophila melanogaster brain known as lamina cartridge [5-13]. We used newly developed techniques for semiautomated reconstruction from serial electron microscopy (EM) [14] to obtain the shapes of neurons, the location of synapses, and the resultant synaptic connectivity. We show that wiring length minimization and volume exclusion together can explain the structure of the lamina microcircuit. Therefore, even in brains larger than that of C. elegans, at least for some circuits, optimization can play an important role in individual neuron placement.