Filter
Associated Lab
- Ahrens Lab (3) Apply Ahrens Lab filter
- Aso Lab (6) Apply Aso Lab filter
- Baker Lab (2) Apply Baker Lab filter
- Betzig Lab (11) Apply Betzig Lab filter
- Branson Lab (6) Apply Branson Lab filter
- Cardona Lab (5) Apply Cardona Lab filter
- Chklovskii Lab (2) Apply Chklovskii Lab filter
- Cui Lab (5) Apply Cui Lab filter
- Dickson Lab (1) Apply Dickson Lab filter
- Druckmann Lab (3) Apply Druckmann Lab filter
- Dudman Lab (3) Apply Dudman Lab filter
- Eddy/Rivas Lab (4) Apply Eddy/Rivas Lab filter
- Egnor Lab (1) Apply Egnor Lab filter
- Fetter Lab (5) Apply Fetter Lab filter
- Freeman Lab (7) Apply Freeman Lab filter
- Funke Lab (1) Apply Funke Lab filter
- Gonen Lab (5) Apply Gonen Lab filter
- Grigorieff Lab (7) Apply Grigorieff Lab filter
- Harris Lab (7) Apply Harris Lab filter
- Heberlein Lab (1) Apply Heberlein Lab filter
- Hess Lab (7) Apply Hess Lab filter
- Jayaraman Lab (4) Apply Jayaraman Lab filter
- Ji Lab (4) Apply Ji Lab filter
- Karpova Lab (1) Apply Karpova Lab filter
- Keleman Lab (2) Apply Keleman Lab filter
- Keller Lab (7) Apply Keller Lab filter
- Lavis Lab (5) Apply Lavis Lab filter
- Leonardo Lab (2) Apply Leonardo Lab filter
- Liu (Zhe) Lab (4) Apply Liu (Zhe) Lab filter
- Looger Lab (9) Apply Looger Lab filter
- Magee Lab (5) Apply Magee Lab filter
- Murphy Lab (1) Apply Murphy Lab filter
- Pastalkova Lab (3) Apply Pastalkova Lab filter
- Reiser Lab (2) Apply Reiser Lab filter
- Romani Lab (2) Apply Romani Lab filter
- Rubin Lab (16) Apply Rubin Lab filter
- Saalfeld Lab (3) Apply Saalfeld Lab filter
- Scheffer Lab (2) Apply Scheffer Lab filter
- Schreiter Lab (4) Apply Schreiter Lab filter
- Shroff Lab (1) Apply Shroff Lab filter
- Simpson Lab (4) Apply Simpson Lab filter
- Singer Lab (10) Apply Singer Lab filter
- Spruston Lab (7) Apply Spruston Lab filter
- Stern Lab (4) Apply Stern Lab filter
- Sternson Lab (7) Apply Sternson Lab filter
- Svoboda Lab (9) Apply Svoboda Lab filter
- Tjian Lab (6) Apply Tjian Lab filter
- Truman Lab (6) Apply Truman Lab filter
- Turaga Lab (1) Apply Turaga Lab filter
- Turner Lab (2) Apply Turner Lab filter
- Wu Lab (1) Apply Wu Lab filter
- Zlatic Lab (4) Apply Zlatic Lab filter
- Zuker Lab (2) Apply Zuker Lab filter
Associated Project Team
Associated Support Team
- Electron Microscopy (1) Apply Electron Microscopy filter
- Fly Facility (1) Apply Fly Facility filter
- Gene Targeting and Transgenics (3) Apply Gene Targeting and Transgenics filter
- Janelia Experimental Technology (2) Apply Janelia Experimental Technology filter
- Management Team (1) Apply Management Team filter
- Molecular Genomics (1) Apply Molecular Genomics filter
- Primary & iPS Cell Culture (2) Apply Primary & iPS Cell Culture filter
- Project Technical Resources (1) Apply Project Technical Resources filter
- Scientific Computing Software (11) Apply Scientific Computing Software filter
Publication Date
- December 2015 (15) Apply December 2015 filter
- November 2015 (22) Apply November 2015 filter
- October 2015 (16) Apply October 2015 filter
- September 2015 (16) Apply September 2015 filter
- August 2015 (17) Apply August 2015 filter
- July 2015 (18) Apply July 2015 filter
- June 2015 (16) Apply June 2015 filter
- May 2015 (16) Apply May 2015 filter
- April 2015 (18) Apply April 2015 filter
- March 2015 (16) Apply March 2015 filter
- February 2015 (15) Apply February 2015 filter
- January 2015 (10) Apply January 2015 filter
- Remove 2015 filter 2015
195 Janelia Publications
Showing 61-70 of 195 resultsWe describe a general approach to designing two-dimensional (2D) protein arrays mediated by noncovalent protein-protein interfaces. Protein homo-oligomers are placed into one of the seventeen 2D layer groups, the degrees of freedom of the lattice are sampled to identify configurations with shape-complementary interacting surfaces, and the interaction energy is minimized using sequence design calculations. We used the method to design proteins that self-assemble into layer groups P 3 2 1, P 4 21 2, and P 6. Projection maps of micrometer-scale arrays, assembled both in vitro and in vivo, are consistent with the design models and display the target layer group symmetry. Such programmable 2D protein lattices should enable new approaches to structure determination, sensing, and nanomaterial engineering.
Calcium signaling has long been associated with key events of immunity, including chemotaxis, phagocytosis, and activation. However, imaging and manipulation of calcium flux in motile immune cells in live animals remain challenging. Using light-sheet microscopy for in vivo calcium imaging in zebrafish, we observe characteristic patterns of calcium flux triggered by distinct events, including phagocytosis of pathogenic bacteria and migration of neutrophils toward inflammatory stimuli. In contrast to findings from ex vivo studies, we observe enriched calcium influx at the leading edge of migrating neutrophils. To directly manipulate calcium dynamics in vivo, we have developed transgenic lines with cell-specific expression of the mammalian TRPV1 channel, enabling ligand-gated, reversible, and spatiotemporal control of calcium influx. We find that controlled calcium influx can function to help define the neutrophil's leading edge. Cell-specific TRPV1 expression may have broad utility for precise control of calcium dynamics in other immune cell types and organisms.
Adaptive optics by direct imaging of the wavefront distortions of a laser-induced guide star has long been used in astronomy, and more recently in microscopy to compensate for aberrations in transparent specimens. Here we extend this approach to tissues that strongly scatter visible light by exploiting the reduced scattering of near-infrared guide stars. The method enables in vivo two-photon morphological and functional imaging down to 700 μm inside the mouse brain.
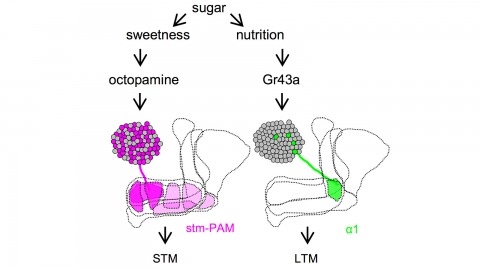
Drosophila melanogaster can acquire a stable appetitive olfactory memory when the presentation of a sugar reward and an odor are paired. However, the neuronal mechanisms by which a single training induces long-term memory are poorly understood. Here we show that two distinct subsets of dopamine neurons in the fly brain signal reward for short-term (STM) and long-term memories (LTM). One subset induces memory that decays within several hours, whereas the other induces memory that gradually develops after training. They convey reward signals to spatially segregated synaptic domains of the mushroom body (MB), a potential site for convergence. Furthermore, we identified a single type of dopamine neuron that conveys the reward signal to restricted subdomains of the mushroom body lobes and induces long-term memory. Constant appetitive memory retention after a single training session thus comprises two memory components triggered by distinct dopamine neurons.
The apical tuft is the most remote area of the dendritic tree of neocortical pyramidal neurons. Despite its distal location, the apical dendritic tuft of layer 5 pyramidal neurons receives substantial excitatory synaptic drive and actively processes corticocortical input during behavior. The properties of the voltage-activated ion channels that regulate synaptic integration in tuft dendrites have, however, not been thoroughly investigated. Here, we use electrophysiological and optical approaches to examine the subcellular distribution and function of hyperpolarization-activated cyclic nucleotide-gated nonselective cation (HCN) channels in rat layer 5B pyramidal neurons. Outside-out patch recordings demonstrated that the amplitude and properties of ensemble HCN channel activity were uniform in patches excised from distal apical dendritic trunk and tuft sites. Simultaneous apical dendritic tuft and trunk whole-cell current-clamp recordings revealed that the pharmacological blockade of HCN channels decreased voltage compartmentalization and enhanced the generation and spread of apical dendritic tuft and trunk regenerative activity. Furthermore, multisite two-photon glutamate uncaging demonstrated that HCN channels control the amplitude and duration of synaptically evoked regenerative activity in the distal apical dendritic tuft. In contrast, at proximal apical dendritic trunk and somatic recording sites, the blockade of HCN channels decreased excitability. Dynamic-clamp experiments revealed that these compartment-specific actions of HCN channels were heavily influenced by the local and distributed impact of the high density of HCN channels in the distal apical dendritic arbor. The properties and subcellular distribution pattern of HCN channels are therefore tuned to regulate the interaction between integration compartments in layer 5B pyramidal neurons.
Progressive depletion of midbrain dopamine neurons (PDD) is associated with deficits in the initiation, speed, and fluidity of voluntary movement. Models of basal ganglia function focus on initiation deficits; however, it is unclear how they account for deficits in the speed or amplitude of movement (vigor). Using an effort-based operant conditioning task for head-fixed mice, we discovered distinct functional classes of neurons in the dorsal striatum that represent movement vigor. Mice with PDD exhibited a progressive reduction in vigor, along with a selective impairment of its neural representation in striatum. Restoration of dopaminergic tone with a synthetic precursor ameliorated deficits in movement vigor and its neural representation, while suppression of striatal activity during movement was sufficient to reduce vigor. Thus, dopaminergic input to the dorsal striatum is indispensable for the emergence of striatal activity that mediates adaptive changes in movement vigor. These results suggest refined intervention strategies for Parkinson’s disease.
Germ granules, specialized ribonucleoprotein particles, are a hallmark of all germ cells. In Drosophila, an estimated 200 mRNAs are enriched in the germ plasm, and some of these have important, often conserved roles in germ cell formation, specification, survival and migration. How mRNAs are spatially distributed within a germ granule and whether their position defines functional properties is unclear. Here we show, using single-molecule FISH and structured illumination microscopy, a super-resolution approach, that mRNAs are spatially organized within the granule whereas core germ plasm proteins are distributed evenly throughout the granule. Multiple copies of single mRNAs organize into 'homotypic clusters' that occupy defined positions within the center or periphery of the granule. This organization, which is maintained during embryogenesis and independent of the translational or degradation activity of mRNAs, reveals new regulatory mechanisms for germ plasm mRNAs that may be applicable to other mRNA granules.
Early transplantation and grafting experiments suggest that body organs follow autonomous growth programs [1-3], therefore pointing to a need for coordination mechanisms to produce fit individuals with proper proportions. We recently identified Drosophila insulin-like peptide 8 (Dilp8) as a relaxin and insulin-like molecule secreted from growing tissues that plays a central role in coordinating growth between organs and coupling organ growth with animal maturation [4, 5]. Deciphering the function of Dilp8 in growth coordination relies on the identification of the receptor and tissues relaying Dilp8 signaling. We show here that the orphan receptor leucine-rich repeat-containing G protein-coupled receptor 3 (Lgr3), a member of the highly conserved family of relaxin family peptide receptors (RXFPs), mediates the checkpoint function of Dilp8 for entry into maturation. We functionally identify two Lgr3-positive neurons in each brain lobe that are required to induce a developmental delay upon overexpression of Dilp8. These neurons are located in the pars intercerebralis, an important neuroendocrine area in the brain, and make physical contacts with the PTTH neurons that ultimately control the production and release of the molting steroid ecdysone. Reducing Lgr3 levels in these neurons results in adult flies exhibiting increased fluctuating bilateral asymmetry, therefore recapitulating the phenotype of dilp8 mutants. Our work reveals a novel Dilp8/Lgr3 neuronal circuitry involved in a feedback mechanism that ensures coordination between organ growth and developmental transitions and prevents developmental variability.
Cortical cells integrate synaptic input from multiple sources, but how these different inputs are distributed across individual neurons is largely unknown. Differences in input might account for diverse responses in neighboring neurons during behavior. We present a strategy for comparing the strengths of multiple types of input onto the same neuron. We developed methods for independent dual-channel photostimulation of synaptic inputs using ChR2 together with ReaChR, a red-shifted channelrhodopsin. We used dual-channel photostimulation to probe convergence of sensory information in the mouse primary motor cortex. Input from somatosensory cortex and thalamus converges in individual neurons. Similarly, inputs from distinct somatotopic regions of the somatosensory cortex are integrated at the level of single motor cortex neurons. We next developed a ReaChR transgenic mouse under the control of both Flp- and Cre-recombinases that is an effective tool for circuit mapping. Our approach to dual-channel photostimulation enables quantitative comparison of the strengths of multiple pathways across all length scales of the brain.
Behavioral strategies employed for chemotaxis have been described across phyla, but the sensorimotor basis of this phenomenon has seldom been studied in naturalistic contexts. Here, we examine how signals experienced during free olfactory behaviors are processed by first-order olfactory sensory neurons (OSNs) of the Drosophila larva. We find that OSNs can act as differentiators that transiently normalize stimulus intensity-a property potentially derived from a combination of integral feedback and feed-forward regulation of olfactory transduction. In olfactory virtual reality experiments, we report that high activity levels of the OSN suppress turning, whereas low activity levels facilitate turning. Using a generalized linear model, we explain how peripheral encoding of olfactory stimuli modulates the probability of switching from a run to a turn. Our work clarifies the link between computations carried out at the sensory periphery and action selection underlying navigation in odor gradients.