The method melds the best of super-resolution fluorescence and electron microscopy to show how proteins relate to cells’ fine structure.
Inside a cell, tentacled vesicles shuttle cargo for sorting. Neighboring neurons cling to one another through a web-like interface. DNA rearranges in the nucleus as stem cells differentiate into neurons. And a new microscopy technique shows it all, in exquisite detail.
The technique, called cryo-SR/EM, melds images captured from electron microscopes and super-resolution light microscopes, resulting in brilliant, clear, detailed views of the inside of cells – in 3-D.
For years, scientists have probed the microscopic world inside cells, developing new tools to view these basic units of life. But each tool comes with a tradeoff. Light microscopy makes it simple to identify specific cellular structures by tagging them with easy-to-see fluorescent molecules. With the development of super-resolution (SR) fluorescence microscopy, these structures can be viewed with even greater clarity. But fluorescence can reveal only a few of the more than 10,000 proteins in a cell at a given time, making it difficult to understand how these few relate to everything else. Electron microscopy (EM), on the other hand, reveals all cellular structures in high-resolution pictures – but delineating one feature from all others by EM alone can be difficult because the space inside cells is so crowded.
Combining the two techniques gives scientists a clear picture of how specific cellular features relate to their surroundings, says Harald Hess, a senior group leader at the Howard Hughes Medical Institute’s Janelia Research Campus. “This is a very powerful method.”
Janelia Research Scientist David Hoffman and Senior Scientist Gleb Shtengel spearheaded the project under the leadership of Hess and Janelia senior fellow Eric Betzig, an HHMI Investigator at the University of California, Berkeley. The work is described January 16, 2020, in the journal Science.
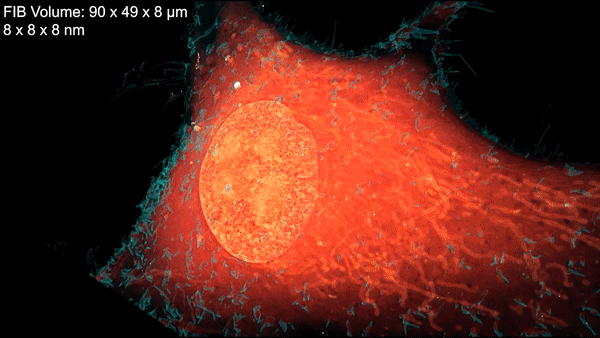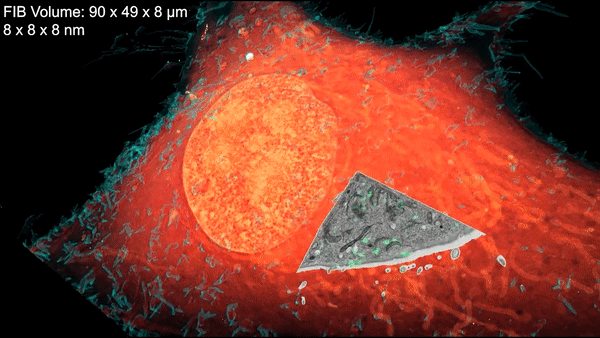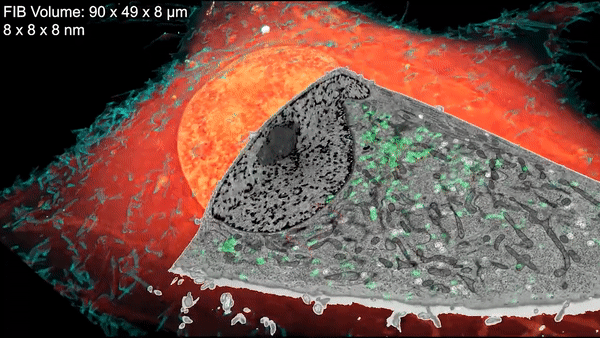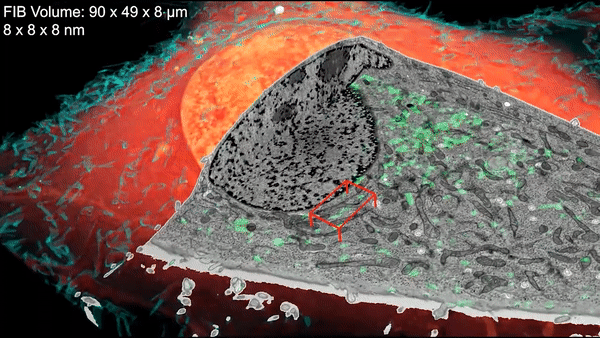
First, the scientists freeze cells under high pressure. That halts the cells’ activity quickly and prevents the formation of ice crystals that can damage cells and disrupt the structures being imaged. Next, the researchers place samples in a cryogenic chamber, where they’re imaged in 3-D by super-resolution fluorescence microscopy at temperatures less than ten degrees above absolute zero. Then, they’re removed, embedded in resin, and imaged in a powerful electron microscope developed by the Hess lab. This scope shoots a beam of ions at the cells’ surface, milling away bit by bit while taking pictures of each newly exposed layer. A computer program then stitches the images together into a 3-D reconstruction.
Finally, researchers overlay the 3-D image data from both microscopes. The result: stunning imagery that reveals cells’ inner details with remarkable clarity.
Below, a few examples of this imagery illustrate how scientists are using the technique. “There’s already been a lot of interest,” says Hess. “There are so many more experiments to do — a whole world of cells out there to study.”
The nucleus of a neuron looks dramatically different before (left) and after (right) the cell begins to assume its final adult role. As the cell matures, DNA is repackaged within the nucleus to turn on a new set of genes. These changes are reflected in the different patterns of gray mottling and colored fluorescence inside the two cells. “The technique provided an amazingly detailed snapshot of the state of the nucleus before and after differentiation,” says David Solecki of St. Jude Children’s Research Hospital, who collaborated on the project. Credit: D. Hoffman et al./Science 2020
Developing neurons stick together. This video shows exactly how those cells adhere to each other, revealing Swiss-cheese-like linkages that help young neurons properly migrate to their final destinations in the nervous system. Purple and green super-resolution fluorescence images of adhesion proteins at these linkages correlate with electron microscopy images showing the membrane’s structure in detail. Credit: D. Hoffman et al./Science 2020
Cells are filled with small vesicles – membrane-bound sacks that help cells store proteins, break down cellular garbage, and carry cargo. These many varieties of vesicles are indistinguishable from each other under an electron microscope alone. But with cryo-SR/EM, their distinct features become clear. This clip zooms in on endosomes, which shuttle cargo to different regions within the cell. Credit: D. Hoffman et al./Science 2020
Researchers from outside Janelia can access the technique through Janelia’s Advanced Imaging Center.
###
Citation
David P. Hoffman, Gleb Shtengel, C. Shan Xu, Kirby R. Campbell, Melanie Freeman, Lei Wang, Daniel E. Milkie, H. Amalia Pasolli, Nirmala Iyer, John A. Bogovic, Daniel R. Stabley, Abbas Shirinifard, Song Pang, David Peale, Kathy Schaefer, Wim Pomp, Chi-Lun Chang, Jennifer Lippincott-Schwartz, Tom Kirchhausen, David J. Solecki, Eric Betzig, and Harald F. Hess. “Correlative three-dimensional super-resolution and block-face electron microscopy of whole vitreously frozen cells.” Science. Published online January 16, 2020. Doi: 10.1126/science.aaz5357
This paper was first published as a preprint on bioRxiv: https://www.biorxiv.org/content/10.1101/773986v1
View full video gallery
Media Contacts
Laurel Hamers
hamersl@janelia.hhmi.org