Filter
Associated Lab
- Ahrens Lab (7) Apply Ahrens Lab filter
- Aso Lab (2) Apply Aso Lab filter
- Baker Lab (2) Apply Baker Lab filter
- Betzig Lab (12) Apply Betzig Lab filter
- Bock Lab (1) Apply Bock Lab filter
- Branson Lab (5) Apply Branson Lab filter
- Card Lab (3) Apply Card Lab filter
- Cardona Lab (8) Apply Cardona Lab filter
- Dickson Lab (4) Apply Dickson Lab filter
- Druckmann Lab (3) Apply Druckmann Lab filter
- Dudman Lab (3) Apply Dudman Lab filter
- Eddy/Rivas Lab (1) Apply Eddy/Rivas Lab filter
- Egnor Lab (1) Apply Egnor Lab filter
- Fetter Lab (6) Apply Fetter Lab filter
- Fitzgerald Lab (1) Apply Fitzgerald Lab filter
- Freeman Lab (4) Apply Freeman Lab filter
- Funke Lab (2) Apply Funke Lab filter
- Gonen Lab (8) Apply Gonen Lab filter
- Grigorieff Lab (6) Apply Grigorieff Lab filter
- Harris Lab (5) Apply Harris Lab filter
- Hess Lab (2) Apply Hess Lab filter
- Jayaraman Lab (3) Apply Jayaraman Lab filter
- Ji Lab (4) Apply Ji Lab filter
- Kainmueller Lab (3) Apply Kainmueller Lab filter
- Karpova Lab (3) Apply Karpova Lab filter
- Keleman Lab (1) Apply Keleman Lab filter
- Keller Lab (6) Apply Keller Lab filter
- Lavis Lab (13) Apply Lavis Lab filter
- Lee (Albert) Lab (1) Apply Lee (Albert) Lab filter
- Leonardo Lab (2) Apply Leonardo Lab filter
- Li Lab (2) Apply Li Lab filter
- Lippincott-Schwartz Lab (9) Apply Lippincott-Schwartz Lab filter
- Liu (Zhe) Lab (4) Apply Liu (Zhe) Lab filter
- Looger Lab (10) Apply Looger Lab filter
- Magee Lab (2) Apply Magee Lab filter
- Menon Lab (2) Apply Menon Lab filter
- Murphy Lab (2) Apply Murphy Lab filter
- Pachitariu Lab (2) Apply Pachitariu Lab filter
- Pastalkova Lab (1) Apply Pastalkova Lab filter
- Pavlopoulos Lab (3) Apply Pavlopoulos Lab filter
- Pedram Lab (1) Apply Pedram Lab filter
- Podgorski Lab (1) Apply Podgorski Lab filter
- Reiser Lab (3) Apply Reiser Lab filter
- Romani Lab (2) Apply Romani Lab filter
- Rubin Lab (3) Apply Rubin Lab filter
- Saalfeld Lab (3) Apply Saalfeld Lab filter
- Schreiter Lab (2) Apply Schreiter Lab filter
- Simpson Lab (1) Apply Simpson Lab filter
- Singer Lab (5) Apply Singer Lab filter
- Spruston Lab (6) Apply Spruston Lab filter
- Stern Lab (7) Apply Stern Lab filter
- Sternson Lab (3) Apply Sternson Lab filter
- Stringer Lab (1) Apply Stringer Lab filter
- Svoboda Lab (8) Apply Svoboda Lab filter
- Tervo Lab (2) Apply Tervo Lab filter
- Tillberg Lab (1) Apply Tillberg Lab filter
- Tjian Lab (3) Apply Tjian Lab filter
- Truman Lab (11) Apply Truman Lab filter
- Turaga Lab (2) Apply Turaga Lab filter
- Turner Lab (1) Apply Turner Lab filter
- Wang (Shaohe) Lab (2) Apply Wang (Shaohe) Lab filter
- Wu Lab (1) Apply Wu Lab filter
- Zlatic Lab (2) Apply Zlatic Lab filter
Associated Project Team
- Fly Functional Connectome (2) Apply Fly Functional Connectome filter
- Fly Olympiad (1) Apply Fly Olympiad filter
- FlyEM (1) Apply FlyEM filter
- GENIE (2) Apply GENIE filter
- MouseLight (2) Apply MouseLight filter
- Tool Translation Team (T3) (1) Apply Tool Translation Team (T3) filter
- Transcription Imaging (11) Apply Transcription Imaging filter
Publication Date
- December 2016 (18) Apply December 2016 filter
- November 2016 (15) Apply November 2016 filter
- October 2016 (22) Apply October 2016 filter
- September 2016 (11) Apply September 2016 filter
- August 2016 (13) Apply August 2016 filter
- July 2016 (15) Apply July 2016 filter
- June 2016 (25) Apply June 2016 filter
- May 2016 (23) Apply May 2016 filter
- April 2016 (14) Apply April 2016 filter
- March 2016 (15) Apply March 2016 filter
- February 2016 (23) Apply February 2016 filter
- January 2016 (15) Apply January 2016 filter
- Remove 2016 filter 2016
Type of Publication
209 Publications
Showing 101-110 of 209 resultsElectrical coupling in circuits can produce non-intuitive circuit dynamics, as seen in both experimental work from the crustacean stomatogastric ganglion and in computational models inspired by the connectivity in this preparation. Ambiguities in interpreting the results of electrophysiological recordings can arise if sets of pre- or postsynaptic neurons are electrically coupled, or if the electrical coupling exhibits some specificity (e.g. rectifying, or voltage-dependent). Even in small circuits, electrical coupling can produce parallel pathways that can allow information to travel by monosynaptic and/or polysynaptic pathways. Consequently, similar changes in circuit dynamics can arise from entirely different underlying mechanisms. When neurons are coupled both chemically and electrically, modifying the relative strengths of the two interactions provides a mechanism for flexibility in circuit outputs. This, together with neuromodulation of gap junctions and coupled neurons is important both in developing and adult circuits. This article is protected by copyright. All rights reserved.
Microelectron diffraction (MicroED) is a new cryo-electron microscopy (cryo-EM) method capable of determining macromolecular structures at atomic resolution from vanishingly small 3D crystals. MicroED promises to solve atomic resolution structures from even the tiniest of crystals, less than a few hundred nanometers thick. MicroED complements frontier advances in crystallography and represents part of the rebirth of cryo-EM that is making macromolecular structure determination more accessible for all. Here we review the concept and practice of MicroED, for both the electron microscopist and crystallographer. Where other reviews have addressed specific details of the technique (Hattne et al., 2015, Shi et al., 2016 and Shi et al., 2013), we aim to provide context and highlight important features that should be considered when performing a MicroED experiment.
Following considerable progress on the molecular and cellular basis of taste perception in fly sensory neurons, the time is now ripe to explore how taste information, integrated with hunger and satiety, undergo a sensorimotor transformation to lead to the motor actions of feeding behavior. I examine what is known of feeding circuitry in adult flies from more than 250 years of work in larger flies and from newer work in Drosophila. I review the anatomy of the proboscis, its muscles and their functions (where known), its motor neurons, interneurons known to receive taste inputs, interneurons that diverge from taste circuitry to provide information to other circuits, interneurons from other circuits that converge on feeding circuits, proprioceptors that influence the motor control of feeding, and sites of integration of hunger and satiety on feeding circuits. In spite of the several neuron types now known, a connected pathway from taste inputs to feeding motor outputs has yet to be found. We are on the threshold of an era where these individual components will be assembled into circuits, revealing how nervous system architecture leads to the control of behavior.
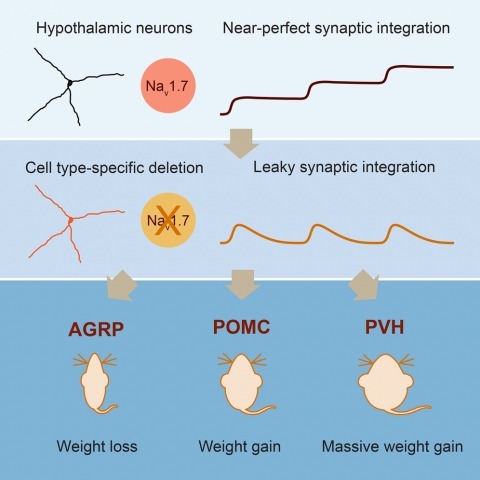
Neurons are well suited for computations on millisecond timescales, but some neuronal circuits set behavioral states over long time periods, such as those involved in energy homeostasis. We found that multiple types of hypothalamic neurons, including those that oppositely regulate body weight, are specialized as near-perfect synaptic integrators that summate inputs over extended timescales. Excitatory postsynaptic potentials (EPSPs) are greatly prolonged, outlasting the neuronal membrane time-constant up to 10-fold. This is due to the voltage-gated sodium channel Nav1.7 (Scn9a), previously associated with pain-sensation but not synaptic integration. Scn9a deletion in AGRP, POMC, or paraventricular hypothalamic neurons reduced EPSP duration, synaptic integration, and altered body weight in mice. In vivo whole-cell recordings in the hypothalamus confirmed near-perfect synaptic integration. These experiments show that integration of synaptic inputs over time by Nav1.7 is critical for body weight regulation and reveal a mechanism for synaptic control of circuits regulating long term homeostatic functions.
The icosahedron is the largest of the Platonic solids, and icosahedral protein structures are widely used in biological systems for packaging and transport. There has been considerable interest in repurposing such structures for applications ranging from targeted delivery to multivalent immunogen presentation. The ability to design proteins that self-assemble into precisely specified, highly ordered icosahedral structures would open the door to a new generation of protein containers with properties custom-tailored to specific applications. Here we describe the computational design of a 25-nanometre icosahedral nanocage that self-assembles from trimeric protein building blocks. The designed protein was produced in Escherichia coli, and found by electron microscopy to assemble into a homogenous population of icosahedral particles nearly identical to the design model. The particles are stable in 6.7 molar guanidine hydrochloride at up to 80 degrees Celsius, and undergo extremely abrupt, but reversible, disassembly between 2 molar and 2.25 molar guanidinium thiocyanate. The icosahedron is robust to genetic fusions: one or two copies of green fluorescent protein (GFP) can be fused to each of the 60 subunits to create highly fluorescent 'standard candles' for use in light microscopy, and a designed protein pentamer can be placed in the centre of each of the 20 pentameric faces to modulate the size of the entrance/exit channels of the cage. Such robust and customizable nanocages should have considerable utility in targeted drug delivery, vaccine design and synthetic biology.
With recent advances in high-throughput Electron Microscopy (EM) imaging it is now possible to image an entire nervous system of organisms like Drosophila melanogaster. One of the bottlenecks to reconstruct a connectome from these large volumes (œ 100 TiB) is the pixel-wise prediction of membranes. The time it would typically take to process such a volume using a convolutional neural network (CNN) with a sliding window approach is in the order of years on a current GPU. With sliding windows, however, a lot of redundant computations are carried out. In this paper, we present an extension to the Caffe library to increase throughput by predicting many pixels at once. On a sliding window network successfully used for membrane classification, we show that our method achieves a speedup of up to 57×, maintaining identical prediction results.
Multi-modal image registration is a challenging task that is vital to fuse complementary signals for subsequent analyses. Despite much research into cost functions addressing this challenge, there exist cases in which these are ineffective. In this work, we show that (1) this is true for the registration of in-vivo Drosophila brain volumes visualizing genetically encoded calcium indicators to an nc82 atlas and (2) that machine learning based contrast synthesis can yield improvements. More specifically, the number of subjects for which the registration outright failed was greatly reduced (from 40% to 15%) by using a synthesized image.
Imaging is used to map activity across populations of neurons. Microscopes with cellular resolution have small (<1 millimeter) fields of view and cannot simultaneously image activity distributed across multiple brain areas. Typical large field of view microscopes do not resolve single cells, especially in the axial dimension. We developed a 2-photon random access mesoscope (2p-RAM) that allows high-resolution imaging anywhere within a volume spanning multiple brain areas (∅ 5 mm x 1 mm cylinder). 2p-RAM resolution is near diffraction limited (lateral, 0.66 μm, axial 4.09 μm at the center; excitation wavelength = 970 nm; numerical aperture = 0.6) over a large range of excitation wavelengths. A fast three-dimensional scanning system allows efficient sampling of neural activity in arbitrary regions of interest across the entire imaging volume. We illustrate the use of the 2p-RAM by imaging neural activity in multiple, non-contiguous brain areas in transgenic mice expressing protein calcium sensors.
Startle behaviors are rapid, high-performance motor responses to threatening stimuli. Startle responses have been identified in a broad range of species across animal diversity. For investigations of neural circuit structure and function, these behaviors offer a number of benefits, including that they are driven by large and identifiable neurons and their neural control is simple in comparison to other behaviors. Among vertebrates, the best-known startle circuit is the Mauthner cell circuit of fishes. In recent years, genetic approaches in zebrafish have provided key tools for morphological and physiological dissection of circuits and greatly extended understanding of their architecture. Here we discuss the startle circuit of fishes, with a focus on the Mauthner cells and associated interneurons called spiral fiber neurons and we add new observations on hindbrain circuit organization. We also briefly review and compare startle circuits of several other taxa, paying particular attention to how movement direction is controlled.
The diffraction limited resolution of two photon and confocal microscope can be recovered using adaptive optics to explore the detailed neuronal network in the brains of zebrafish and mouse in vivo.