Inside of a cell, snaking membranes house tiny factories that pump out proteins.
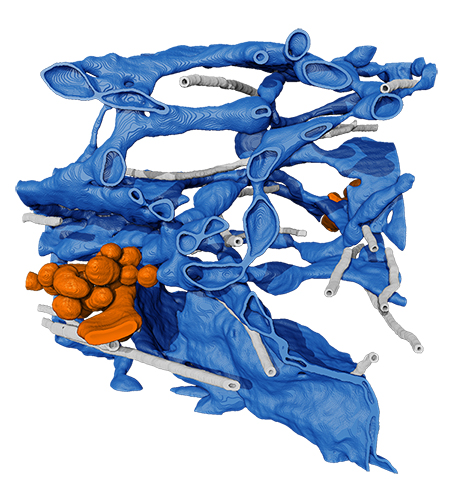
Now, detailed microscope data gives a new view of exactly what’s happening as the proteins made here are sent out into the cell: A network of stretchy tubes delivers the proteins directly to the next stop on their journey. The structure seems to be able to adapt to handle different kinds and quantities of proteins, giving cells flexibility in their protein exporting capabilities, says senior group leader Jennifer Lippincott-Schwartz. Her team describes their findings in a paper published April 13 in the journal Cell.
Cells contain genetic instructions to make thousands of different kinds of proteins, which carry out many different jobs. And about a third of those proteins are produced in this twisting structure called the endoplasmic reticulum. Their journey is a long one — they’re built, folded, sorted, packaged, and then transported to their final destination. Lippincott-Schwartz’s team focused on just one part of the process: What happens as just-made proteins leave the production floor and head to vehicles that distribute them elsewhere in the cell. That export process takes place at specialized spots across the endoplasmic reticulum, called ER exit sites.
“There’s a lot of data and experiments in this area, but they’re inconsistent in that they don’t fit into one cohesive model,” says Aubrey Weigel, a scientist at Janelia who worked on the project as a postdoc in Lippincott-Schwartz’s lab, alongside postdoc Chi-Lun Chang. By surveying the whole cell, rather than looking at small regions, the researchers could get a clearer understanding of what was going on. “It’s a first big step in redefining how this works in a mammalian system,” Weigel says.
To survey the protein export step in detail, the researchers combined recently developed forms of light and electron microscopy. Electron microscopy delivers high-resolution images, but in black and white. Light microscopy is lower resolution, but fluorescent tagging makes it easier to ID certain features. And adding live cell imaging into the mix helped link structure to function.
“I think this is one of the first papers that uses these advanced imaging tools to reveal something about biology,” says Chang. Rather than just capture a snapshot in time, “we can actually study the dynamics using these different tools.”
Previously, cell biologists assumed that proteins left the ER encased within small, spherical, membrane-bound vesicles. “What we're showing is that it's not that simple of a system,” Weigel says.
Instead, the team found that proteins left the ER to their next destination via shape-shifting tubular outgrowths. These tubes can expand and contract and stretch and even inch along microtubules, adapting on the fly depending on what’s being released. “Surprisingly, the exit sites are actually a continuous outgrowth of the ER, connected to it through a narrow neck,” says Lippincott-Schwartz.
Such flexibility has implications for many different kinds of cellular processes. Every cell contains instructions to make thousands of different proteins, written in its DNA. What gets produced at any given moment (and in what quantities) depends on what the cell needs. So, the pathways carrying proteins out of the ER need to be adaptable — to be able to push out proteins at top speed but also to go slow; to package tiny proteins as well as massive ones.
For instance, when immune cells begin producing antibodies against a specific pathogen, the cells “go into full throttle” and start furiously churning out antibodies, Lippincott-Schwartz says. Without some kind of adaptability, such a huge outflow of antibody proteins might cause a traffic jam, like a crowd of people trying to leave a stadium after a sports game. But instead, the tubular membranes of the ER exit sites can stretch and morph to let proteins leave efficiently.
Now, the researchers are taking the findings a step further, examining how this shifting tubular network might help explain cells’ adaptability under different kinds of extreme conditions.
###
Citation:
Aubrey V. Weigel, Chi-Lun Chang, Gleb Shtengel, C. Shan Xu, David P. Hoffman, Melanie Freeman, Nirmala Iyer, Jesse Aaron, Satya Khuon, John Bogovic, Wei Qiu, Harald F. Hess, and Jennifer Lippincott-Schwartz. “ER to Golgi protein delivery through an interwoven, tubular network extending from ER.” Cell, published online April 13, 2021. Doi:10.1016/j.cell.2021.03.035.










