Filter
Associated Lab
- Card Lab (3) Apply Card Lab filter
- Dickson Lab (5) Apply Dickson Lab filter
- Funke Lab (2) Apply Funke Lab filter
- Lavis Lab (2) Apply Lavis Lab filter
- Reiser Lab (1) Apply Reiser Lab filter
- Rubin Lab (1) Apply Rubin Lab filter
- Singer Lab (2) Apply Singer Lab filter
- Stern Lab (160) Apply Stern Lab filter
- Tillberg Lab (2) Apply Tillberg Lab filter
- Truman Lab (4) Apply Truman Lab filter
Associated Project Team
Publication Date
- 2026 (3) Apply 2026 filter
- 2025 (6) Apply 2025 filter
- 2024 (9) Apply 2024 filter
- 2023 (2) Apply 2023 filter
- 2022 (7) Apply 2022 filter
- 2021 (5) Apply 2021 filter
- 2020 (4) Apply 2020 filter
- 2019 (5) Apply 2019 filter
- 2018 (6) Apply 2018 filter
- 2017 (8) Apply 2017 filter
- 2016 (7) Apply 2016 filter
- 2015 (4) Apply 2015 filter
- 2014 (6) Apply 2014 filter
- 2013 (6) Apply 2013 filter
- 2012 (5) Apply 2012 filter
- 2011 (4) Apply 2011 filter
- 2010 (8) Apply 2010 filter
- 2009 (5) Apply 2009 filter
- 2008 (4) Apply 2008 filter
- 2007 (9) Apply 2007 filter
- 2006 (6) Apply 2006 filter
- 2005 (6) Apply 2005 filter
- 2004 (3) Apply 2004 filter
- 2003 (8) Apply 2003 filter
- 2001 (1) Apply 2001 filter
- 2000 (4) Apply 2000 filter
- 1999 (2) Apply 1999 filter
- 1998 (2) Apply 1998 filter
- 1997 (3) Apply 1997 filter
- 1996 (3) Apply 1996 filter
- 1995 (2) Apply 1995 filter
- 1994 (2) Apply 1994 filter
- 1993 (1) Apply 1993 filter
- 1991 (3) Apply 1991 filter
- 1990 (1) Apply 1990 filter
Type of Publication
160 Publications
Showing 131-140 of 160 resultsInsect dispersal dimorphisms, in which both flight-capable and flightless individuals occur in the same species, are thought to reflect a balance between the benefits and costs of dispersal. Fitness costs and benefits associated with wing dimorphism were investigated in the male pea aphid, Acyrthosiphon pisum (Harris) (Hemiptera: Aphididae). In one-on-one mating competitions in small arenas between winged and wingless males, the winged aphids obtained most of the matings with virgin females. In contrast, during competition experiments in larger cages with multiple individuals of each morph, the winged males no longer had a clear mating advantage over wingless males. In the absence of competition, wingless males had marginally higher lifetime reproductive success than winged males, probably because mating winged males tended to die faster than wingless males. In the absence of females, winged males survived longer than wingless males and this difference disappeared under starvation conditions. Mating males of both morphs died significantly faster than males without access to females. There does not appear to be a direct tradeoff of dispersal ability with life history characteristics in pea aphid males, suggesting that the advantages of producing winged males may result from outbreeding.
The pea aphid, Acyrthosiphon pisum, is an emerging genomic model system for studies of polyphenisms, bacterial symbioses, host-plant specialization, and the vectoring of plant viruses. Here we provide estimates of nucleotide diversity and linkage disequilibrium (LD) in native (European) and introduced (United States) populations of the pea aphid. Because introductions can cause population bottlenecks, we hypothesized that U.S. populations harbor lower levels of nucleotide diversity and higher levels of LD than native populations.
To perform most behaviors, animals must send commands from higher-order processing centers in the brain to premotor circuits that reside in ganglia distinct from the brain, such as the mammalian spinal cord or insect ventral nerve cord. How these circuits are functionally organized to generate the great diversity of animal behavior remains unclear. An important first step in unraveling the organization of premotor circuits is to identify their constituent cell types and create tools to monitor and manipulate these with high specificity to assess their functions. This is possible in the tractable ventral nerve cord of the fly. To generate such a toolkit, we used a combinatorial genetic technique (split-GAL4) to create 195 sparse transgenic driver lines targeting 196 individual cell types in the ventral nerve cord. These included wing and haltere motoneurons, modulatory neurons, and interneurons. Using a combination of behavioral, developmental, and anatomical analyses, we systematically characterized the cell types targeted in our collection. In addition, we identified correspondences between the cells in this collection and a recent connectomic data set of the ventral nerve cord. Taken together, the resources and results presented here form a powerful toolkit for future investigations of neuronal circuits and connectivity of premotor circuits while linking them to behavioral outputs.
Background Many Drosophila species use acoustic communication during courtship and studies of these communication systems have provided insight into neurobiology, behavioral ecology, ethology, and evolution. Recording Drosophila courtship sounds and associated behavior is challenging, especially at high throughput, and previously designed devices are relatively expensive and complex to assemble. Results We present construction plans for a modular system utilizing mostly off-the-shelf, relatively inexpensive components that provides simultaneous high-resolution audio and video recording of 96 isolated or paired Drosophila individuals. We provide open-source control software to record audio and video. We designed high intensity LED arrays that can be used to perform optogenetic activation and inactivation of labelled neurons. The basic design can be modified to facilitate novel study designs or to record insects larger than Drosophila. Fewer than 96 microphones can be used in the system if the full array is not required or to reduce costs. Implications Our hardware design and software provide an improved platform for reliable and comparatively inexpensive high-throughput recording of Drosophila courtship acoustic and visual behavior and perhaps for recording acoustic signals of other small animals.
Multiple methods have been introduced over the past 30 years to identify the genomic insertion sites of transposable elements and other DNA elements that integrate into genomes. However, each of these methods suffer from limitations that can frustrate attempts to map multiple insertions in a single genome and to map insertions in genomes of high complexity that contain extensive repetitive DNA. I introduce a new method for transposon mapping that is simple to perform, can accurately map multiple insertions per genome, and generates long sequence reads that facilitate mapping to complex genomes. The method, called TagMap, for Tagmentation-based Mapping, relies on a modified Tn5 tagmentation protocol with a single tagmentation adaptor followed by PCR using primers specific to the tranposable element and the adaptor sequence. Several minor modifications to normal tagmentation reagents and protocols allow easy and rapid preparation of TagMap libraries. Short read sequencing starting from the adaptor sequence generates oriented reads that flank and are oriented toward the transposable element insertion site. The convergent orientation of adjacent reads at the insertion site allows straightforward prediction of the precise insertion site(s). A Linux shell script is provided to identify insertion sites from fastq files.
We tested whether transcription activator-like effectors (TALEs) could mediate repression and activation of endogenous enhancers in the Drosophila genome. TALE repressors (TALERs) targeting each of the five even-skipped (eve) stripe enhancers generated repression specifically of the focal stripes. TALE activators (TALEAs) targeting the eve promoter or enhancers caused increased expression primarily in cells normally activated by the promoter or targeted enhancer, respectively. This effect supports the view that repression acts in a dominant fashion on transcriptional activators and that the activity state of an enhancer influences TALE binding or the ability of the VP16 domain to enhance transcription. In these assays, the Hairy repression domain did not exhibit previously described long-range transcriptional repression activity. The phenotypic effects of TALER and TALEA expression in larvae and adults are consistent with the observed modulations of eve expression. TALEs thus provide a novel tool for detection and functional modulation of transcriptional enhancers in their native genomic context.
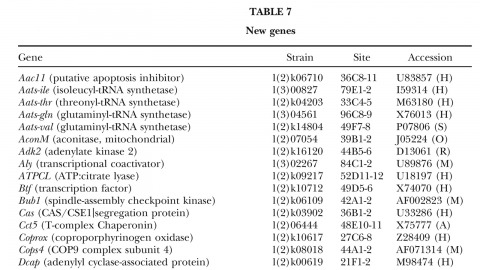
A fundamental goal of genetics and functional genomics is to identify and mutate every gene in model organisms such as Drosophila melanogaster. The Berkeley Drosophila Genome Project (BDGP) gene disruption project generates single P-element insertion strains that each mutate unique genomic open reading frames. Such strains strongly facilitate further genetic and molecular studies of the disrupted loci, but it has remained unclear if P elements can be used to mutate all Drosophila genes. We now report that the primary collection has grown to contain 1045 strains that disrupt more than 25% of the estimated 3600 Drosophila genes that are essential for adult viability. Of these P insertions, 67% have been verified by genetic tests to cause the associated recessive mutant phenotypes, and the validity of most of the remaining lines is predicted on statistical grounds. Sequences flanking >920 insertions have been determined to exactly position them in the genome and to identify 376 potentially affected transcripts from collections of EST sequences. Strains in the BDGP collection are available from the Bloomington Stock Center and have already assisted the research community in characterizing >250 Drosophila genes. The likely identity of 131 additional genes in the collection is reported here. Our results show that Drosophila genes have a wide range of sensitivity to inactivation by P elements, and provide a rationale for greatly expanding the BDGP primary collection based entirely on insertion site sequencing. We predict that this approach can bring >85% of all Drosophila open reading frames under experimental control.
The density and distribution of regulatory information in non-coding DNA of eukaryotic genomes is largely unknown. Evolutionary analyses have estimated that ∼60% of nucleotides in intergenic regions of the D. melanogaster genome is functionally relevant. This estimate is difficult to reconcile with the commonly accepted idea that enhancers are compact regulatory elements that generally encompass less than 1 kilobase of DNA. Here, we approached this issue through a functional dissection of the regulatory region of the gene shavenbaby (svb). Most of the ∼90 kilobases of this large regulatory region is highly conserved in the genus Drosophila, though characterized enhancers occupy a small fraction of this region. By analyzing the regulation of svb in different contexts of Drosophila development, we found that the regulatory architecture that drives svb expression in the abdominal pupal epidermis is organized in a dramatically different way than the information that drives svb expression in the embryonic epidermis. While in the embryonic epidermis svb is activated by compact and dispersed enhancers, svb expression in the pupal epidermis is driven by large regions with enhancer activity, which occupy a great portion of the svb cis-regulatory DNA. We observed that other developmental genes also display a dense distribution of putative regulatory elements in their regulatory regions. Furthermore, we found that a large percentage of conserved non-coding DNA of the Drosophila genome is contained within putative regulatory DNA. These results suggest that part of the evolutionary constraint on non-coding DNA of Drosophila is explained by the density of regulatory information.
Within all species of animals, the size of each organ bears a specific relationship to overall body size. These patterns of organ size relative to total body size are called static allometry and have enchanted biologists for centuries, yet the mechanisms generating these patterns have attracted little experimental study. We review recent and older work on holometabolous insect development that sheds light on these mechanisms. In insects, static allometry can be divided into at least two processes: (1) the autonomous specification of organ identity, perhaps including the approximate size of the organ, and (2) the determination of the final size of organs based on total body size. We present three models to explain the second process: (1) all organs autonomously absorb nutrients and grow at organ-specific rates, (2) a centralized system measures a close correlate of total body size and distributes this information to all organs, and (3) autonomous organ growth is combined with feedback between growing organs to modulate final sizes. We provide evidence supporting models 2 and 3 and also suggest that hormones are the messengers of size information. Advances in our understanding of the mechanisms of allometry will come through the integrated study of whole tissues using techniques from development, genetics, endocrinology and population biology.
What is the relationship between variation that segregates within natural populations and the differences that distinguish species? Many studies over the past century have demonstrated that most of the genetic variation within natural populations that contributes to quantitative traits causes relatively small phenotypic effects. In contrast, the genetic causes of quantitative differences between species are at least sometimes caused by few loci of relatively large effect. In addition, most of the results from evolutionary developmental biology are often discussed as though changes at just a few important 'molecular toolbox' genes provide the key clues to morphological evolution. On the face of it, these divergent results seem incompatible and call into question the neo-Darwinian view that differences between species emerge from precisely the same kinds of variants that segregate much of the time in natural populations. One prediction from the classical model is that many different genes can evolve to generate similar phenotypes. I discuss our studies that demonstrate that similar phenotypes have evolved in multiple lineages of Drosophila by evolution of the same gene, shavenbaby/ovo. This evidence for parallel evolution suggests that svb occupies a privileged position in the developmental network patterning larval trichomes that makes it a favourable target of evolutionary change.