Filter
Associated Lab
- Aguilera Castrejon Lab (1) Apply Aguilera Castrejon Lab filter
- Ahrens Lab (53) Apply Ahrens Lab filter
- Aso Lab (40) Apply Aso Lab filter
- Baker Lab (19) Apply Baker Lab filter
- Betzig Lab (101) Apply Betzig Lab filter
- Beyene Lab (8) Apply Beyene Lab filter
- Bock Lab (14) Apply Bock Lab filter
- Branson Lab (50) Apply Branson Lab filter
- Card Lab (36) Apply Card Lab filter
- Cardona Lab (45) Apply Cardona Lab filter
- Chklovskii Lab (10) Apply Chklovskii Lab filter
- Clapham Lab (14) Apply Clapham Lab filter
- Cui Lab (19) Apply Cui Lab filter
- Darshan Lab (8) Apply Darshan Lab filter
- Dickson Lab (32) Apply Dickson Lab filter
- Druckmann Lab (21) Apply Druckmann Lab filter
- Dudman Lab (38) Apply Dudman Lab filter
- Eddy/Rivas Lab (30) Apply Eddy/Rivas Lab filter
- Egnor Lab (4) Apply Egnor Lab filter
- Espinosa Medina Lab (15) Apply Espinosa Medina Lab filter
- Feliciano Lab (7) Apply Feliciano Lab filter
- Fetter Lab (31) Apply Fetter Lab filter
- Fitzgerald Lab (16) Apply Fitzgerald Lab filter
- Freeman Lab (15) Apply Freeman Lab filter
- Funke Lab (38) Apply Funke Lab filter
- Gonen Lab (59) Apply Gonen Lab filter
- Grigorieff Lab (34) Apply Grigorieff Lab filter
- Harris Lab (53) Apply Harris Lab filter
- Heberlein Lab (13) Apply Heberlein Lab filter
- Hermundstad Lab (23) Apply Hermundstad Lab filter
- Hess Lab (74) Apply Hess Lab filter
- Ilanges Lab (2) Apply Ilanges Lab filter
- Jayaraman Lab (42) Apply Jayaraman Lab filter
- Ji Lab (33) Apply Ji Lab filter
- Johnson Lab (1) Apply Johnson Lab filter
- Karpova Lab (13) Apply Karpova Lab filter
- Keleman Lab (8) Apply Keleman Lab filter
- Keller Lab (61) Apply Keller Lab filter
- Koay Lab (2) Apply Koay Lab filter
- Lavis Lab (137) Apply Lavis Lab filter
- Lee (Albert) Lab (29) Apply Lee (Albert) Lab filter
- Leonardo Lab (19) Apply Leonardo Lab filter
- Li Lab (4) Apply Li Lab filter
- Lippincott-Schwartz Lab (97) Apply Lippincott-Schwartz Lab filter
- Liu (Yin) Lab (1) Apply Liu (Yin) Lab filter
- Liu (Zhe) Lab (58) Apply Liu (Zhe) Lab filter
- Looger Lab (137) Apply Looger Lab filter
- Magee Lab (31) Apply Magee Lab filter
- Menon Lab (12) Apply Menon Lab filter
- Murphy Lab (6) Apply Murphy Lab filter
- O'Shea Lab (6) Apply O'Shea Lab filter
- Otopalik Lab (1) Apply Otopalik Lab filter
- Pachitariu Lab (36) Apply Pachitariu Lab filter
- Pastalkova Lab (5) Apply Pastalkova Lab filter
- Pavlopoulos Lab (7) Apply Pavlopoulos Lab filter
- Pedram Lab (4) Apply Pedram Lab filter
- Podgorski Lab (16) Apply Podgorski Lab filter
- Reiser Lab (45) Apply Reiser Lab filter
- Riddiford Lab (20) Apply Riddiford Lab filter
- Romani Lab (31) Apply Romani Lab filter
- Rubin Lab (105) Apply Rubin Lab filter
- Saalfeld Lab (46) Apply Saalfeld Lab filter
- Satou Lab (1) Apply Satou Lab filter
- Scheffer Lab (36) Apply Scheffer Lab filter
- Schreiter Lab (50) Apply Schreiter Lab filter
- Sgro Lab (1) Apply Sgro Lab filter
- Shroff Lab (31) Apply Shroff Lab filter
- Simpson Lab (18) Apply Simpson Lab filter
- Singer Lab (37) Apply Singer Lab filter
- Spruston Lab (57) Apply Spruston Lab filter
- Stern Lab (73) Apply Stern Lab filter
- Sternson Lab (47) Apply Sternson Lab filter
- Stringer Lab (32) Apply Stringer Lab filter
- Svoboda Lab (131) Apply Svoboda Lab filter
- Tebo Lab (9) Apply Tebo Lab filter
- Tervo Lab (9) Apply Tervo Lab filter
- Tillberg Lab (18) Apply Tillberg Lab filter
- Tjian Lab (17) Apply Tjian Lab filter
- Truman Lab (58) Apply Truman Lab filter
- Turaga Lab (39) Apply Turaga Lab filter
- Turner Lab (27) Apply Turner Lab filter
- Vale Lab (7) Apply Vale Lab filter
- Voigts Lab (3) Apply Voigts Lab filter
- Wang (Meng) Lab (21) Apply Wang (Meng) Lab filter
- Wang (Shaohe) Lab (6) Apply Wang (Shaohe) Lab filter
- Wu Lab (8) Apply Wu Lab filter
- Zlatic Lab (26) Apply Zlatic Lab filter
- Zuker Lab (5) Apply Zuker Lab filter
Associated Project Team
- CellMap (12) Apply CellMap filter
- COSEM (3) Apply COSEM filter
- FIB-SEM Technology (3) Apply FIB-SEM Technology filter
- Fly Descending Interneuron (11) Apply Fly Descending Interneuron filter
- Fly Functional Connectome (14) Apply Fly Functional Connectome filter
- Fly Olympiad (5) Apply Fly Olympiad filter
- FlyEM (53) Apply FlyEM filter
- FlyLight (49) Apply FlyLight filter
- GENIE (46) Apply GENIE filter
- Integrative Imaging (4) Apply Integrative Imaging filter
- Larval Olympiad (2) Apply Larval Olympiad filter
- MouseLight (18) Apply MouseLight filter
- NeuroSeq (1) Apply NeuroSeq filter
- ThalamoSeq (1) Apply ThalamoSeq filter
- Tool Translation Team (T3) (26) Apply Tool Translation Team (T3) filter
- Transcription Imaging (45) Apply Transcription Imaging filter
Associated Support Team
- Project Pipeline Support (5) Apply Project Pipeline Support filter
- Anatomy and Histology (18) Apply Anatomy and Histology filter
- Cryo-Electron Microscopy (36) Apply Cryo-Electron Microscopy filter
- Electron Microscopy (16) Apply Electron Microscopy filter
- Gene Targeting and Transgenics (11) Apply Gene Targeting and Transgenics filter
- Integrative Imaging (17) Apply Integrative Imaging filter
- Invertebrate Shared Resource (40) Apply Invertebrate Shared Resource filter
- Janelia Experimental Technology (37) Apply Janelia Experimental Technology filter
- Management Team (1) Apply Management Team filter
- Molecular Genomics (15) Apply Molecular Genomics filter
- Primary & iPS Cell Culture (14) Apply Primary & iPS Cell Culture filter
- Project Technical Resources (50) Apply Project Technical Resources filter
- Quantitative Genomics (19) Apply Quantitative Genomics filter
- Scientific Computing Software (92) Apply Scientific Computing Software filter
- Scientific Computing Systems (7) Apply Scientific Computing Systems filter
- Viral Tools (14) Apply Viral Tools filter
- Vivarium (7) Apply Vivarium filter
Publication Date
- 2025 (126) Apply 2025 filter
- 2024 (215) Apply 2024 filter
- 2023 (159) Apply 2023 filter
- 2022 (167) Apply 2022 filter
- 2021 (175) Apply 2021 filter
- 2020 (177) Apply 2020 filter
- 2019 (177) Apply 2019 filter
- 2018 (206) Apply 2018 filter
- 2017 (186) Apply 2017 filter
- 2016 (191) Apply 2016 filter
- 2015 (195) Apply 2015 filter
- 2014 (190) Apply 2014 filter
- 2013 (136) Apply 2013 filter
- 2012 (112) Apply 2012 filter
- 2011 (98) Apply 2011 filter
- 2010 (61) Apply 2010 filter
- 2009 (56) Apply 2009 filter
- 2008 (40) Apply 2008 filter
- 2007 (21) Apply 2007 filter
- 2006 (3) Apply 2006 filter
2691 Janelia Publications
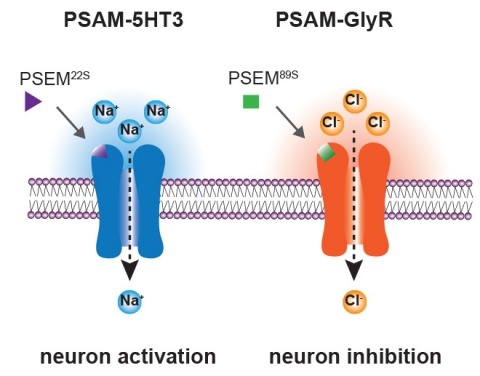
Showing 2441-2450 of 2691 resultsIonic flux mediates essential physiological and behavioral functions in defined cell populations. Cell type-specific activators of diverse ionic conductances are needed for probing these effects. We combined chemistry and protein engineering to enable the systematic creation of a toolbox of ligand-gated ion channels (LGICs) with orthogonal pharmacologic selectivity and divergent functional properties. The LGICs and their small-molecule effectors were able to activate a range of ionic conductances in genetically specified cell types. LGICs constructed for neuronal perturbation could be used to selectively manipulate neuron activity in mammalian brains in vivo. The diversity of ion channel tools accessible from this approach will be useful for examining the relationship between neuronal activity and animal behavior, as well as for cell biological and physiological applications requiring chemical control of ion conductance.
Concomitant with the publication of this Special Issue of Neuroinformatics, a substantially updated version of the DIADEM web site has been released at http://diademchallenge.org. This web site was originally designed to host the challenge for automating the digital reconstruction of axonal and dendritic morphology (hence the DIADEM acronym). This post-competition version features additional content for continued use as the access point for DIADEM-related material. From the very beginning, one of the spirits of DIADEM has been to share data and resources with the neuroscience research community at large. The resources available from or linked to the DIADEM website constitute a substantial scientific legacy of the 2009/2010 competition. The new content includes finalist algorithms, image stack data, gold standard reconstructions, an updated DIADEM metric, and a retrospective on the competition in text and images.
We took advantage of the unusual genomic organization of the ciliate Oxytricha trifallax to screen for eukaryotic non-coding RNA (ncRNA) genes. Ciliates have two types of nuclei: a germ line micronucleus that is usually transcriptionally inactive, and a somatic macronucleus that contains a reduced, fragmented and rearranged genome that expresses all genes required for growth and asexual reproduction. In some ciliates including Oxytricha, the macronuclear genome is particularly extreme, consisting of thousands of tiny ’nanochromosomes’, each of which usually contains only a single gene. Because the organism itself identifies and isolates most of its genes on single-gene nanochromosomes, nanochromosome structure could facilitate the discovery of unusual genes or gene classes, such as ncRNA genes. Using a draft Oxytricha genome assembly and a custom-written protein-coding genefinding program, we identified a subset of nanochromosomes that lack any detectable protein-coding gene, thereby strongly enriching for nanochromosomes that carry ncRNA genes. We found only a small proportion of non-coding nanochromosomes, suggesting that Oxytricha has few independent ncRNA genes besides homologs of already known RNAs. Other than new members of known ncRNA classes including C/D and H/ACA snoRNAs, our screen identified one new family of small RNA genes, named the Arisong RNAs, which share some of the features of small nuclear RNAs.
Because of its genetic, molecular, and behavioral tractability, Drosophila has emerged as a powerful model system for studying molecular and cellular mechanisms underlying the development and function of nervous systems. The Drosophila nervous system has fewer neurons and exhibits a lower glia:neuron ratio than is seen in vertebrate nervous systems. Despite the simplicity of the Drosophila nervous system, glial organization in flies is as sophisticated as it is in vertebrates. Furthermore, fly glial cells play vital roles in neural development and behavior. In addition, powerful genetic tools are continuously being created to explore cell function in vivo. In taking advantage of these features, the fly nervous system serves as an excellent model system to study general aspects of glial cell development and function in vivo. In this article, we review and discuss advanced genetic tools that are potentially useful for understanding glial cell biology in Drosophila.
Parallel circuits throughout the CNS exhibit distinct sensitivities and responses to sensory stimuli. Ambiguities in the source and properties of signals elicited by physiological stimuli, however, frequently obscure the mechanisms underlying these distinctions. We found that differences in the degree to which activity in two classes of Off retinal ganglion cell (RGC) encode information about light stimuli near detection threshold were not due to obvious differences in the cells’ intrinsic properties or the chemical synaptic input the cells received; indeed, differences in the cells’ light responses were largely insensitive to block of fast ionotropic glutamate receptors. Instead, the distinct responses of the two types of RGCs likely reflect differences in light-evoked electrical synaptic input. These results highlight a surprising strategy by which the retina differentially processes and routes visual information and provide new insight into the circuits that underlie responses to stimuli near detection threshold.
Site-specific recombinases have been used for two decades to manipulate the structure of animal genomes in highly predictable ways and have become major research tools. However, the small number of recombinases demonstrated to have distinct specificities, low toxicity, and sufficient activity to drive reactions to completion in animals has been a limitation. In this report we show that four recombinases derived from yeast-KD, B2, B3, and R-are highly active and nontoxic in Drosophila and that KD, B2, B3, and the widely used FLP recombinase have distinct target specificities. We also show that the KD and B3 recombinases are active in mice.
MOTIVATION: Automatic recognition of cell identities is critical for quantitative measurement, targeting, and manipulation of cells of model animals at single-cell resolution. It has been shown to be a powerful tool for studying gene expression and regulation, cell lineages, and cell fates. Existing methods first segment cells, before applying a recognition algorithm in the second step. As a result, the segmentation errors in the first step directly affect and complicate the subsequent cell recognition step. Moreover, in new experimental settings, some of the image features that have been previously relied upon to recognize cells may not be easy to reproduce, due to limitations on the number of color channels available for fluorescent imaging or to the cost of building transgenic animals. An approach that is more accurate and relies on only a single signal channel is clearly desirable. RESULTS: We have developed a new method, called SRS (for Simultaneous Recognition and Segmentation of cells), and applied it to 3D image stacks of the model organism C. elegans. Given a 3D image stack of the animal and a 3D atlas of target cells, SRS is effectively an atlas-guided voxel classification process: cell recognition is realized by smoothly deforming the atlas to best fit the image, where the segmentation is obtained naturally via classification of all image voxels. The method achieved a 97.7% overall recognition accuracy in recognizing a key class of marker cells, the body wall muscle (BWM) cells, on a data set of 175 C. elegans image stacks containing 14,118 manually curated BWM cells providing the "ground-truth" for accuracy. This result was achieved without any additional fiducial image features. SRS also automatically identified 14 of the image stacks as involving ±90-degree rotations. With these stacks excluded from the data set, the recognition accuracy rose to 99.1%. We also show SRS is generally applicable to other cell-types, e.g. intestinal cells. AVAILABILITY: The supplementary movies can be downloaded from our website http://penglab.janelia.org/proj/celegans_seganno. The method has been implemented as a plug-in program within the V3D system (http://penglab.janelia.org/proj/v3d) and will be released in the V3D plugin source code repository.
Pavlovian olfactory learning in Drosophila produces two genetically distinct forms of intermediate-term memories: anesthesia-sensitive memory, which requires the amnesiac gene, and anesthesia-resistant memory (ARM), which requires the radish gene. Here, we report that ARM is specifically enhanced or inhibited in flies with elevated or reduced serotonin (5HT) levels, respectively. The requirement for 5HT was additive with the memory defect of the amnesiac mutation but was occluded by the radish mutation. This result suggests that 5HT and Radish protein act on the same pathway for ARM formation. Three supporting lines of evidence indicate that ARM formation requires 5HT released from only two dorsal paired medial (DPM) neurons onto the mushroom bodies (MBs), the olfactory learning and memory center in Drosophila: (i) DPM neurons were 5HT-antibody immunopositive; (ii) temporal inhibition of 5HT synthesis or release from DPM neurons, but not from other serotonergic neurons, impaired ARM formation; (iii) knocking down the expression of d5HT1A serotonin receptors in α/β MB neurons, which are innervated by DPM neurons, inhibited ARM formation. Thus, in addition to the Amnesiac peptide required for anesthesia-sensitive memory formation, the two DPM neurons also release 5HT acting on MB neurons for ARM formation.