Filter
Associated Lab
- Betzig Lab (8) Apply Betzig Lab filter
- Eddy/Rivas Lab (2) Apply Eddy/Rivas Lab filter
- Fetter Lab (1) Apply Fetter Lab filter
- Hess Lab (2) Apply Hess Lab filter
- Ji Lab (4) Apply Ji Lab filter
- Looger Lab (3) Apply Looger Lab filter
- Magee Lab (4) Apply Magee Lab filter
- Riddiford Lab (1) Apply Riddiford Lab filter
- Rubin Lab (2) Apply Rubin Lab filter
- Schreiter Lab (1) Apply Schreiter Lab filter
- Shroff Lab (6) Apply Shroff Lab filter
- Spruston Lab (3) Apply Spruston Lab filter
- Sternson Lab (1) Apply Sternson Lab filter
- Svoboda Lab (6) Apply Svoboda Lab filter
- Truman Lab (1) Apply Truman Lab filter
Publication Date
- December 2008 (8) Apply December 2008 filter
- November 2008 (1) Apply November 2008 filter
- October 2008 (1) Apply October 2008 filter
- September 2008 (2) Apply September 2008 filter
- August 2008 (3) Apply August 2008 filter
- July 2008 (4) Apply July 2008 filter
- June 2008 (1) Apply June 2008 filter
- May 2008 (3) Apply May 2008 filter
- April 2008 (1) Apply April 2008 filter
- March 2008 (5) Apply March 2008 filter
- February 2008 (5) Apply February 2008 filter
- January 2008 (6) Apply January 2008 filter
- Remove 2008 filter 2008
40 Janelia Publications
Showing 21-30 of 40 resultsThe identification of synaptic partners is challenging in dense nerve bundles, where many processes occupy regions beneath the resolution of conventional light microscopy. To address this difficulty, we have developed GRASP, a system to label membrane contacts and synapses between two cells in living animals. Two complementary fragments of GFP are expressed on different cells, tethered to extracellular domains of transmembrane carrier proteins. When the complementary GFP fragments are fused to ubiquitous transmembrane proteins, GFP fluorescence appears uniformly along membrane contacts between the two cells. When one or both GFP fragments are fused to synaptic transmembrane proteins, GFP fluorescence is tightly localized to synapses. GRASP marks known synaptic contacts in C. elegans, correctly identifies changes in mutants with altered synaptic specificity, and can uncover new information about synaptic locations as confirmed by electron microscopy. GRASP may prove particularly useful for defining connectivity in complex nervous systems.
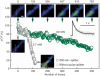
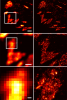
We combined photoactivated localization microscopy (PALM) with live-cell single-particle tracking to create a new method termed sptPALM. We created spatially resolved maps of single-molecule motions by imaging the membrane proteins Gag and VSVG, and obtained several orders of magnitude more trajectories per cell than traditional single-particle tracking enables. By probing distinct subsets of molecules, sptPALM can provide insight into the origins of spatial and temporal heterogeneities in membranes.
Commentary: As a stepping stone to true live cell PALM (see above), our collaborator Jennifer Lippincott-Schwartz suggested using the sparse photoactivation principle of PALM to track the nanoscale motion of thousands of individual molecules within a single living cell. Termed single particle tracking PALM (sptPALM), Jennifer’s postdocs Suliana Manley and Jen Gillette used the method in our PALM rig to create spatially resolved maps of diffusion rates in the plasma membrane of live cells. sptPALM is a powerful tool to study the active cytoskeletal or passive diffusional transport of individual molecules with far more measurements per cell than is possible without sparse photoactivation.
Pulsed lasers are key elements in nonlinear bioimaging techniques such as two-photon fluorescence excitation (TPE) microscopy. Typically, however, only a percent or less of the laser power available can be delivered to the sample before photoinduced damage becomes excessive. Here we describe a passive pulse splitter that converts each laser pulse into a fixed number of sub-pulses of equal energy. We applied the splitter to TPE imaging of fixed mouse brain slices labeled with GFP and show that, in different power regimes, the splitter can be used either to increase the signal rate more than 100-fold or to reduce the rate of photobleaching by over fourfold. In living specimens, the gains were even greater: a ninefold reduction in photobleaching during in vivo imaging of Caenorhabditis elegans larvae, and a six- to 20-fold decrease in the rate of photodamage during calcium imaging of rat hippocampal brain slices.
Pulsed lasers are key elements in nonlinear bioimaging techniques such as two-photon fluorescence excitation (TPE) microscopy. Typically, however, only a percent or less of the laser power available can be delivered to the sample before photoinduced damage becomes excessive. Here we describe a passive pulse splitter that converts each laser pulse into a fixed number of sub-pulses of equal energy. We applied the splitter to TPE imaging of fixed mouse brain slices labeled with GFP and show that, in different power regimes, the splitter can be used either to increase the signal rate more than 100-fold or to reduce the rate of photobleaching by over fourfold. In living specimens, the gains were even greater: a ninefold reduction in photobleaching during in vivo imaging of Caenorhabditis elegans larvae, and a six- to 20-fold decrease in the rate of photodamage during calcium imaging of rat hippocampal brain slices.
Commentary: Na Ji came to me early in her postdoc with an idea to reduce photodamage in nonlinear microscopy by splitting the pulses from an ultrafast laser into multiple subpulses of reduced energy. In six weeks, we constructed a prototype pulse splitter and obtained initial results confirming the validity of her vision. Further experiments with Jeff Magee demonstrated that the splitter could be used to increase imaging speed or reduce photodamage in two photon microscopy by one to two orders of magnitude. This project is a great example of how quickly one can react and exploit new ideas in the Janelia environment.
We demonstrate live-cell super-resolution imaging using photoactivated localization microscopy (PALM). The use of photon-tolerant cell lines in combination with the high resolution and molecular sensitivity of PALM permitted us to investigate the nanoscale dynamics within individual adhesion complexes (ACs) in living cells under physiological conditions for as long as 25 min, with half of the time spent collecting the PALM images at spatial resolutions down to approximately 60 nm and frame rates as short as 25 s. We visualized the formation of ACs and measured the fractional gain and loss of individual paxillin molecules as each AC evolved. By allowing observation of a wide variety of nanoscale dynamics, live-cell PALM provides insights into molecular assembly during the initiation, maturation and dissolution of cellular processes.
Commentary: The first example of true live cell and time lapse imaging by localization microscopy (as opposed to particle tracking), this paper uses the Nyquist criterion to establish a necessary condition for true spatial resolution based on the density of localized molecules – a condition often unmet in claims elsewhere in the superresolution literature.
By any method, higher spatiotemporal resolution requires increasing light exposure at the specimen, making noninvasive imaging increasingly difficult. Here, simultaneous differential interference contrast imaging is used to establish that cells behave physiologically before, during, and after PALM imaging. Similar controls are lacking from many supposed “live cell” superresolution demonstrations.
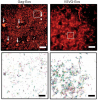
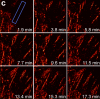
Recent advances in optical microscopy have enabled biological imaging beyond the diffraction limit at nanometer resolution. A general feature of most of the techniques based on photoactivated localization microscopy (PALM) or stochastic optical reconstruction microscopy (STORM) has been the use of thin biological samples in combination with total internal reflection, thus limiting the imaging depth to a fraction of an optical wavelength. However, to study whole cells or organelles that are typically up to 15 microm deep into the cell, the extension of these methods to a three-dimensional (3D) super resolution technique is required. Here, we report an advance in optical microscopy that enables imaging of protein distributions in cells with a lateral localization precision better than 50 nm at multiple imaging planes deep in biological samples. The approach is based on combining the lateral super resolution provided by PALM with two-photon temporal focusing that provides optical sectioning. We have generated super-resolution images over an axial range of approximately 10 microm in both mitochondrially labeled fixed cells, and in the membranes of living S2 Drosophila cells.
Mammalian mtDNA has been found here to harbor RNA-DNA hybrids at a variety of locations throughout the genome. The R-loop, previously characterized in vitro at the leading strand replication origin (OH), is isolated as a native RNA-DNA hybrid copurifying with mtDNA. Surprisingly, other mitochondrial transcripts also form stable partial R-loops. These are abundant and affect mtDNA conformation. Current models regarding the mechanism of mammalian mtDNA replication have been expanded by recent data and discordant hypotheses. The presence of stable, nonreplicative, and partially hybridized RNA on the mtDNA template is significant for the reevaluation of replication models based on two-dimensional agarose gel analyses. In addition, the close association of a subpopulation of mtRNA with the DNA template has further implications regarding the structure, maintenance, and expression of the mitochondrial genome. These results demonstrate that variously processed and targeted mtRNAs within mammalian mitochondria likely have multiple functions in addition to their conventional roles.
Key to understanding a protein’s biological function is the accurate determination of its spatial distribution inside a cell. Although fluorescent protein markers allow the targeting of specific proteins with molecular precision, much of this information is lost when the resultant fusion proteins are imaged with conventional, diffraction-limited optics. In response, several imaging modalities that are capable of resolution below the diffraction limit (approximately 200 nm) have emerged. Here, both single- and dual-color superresolution imaging of biological structures using photoactivated localization microscopy (PALM) are described. The examples discussed focus on adhesion complexes: dense, protein-filled assemblies that form at the interface between cells and their substrata. A particular emphasis is placed on the instrumentation and photoactivatable fluorescent protein (PA-FP) tags necessary to achieve PALM images at approximately 20 nm resolution in 5 to 30 min in fixed cells.
Commentary: A paper spearheaded by Hari which gives a thorough description of the methods and hardware needed to successfully practice PALM, including cover slip preparation, cell transfection and fixation, drift correction with fiducials, characterization of on/off contrast ratios for different photoactivted fluorescent proteins, identifying PALM-suitable cells, and mechanical and optical components of a PALM system.
A fundamental task in sequence analysis is to calculate the probability of a multiple alignment given a phylogenetic tree relating the sequences and an evolutionary model describing how sequences change over time. However, the most widely used phylogenetic models only account for residue substitution events. We describe a probabilistic model of a multiple sequence alignment that accounts for insertion and deletion events in addition to substitutions, given a phylogenetic tree, using a rate matrix augmented by the gap character. Starting from a continuous Markov process, we construct a non-reversible generative (birth-death) evolutionary model for insertions and deletions. The model assumes that insertion and deletion events occur one residue at a time. We apply this model to phylogenetic tree inference by extending the program dnaml in phylip. Using standard benchmarking methods on simulated data and a new "concordance test" benchmark on real ribosomal RNA alignments, we show that the extended program dnamlepsilon improves accuracy relative to the usual approach of ignoring gaps, while retaining the computational efficiency of the Felsenstein peeling algorithm.
Pyramidal neurons are characterized by their distinct apical and basal dendritic trees and the pyramidal shape of their soma. They are found in several regions of the CNS and, although the reasons for their abundance remain unclear, functional studies--especially of CA1 hippocampal and layer V neocortical pyramidal neurons--have offered insights into the functions of their unique cellular architecture. Pyramidal neurons are not all identical, but some shared functional principles can be identified. In particular, the existence of dendritic domains with distinct synaptic inputs, excitability, modulation and plasticity appears to be a common feature that allows synapses throughout the dendritic tree to contribute to action-potential generation. These properties support a variety of coincidence-detection mechanisms, which are likely to be crucial for synaptic integration and plasticity.