Filter
Associated Lab
- Ahrens Lab (1) Apply Ahrens Lab filter
- Aso Lab (1) Apply Aso Lab filter
- Baker Lab (3) Apply Baker Lab filter
- Betzig Lab (7) Apply Betzig Lab filter
- Bock Lab (2) Apply Bock Lab filter
- Branson Lab (1) Apply Branson Lab filter
- Cardona Lab (1) Apply Cardona Lab filter
- Cui Lab (3) Apply Cui Lab filter
- Dickson Lab (2) Apply Dickson Lab filter
- Druckmann Lab (1) Apply Druckmann Lab filter
- Dudman Lab (1) Apply Dudman Lab filter
- Eddy/Rivas Lab (5) Apply Eddy/Rivas Lab filter
- Fetter Lab (4) Apply Fetter Lab filter
- Fitzgerald Lab (1) Apply Fitzgerald Lab filter
- Gonen Lab (5) Apply Gonen Lab filter
- Grigorieff Lab (6) Apply Grigorieff Lab filter
- Heberlein Lab (12) Apply Heberlein Lab filter
- Hermundstad Lab (1) Apply Hermundstad Lab filter
- Hess Lab (2) Apply Hess Lab filter
- Jayaraman Lab (2) Apply Jayaraman Lab filter
- Ji Lab (2) Apply Ji Lab filter
- Kainmueller Lab (1) Apply Kainmueller Lab filter
- Keller Lab (4) Apply Keller Lab filter
- Lavis Lab (5) Apply Lavis Lab filter
- Lee (Albert) Lab (1) Apply Lee (Albert) Lab filter
- Leonardo Lab (2) Apply Leonardo Lab filter
- Lippincott-Schwartz Lab (18) Apply Lippincott-Schwartz Lab filter
- Liu (Zhe) Lab (1) Apply Liu (Zhe) Lab filter
- Looger Lab (7) Apply Looger Lab filter
- Magee Lab (1) Apply Magee Lab filter
- Menon Lab (3) Apply Menon Lab filter
- Murphy Lab (1) Apply Murphy Lab filter
- Pastalkova Lab (1) Apply Pastalkova Lab filter
- Pavlopoulos Lab (2) Apply Pavlopoulos Lab filter
- Reiser Lab (2) Apply Reiser Lab filter
- Riddiford Lab (1) Apply Riddiford Lab filter
- Romani Lab (1) Apply Romani Lab filter
- Rubin Lab (4) Apply Rubin Lab filter
- Satou Lab (3) Apply Satou Lab filter
- Scheffer Lab (2) Apply Scheffer Lab filter
- Schreiter Lab (3) Apply Schreiter Lab filter
- Sgro Lab (2) Apply Sgro Lab filter
- Simpson Lab (3) Apply Simpson Lab filter
- Singer Lab (10) Apply Singer Lab filter
- Spruston Lab (1) Apply Spruston Lab filter
- Stern Lab (4) Apply Stern Lab filter
- Sternson Lab (6) Apply Sternson Lab filter
- Svoboda Lab (7) Apply Svoboda Lab filter
- Tjian Lab (4) Apply Tjian Lab filter
- Truman Lab (1) Apply Truman Lab filter
- Turaga Lab (1) Apply Turaga Lab filter
- Turner Lab (2) Apply Turner Lab filter
- Zlatic Lab (1) Apply Zlatic Lab filter
- Zuker Lab (3) Apply Zuker Lab filter
Associated Project Team
Publication Date
- December 2011 (22) Apply December 2011 filter
- November 2011 (15) Apply November 2011 filter
- October 2011 (14) Apply October 2011 filter
- September 2011 (17) Apply September 2011 filter
- August 2011 (14) Apply August 2011 filter
- July 2011 (10) Apply July 2011 filter
- June 2011 (17) Apply June 2011 filter
- May 2011 (13) Apply May 2011 filter
- April 2011 (11) Apply April 2011 filter
- March 2011 (14) Apply March 2011 filter
- February 2011 (16) Apply February 2011 filter
- January 2011 (27) Apply January 2011 filter
- Remove 2011 filter 2011
Type of Publication
190 Publications
Showing 41-50 of 190 resultsHippocampal neurons can display reliable and long-lasting sequences of transient firing patterns, even in the absence of changing external stimuli. We suggest that time-keeping is an important function of these sequences, and propose a network mechanism for their generation. We show that sequences of neuronal assemblies recorded from rat hippocampal CA1 pyramidal cells can reliably predict elapsed time (15-20 s) during wheel running with a precision of 0.5 s. In addition, we demonstrate the generation of multiple reliable, long-lasting sequences in a recurrent network model. These sequences are generated in the presence of noisy, unstructured inputs to the network, mimicking stationary sensory input. Identical initial conditions generate similar sequences, whereas different initial conditions give rise to distinct sequences. The key ingredients responsible for sequence generation in the model are threshold-adaptation and a Mexican-hat-like pattern of connectivity among pyramidal cells. This pattern may arise from recurrent systems such as the hippocampal CA3 region or the entorhinal cortex. We hypothesize that mechanisms that evolved for spatial navigation also support tracking of elapsed time in behaviorally relevant contexts.
We recently discovered that the major mammalian bile acid, taurocholate, accelerated polarity in primary rat hepatocytes. Taurocholate increased cellular cAMP and signals through an Epac-Rap1-MEK-LKB1-AMPK pathway for its polarity effect. This review discusses possible mechanisms for how taurocholate affects different cell polarity factors, particularly AMPK, and thereby regulates events that generate polarity. These include tight junction formation, apical trafficking, recycling endosome dynamics, and cytoskeleton rearrangement. We also discuss whether the effects of taurocholate are mediated by other LKB1 downstream kinases, such as Par1 and NUAK1.
Sensory stimuli are represented in the brain by the activity of populations of neurons. In most biological systems, studying population coding is challenging since only a tiny proportion of cells can be recorded simultaneously. Here we used two-photon imaging to record neural activity in the relatively simple Drosophila mushroom body (MB), an area involved in olfactory learning and memory. Using the highly sensitive calcium indicator GCaMP3, we simultaneously monitored the activity of >100 MB neurons in vivo (∼5% of the total population). The MB is thought to encode odors in sparse patterns of activity, but the code has yet to be explored either on a population level or with a wide variety of stimuli. We therefore imaged responses to odors chosen to evaluate the robustness of sparse representations. Different odors activated distinct patterns of MB neurons; however, we found no evidence for spatial organization of neurons by either response probability or odor tuning within the cell body layer. The degree of sparseness was consistent across a wide range of stimuli, from monomolecular odors to artificial blends and even complex natural smells. Sparseness was mainly invariant across concentrations, largely because of the influence of recent odor experience. Finally, in contrast to sensory processing in other systems, no response features distinguished natural stimuli from monomolecular odors. Our results indicate that the fundamental feature of odor processing in the MB is to create sparse stimulus representations in a format that facilitates arbitrary associations between odor and punishment or reward.
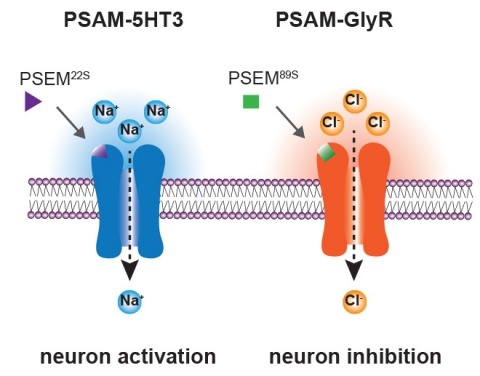
Ionic flux mediates essential physiological and behavioral functions in defined cell populations. Cell type-specific activators of diverse ionic conductances are needed for probing these effects. We combined chemistry and protein engineering to enable the systematic creation of a toolbox of ligand-gated ion channels (LGICs) with orthogonal pharmacologic selectivity and divergent functional properties. The LGICs and their small-molecule effectors were able to activate a range of ionic conductances in genetically specified cell types. LGICs constructed for neuronal perturbation could be used to selectively manipulate neuron activity in mammalian brains in vivo. The diversity of ion channel tools accessible from this approach will be useful for examining the relationship between neuronal activity and animal behavior, as well as for cell biological and physiological applications requiring chemical control of ion conductance.
Establishing visual correspondences is a critical step in many computer vision tasks involving multiple views of a scene. In a dynamic environment and when cameras are mobile, visual correspondences need to be updated on a recurring basis. At the same time, the use of wireless links between camera motes imposes tight rate constraints. This combination of issues motivates us to consider the problem of establishing visual correspondences in a distributed fashion between cameras operating under rate constraints. We propose a solution based on constructing distance preserving hashes using binarized random projections. By exploiting the fact that descriptors of regions in correspondence are highly correlated, we propose a novel use of distributed source coding via linear codes on the binary hashes to more efficiently exchange feature descriptors for establishing correspondences across multiple camera views. A systematic approach is used to evaluate rate vs visual correspondences retrieval performance; under a stringent matching criterion, our proposed methods demonstrate superior performance to a baseline scheme employing transform coding of descriptors.
Bilateral symmetric tissues must interpret axial references to maintain their global architecture during growth or repair. The regeneration of hair cells in the zebrafish lateral line, for example, forms a vertical midline that bisects the neuromast epithelium into perfect mirror-symmetric plane-polarized halves. Each half contains hair cells of identical planar orientation but opposite to that of the confronting half. The establishment of bilateral symmetry in this organ is poorly understood. Here, we show that hair-cell regeneration is strongly directional along an axis perpendicular to that of epithelial planar polarity. We demonstrate compartmentalized Notch signaling in neuromasts, and show that directional regeneration depends on the development of hair-cell progenitors in polar compartments that have low Notch activity. High-resolution live cell tracking reveals a novel process of planar cell inversions whereby sibling hair cells invert positions immediately after progenitor cytokinesis, demonstrating that oriented progenitor divisions are dispensable for bilateral symmetry. Notwithstanding the invariably directional regeneration, the planar polarization of the epithelium eventually propagates symmetrically because mature hair cells move away from the midline towards the periphery of the neuromast. We conclude that a strongly anisotropic regeneration process that relies on the dynamic stabilization of progenitor identity in permissive polar compartments sustains bilateral symmetry in the lateral line.
Deciphering the molecular basis of pluripotency is fundamental to our understanding of development and embryonic stem cell function. Here, we report that TAF3, a TBP-associated core promoter factor, is highly enriched in ES cells. In this context, TAF3 is required for endoderm lineage differentiation and prevents premature specification of neuroectoderm and mesoderm. In addition to its role in the core promoter recognition complex TFIID, genome-wide binding studies reveal that TAF3 localizes to a subset of chromosomal regions bound by CTCF/cohesin that are selectively associated with genes upregulated by TAF3. Notably, CTCF directly recruits TAF3 to promoter distal sites and TAF3-dependent DNA looping is observed between the promoter distal sites and core promoters occupied by TAF3/CTCF/cohesin. Together, our findings support a new role of TAF3 in mediating long-range chromatin regulatory interactions that safeguard the finely-balanced transcriptional programs underlying pluripotency.
Recent studies of several key developmental transitions have brought into question the long held view of the basal transcriptional apparatus as ubiquitous and invariant. In an effort to better understand the role of core promoter recognition and coactivator complex switching in cellular differentiation, we have examined changes in transcription factor IID (TFIID) and cofactor required for Sp1 activation/Mediator during mouse liver development. Here we show that the differentiation of fetal liver progenitors to adult hepatocytes involves a wholesale depletion of canonical cofactor required for Sp1 activation/Mediator and TFIID complexes at both the RNA and protein level, and that this alteration likely involves silencing of transcription factor promoters as well as protein degradation. It will be intriguing for future studies to determine if a novel and as yet unknown core promoter recognition complex takes the place of TFIID in adult hepatocytes and to uncover the mechanisms that down-regulate TFIID during this critical developmental transition.
Aberrant mRNAs with premature translation termination codons (PTCs) are recognized and eliminated by the nonsense-mediated mRNA decay (NMD) pathway in eukaryotes. We employed a novel live-cell imaging approach to investigate the kinetics of mRNA synthesis and release at the transcription site of PTC-containing (PTC+) and PTC-free (PTC-) immunoglobulin-μ reporter genes. Fluorescence recovery after photobleaching (FRAP) and photoconversion analyses revealed that PTC+ transcripts are specifically retained at the transcription site. Remarkably, the retained PTC+ transcripts are mainly unspliced, and this RNA retention is dependent upon two important NMD factors, UPF1 and SMG6, since their depletion led to the release of the PTC+ transcripts. Finally, ChIP analysis showed a physical association of UPF1 and SMG6 with both the PTC+ and the PTC- reporter genes in vivo. Collectively, our data support a mechanism for regulation of PTC+ transcripts at the transcription site.