Filter
Associated Lab
- Ahrens Lab (3) Apply Ahrens Lab filter
- Aso Lab (3) Apply Aso Lab filter
- Baker Lab (5) Apply Baker Lab filter
- Betzig Lab (8) Apply Betzig Lab filter
- Branson Lab (6) Apply Branson Lab filter
- Card Lab (1) Apply Card Lab filter
- Cardona Lab (1) Apply Cardona Lab filter
- Chklovskii Lab (2) Apply Chklovskii Lab filter
- Cui Lab (2) Apply Cui Lab filter
- Darshan Lab (1) Apply Darshan Lab filter
- Dickson Lab (6) Apply Dickson Lab filter
- Druckmann Lab (1) Apply Druckmann Lab filter
- Dudman Lab (4) Apply Dudman Lab filter
- Eddy/Rivas Lab (3) Apply Eddy/Rivas Lab filter
- Egnor Lab (1) Apply Egnor Lab filter
- Fetter Lab (1) Apply Fetter Lab filter
- Fitzgerald Lab (1) Apply Fitzgerald Lab filter
- Freeman Lab (3) Apply Freeman Lab filter
- Gonen Lab (10) Apply Gonen Lab filter
- Grigorieff Lab (3) Apply Grigorieff Lab filter
- Harris Lab (2) Apply Harris Lab filter
- Heberlein Lab (2) Apply Heberlein Lab filter
- Hermundstad Lab (2) Apply Hermundstad Lab filter
- Hess Lab (5) Apply Hess Lab filter
- Jayaraman Lab (1) Apply Jayaraman Lab filter
- Ji Lab (3) Apply Ji Lab filter
- Johnson Lab (1) Apply Johnson Lab filter
- Kainmueller Lab (2) Apply Kainmueller Lab filter
- Karpova Lab (1) Apply Karpova Lab filter
- Keller Lab (8) Apply Keller Lab filter
- Lavis Lab (7) Apply Lavis Lab filter
- Lee (Albert) Lab (4) Apply Lee (Albert) Lab filter
- Leonardo Lab (4) Apply Leonardo Lab filter
- Li Lab (1) Apply Li Lab filter
- Lippincott-Schwartz Lab (12) Apply Lippincott-Schwartz Lab filter
- Liu (Zhe) Lab (4) Apply Liu (Zhe) Lab filter
- Looger Lab (11) Apply Looger Lab filter
- Magee Lab (1) Apply Magee Lab filter
- Menon Lab (3) Apply Menon Lab filter
- Murphy Lab (1) Apply Murphy Lab filter
- Pavlopoulos Lab (2) Apply Pavlopoulos Lab filter
- Reiser Lab (2) Apply Reiser Lab filter
- Riddiford Lab (4) Apply Riddiford Lab filter
- Romani Lab (2) Apply Romani Lab filter
- Rubin Lab (9) Apply Rubin Lab filter
- Saalfeld Lab (1) Apply Saalfeld Lab filter
- Scheffer Lab (7) Apply Scheffer Lab filter
- Sgro Lab (1) Apply Sgro Lab filter
- Simpson Lab (2) Apply Simpson Lab filter
- Singer Lab (10) Apply Singer Lab filter
- Spruston Lab (1) Apply Spruston Lab filter
- Stern Lab (6) Apply Stern Lab filter
- Sternson Lab (5) Apply Sternson Lab filter
- Stringer Lab (1) Apply Stringer Lab filter
- Svoboda Lab (7) Apply Svoboda Lab filter
- Tebo Lab (2) Apply Tebo Lab filter
- Tervo Lab (1) Apply Tervo Lab filter
- Tillberg Lab (1) Apply Tillberg Lab filter
- Tjian Lab (4) Apply Tjian Lab filter
- Truman Lab (1) Apply Truman Lab filter
- Turaga Lab (1) Apply Turaga Lab filter
- Turner Lab (1) Apply Turner Lab filter
- Wang (Shaohe) Lab (1) Apply Wang (Shaohe) Lab filter
- Wu Lab (2) Apply Wu Lab filter
- Zlatic Lab (1) Apply Zlatic Lab filter
- Zuker Lab (1) Apply Zuker Lab filter
Associated Project Team
Publication Date
- December 2014 (35) Apply December 2014 filter
- November 2014 (14) Apply November 2014 filter
- October 2014 (15) Apply October 2014 filter
- September 2014 (17) Apply September 2014 filter
- August 2014 (14) Apply August 2014 filter
- July 2014 (26) Apply July 2014 filter
- June 2014 (14) Apply June 2014 filter
- May 2014 (14) Apply May 2014 filter
- April 2014 (20) Apply April 2014 filter
- March 2014 (18) Apply March 2014 filter
- February 2014 (15) Apply February 2014 filter
- January 2014 (34) Apply January 2014 filter
- Remove 2014 filter 2014
Type of Publication
236 Publications
Showing 191-200 of 236 resultsDespite the importance of the insect nervous system for functional and developmental neuroscience, descriptions of insect brains have suffered from a lack of uniform nomenclature. Ambiguous definitions of brain regions and fiber bundles have contributed to the variation of names used to describe the same structure. The lack of clearly determined neuropil boundaries has made it difficult to document precise locations of neuronal projections for connectomics study. To address such issues, a consortium of neurobiologists studying arthropod brains, the Insect Brain Name Working Group, has established the present hierarchical nomenclature system, using the brain of Drosophila melanogaster as the reference framework, while taking the brains of other taxa into careful consideration for maximum consistency and expandability. The following summarizes the consortium’s nomenclature system and highlights examples of existing ambiguities and remedies for them. This nomenclature is intended to serve as a standard of reference for the study of the brain of Drosophila and other insects.
BACKGROUND: Detecting the direction of visual motion is an essential task of the early visual system. The Reichardt detector has been proven to be a faithful description of the underlying computation in insects. A series of recent studies addressed the neural implementation of the Reichardt detector in Drosophila revealing the overall layout in parallel ON and OFF channels, its input neurons from the lamina (L1→ON, and L2→OFF), and the respective output neurons to the lobula plate (ON→T4, and OFF→T5). While anatomical studies showed that T4 cells receive input from L1 via Mi1 and Tm3 cells, the neurons connecting L2 to T5 cells have not been identified so far. It is, however, known that L2 contacts, among others, two neurons, called Tm2 and L4, which show a pronounced directionality in their wiring. RESULTS: We characterized the visual response properties of both Tm2 and L4 neurons via Ca(2+) imaging. We found that Tm2 and L4 cells respond with an increase in activity to moving OFF edges in a direction-unselective manner. To investigate their participation in motion vision, we blocked their output while recording from downstream tangential cells in the lobula plate. Silencing of Tm2 and L4 completely abolishes the response to moving OFF edges. CONCLUSIONS: Our results demonstrate that both cell types are essential components of the Drosophila OFF motion vision pathway, prior to the computation of directionality in the dendrites of T5 cells.
A number of recent studies have provided compelling demonstrations that both mice and rats can be trained to perform a variety of behavioral tasks while restrained by mechanical elements mounted to the skull. The independent development of this technique by a number of laboratories has led to diverse solutions. We found that these solutions often used expensive materials and impeded future development and modification in the absence of engineering support. In order to address these issues, here we report on the development of a flexible single hardware design for electrophysiology and imaging both in brain tissue in vitro. Our hardware facilitates the rapid conversion of a single preparation between physiology and imaging system and the conversion of a given system between preparations. In addition, our use of rapid prototyping machines ("3D printers") allows for the deployment of new designs within a day. Here, we present specifications for design and manufacturing as well as some data from our lab demonstrating the suitability of the design for physiology in behaving animals and imaging in vitro and in vivo.
Development of multicellular organisms depends on patterning and growth mechanisms encoded in the genome, but also on the physical properties and mechanical interactions of the constituent cells that interpret these genetic cues. This fundamental biological problem requires integrated studies at multiple levels of biological organization: from genes, to cell behaviors, to tissue morphogenesis. We have recently combined functional genetics with live imaging approaches in embryos of the insect Tribolium castaneum, in order to understand their remarkable transformation from a uniform single-layered blastoderm into a condensed multi-layered embryo covered by extensive extra-embryonic tissues. We first developed a quick and reliable methodology to fluorescently label various cell components in entire Tribolium embryos. Live imaging of labeled embryos at single cell resolution provided detailed descriptions of cell behaviors and tissue movements during normal embryogenesis. We then compared cell and tissue dynamics between wild-type and genetically perturbed embryos that exhibited altered relative proportions of constituent tissues. This systematic comparison led to a qualitative model of the molecular, cellular and tissue interactions that orchestrate the observed epithelial rearrangements. We expect this work to establish the Tribolium embryo as a powerful and attractive model system for biologists and biophysicists interested in the molecular, cellular and mechanical control of tissue morphogenesis.
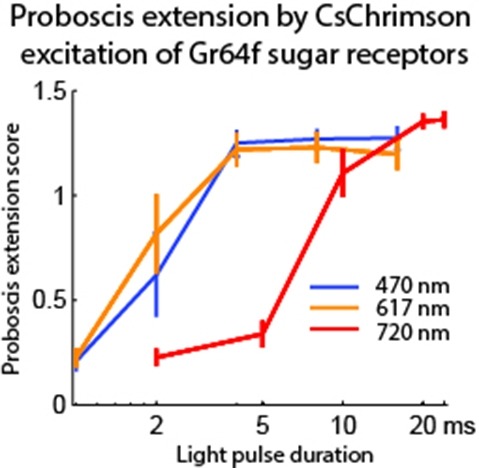
Optogenetic tools enable examination of how specific cell types contribute to brain circuit functions. A long-standing question is whether it is possible to independently activate two distinct neural populations in mammalian brain tissue. Such a capability would enable the study of how different synapses or pathways interact to encode information in the brain. Here we describe two channelrhodopsins, Chronos and Chrimson, discovered through sequencing and physiological characterization of opsins from over 100 species of alga. Chrimson’s excitation spectrum is red shifted by 45 nm relative to previous channelrhodopsins and can enable experiments in which red light is preferred. We show minimal visual system-mediated behavioral interference when using Chrimson in neurobehavioral studies in Drosophila melanogaster. Chronos has faster kinetics than previous channelrhodopsins yet is effectively more light sensitive. Together these two reagents enable two-color activation of neural spiking and downstream synaptic transmission in independent neural populations without detectable cross-talk in mouse brain slice.
The human immunodeficiency virus (HIV) hijacks the endosomal sorting complexes required for transport (ESCRT) to mediate virus release from infected cells. The nanoscale organization of ESCRT machinery necessary for mediating viral abscission is unclear. Here, we applied three-dimensional superresolution microscopy and correlative electron microscopy to delineate the organization of ESCRT components at HIV assembly sites. We observed ESCRT subunits localized within the head of budding virions and released particles, with head-localized levels of CHMP2A decreasing relative to Tsg101 and CHMP4B upon virus abscission. Thus, the driving force for HIV release may derive from initial scaffolding of ESCRT subunits within the viral bud interior followed by plasma membrane association and selective remodeling of ESCRT subunits.
We describe a method for fully automated detection of chemical synapses in serial electron microscopy images with highly anisotropic axial and lateral resolution, such as images taken on transmission electron microscopes. Our pipeline starts from classification of the pixels based on 3D pixel features, which is followed by segmentation with an Ising model MRF and another classification step, based on object-level features. Classifiers are learned on sparse user labels; a fully annotated data subvolume is not required for training. The algorithm was validated on a set of 238 synapses in 20 serial 7197×7351 pixel images (4.5×4.5×45 nm resolution) of mouse visual cortex, manually labeled by three independent human annotators and additionally re-verified by an expert neuroscientist. The error rate of the algorithm (12% false negative, 7% false positive detections) is better than state-of-the-art, even though, unlike the state-of-the-art method, our algorithm does not require a prior segmentation of the image volume into cells. The software is based on the ilastik learning and segmentation toolkit and the vigra image processing library and is freely available on our website, along with the test data and gold standard annotations (http://www.ilastik.org/synapse-detection/sstem).
View Publication PageHistone variant H2A.Z-containing nucleosomes exist at most eukaryotic promoters and play important roles in gene transcription and genome stability. The multisubunit nucleosome-remodeling enzyme complex SWR1, conserved from yeast to mammals, catalyzes the ATP-dependent replacement of histone H2A in canonical nucleosomes with H2A.Z. How SWR1 catalyzes the replacement reaction is largely unknown. Here, we determined the crystal structure of the N-terminal region (599-627) of the catalytic subunit Swr1, termed Swr1-Z domain, in complex with the H2A.Z-H2B dimer at 1.78 Å resolution. The Swr1-Z domain forms a 310 helix and an irregular chain. A conserved LxxLF motif in the Swr1-Z 310 helix specifically recognizes the αC helix of H2A.Z. Our results show that the Swr1-Z domain can deliver the H2A.Z-H2B dimer to the DNA-(H3-H4)2 tetrasome to form the nucleosome by a histone chaperone mechanism.
The organization of synaptic connectivity within a neuronal circuit is a prime determinant of circuit function. We performed a comprehensive fine-scale circuit mapping of hippocampal regions (CA3-CA1) using the newly developed synapse labeling method, mGRASP. This mapping revealed spatially nonuniform and clustered synaptic connectivity patterns. Furthermore, synaptic clustering was enhanced between groups of neurons that shared a similar developmental/migration time window, suggesting a mechanism for establishing the spatial structure of synaptic connectivity. Such connectivity patterns are thought to effectively engage active dendritic processing and storage mechanisms, thereby potentially enhancing neuronal feature selectivity.
BACKGROUND: Male-specific products of the fruitless (fru) gene control the development and function of neuronal circuits that underlie male-specific behaviors in Drosophila, including courtship. Alternative splicing generates at least three distinct Fru isoforms, each containing a different zinc-finger domain. Here, we examine the expression and function of each of these isoforms. RESULTS: We show that most fru(+) cells express all three isoforms, yet each isoform has a distinct function in the elaboration of sexually dimorphic circuitry and behavior. The strongest impairment in courtship behavior is observed in fru(C) mutants, which fail to copulate, lack sine song, and do not generate courtship song in the absence of visual stimuli. Cellular dimorphisms in the fru circuit are dependent on Fru(C) rather than other single Fru isoforms. Removal of Fru(C) from the neuronal classes vAB3 or aSP4 leads to cell-autonomous feminization of arborizations and loss of courtship in the dark. CONCLUSIONS: These data map specific aspects of courtship behavior to the level of single fru isoforms and fru(+) cell types-an important step toward elucidating the chain of causality from gene to circuit to behavior.