Filter
Associated Lab
Associated Project Team
3 Janelia Publications
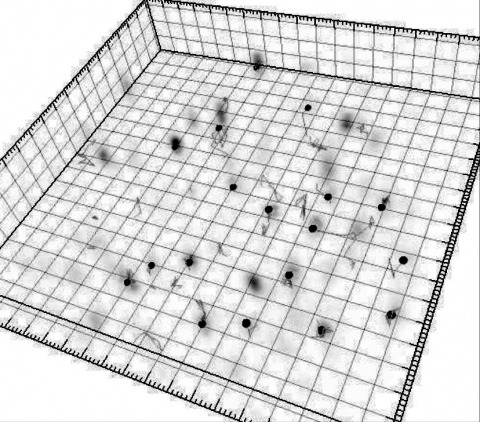
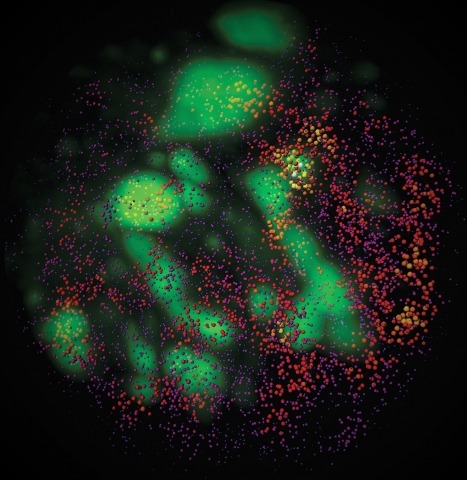
Showing 1-3 of 3 resultsCombinatorial cis-regulatory networks encoded in animal genomes represent the foundational gene expression mechanism for directing cell-fate commitment and maintenance of cell identity by transcription factors (TFs). However, the 3D spatial organization of cis-elements and how such sub-nuclear structures influence TF activity remain poorly understood. Here, we combine lattice light-sheet imaging, single-molecule tracking, numerical simulations, and ChIP-exo mapping to localize and functionally probe Sox2 enhancer-organization in living embryonic stem cells. Sox2 enhancers form 3D-clusters that are segregated from heterochromatin but overlap with a subset of Pol II enriched regions. Sox2 searches for specific binding targets via a 3D-diffusion dominant mode when shuttling long-distances between clusters while chromatin-bound states predominate within individual clusters. Thus, enhancer clustering may reduce global search efficiency but enables rapid local fine-tuning of TF search parameters. Our results suggest an integrated model linking cis-element 3D spatial distribution to local-versus-global target search modalities essential for regulating eukaryotic gene transcription.
Although fluorescence microscopy provides a crucial window into the physiology of living specimens, many biological processes are too fragile, are too small, or occur too rapidly to see clearly with existing tools. We crafted ultrathin light sheets from two-dimensional optical lattices that allowed us to image three-dimensional (3D) dynamics for hundreds of volumes, often at subsecond intervals, at the diffraction limit and beyond. We applied this to systems spanning four orders of magnitude in space and time, including the diffusion of single transcription factor molecules in stem cell spheroids, the dynamic instability of mitotic microtubules, the immunological synapse, neutrophil motility in a 3D matrix, and embryogenesis in Caenorhabditis elegans and Drosophila melanogaster. The results provide a visceral reminder of the beauty and the complexity of living systems.
Enhancer-binding pluripotency regulators (Sox2 and Oct4) play a seminal role in embryonic stem (ES) cell-specific gene regulation. Here, we combine in vivo and in vitro single-molecule imaging, transcription factor (TF) mutagenesis, and ChIP-exo mapping to determine how TFs dynamically search for and assemble on their cognate DNA target sites. We find that enhanceosome assembly is hierarchically ordered with kinetically favored Sox2 engaging the target DNA first, followed by assisted binding of Oct4. Sox2/Oct4 follow a trial-and-error sampling mechanism involving 84-97 events of 3D diffusion (3.3-3.7 s) interspersed with brief nonspecific collisions (0.75-0.9 s) before acquiring and dwelling at specific target DNA (12.0-14.6 s). Sox2 employs a 3D diffusion-dominated search mode facilitated by 1D sliding along open DNA to efficiently locate targets. Our findings also reveal fundamental aspects of gene and developmental regulation by fine-tuning TF dynamics and influence of the epigenome on target search parameters.