Filter
Associated Lab
- Ahrens Lab (3) Apply Ahrens Lab filter
- Aso Lab (3) Apply Aso Lab filter
- Baker Lab (5) Apply Baker Lab filter
- Betzig Lab (8) Apply Betzig Lab filter
- Branson Lab (6) Apply Branson Lab filter
- Card Lab (1) Apply Card Lab filter
- Cardona Lab (1) Apply Cardona Lab filter
- Chklovskii Lab (2) Apply Chklovskii Lab filter
- Cui Lab (2) Apply Cui Lab filter
- Dickson Lab (5) Apply Dickson Lab filter
- Druckmann Lab (1) Apply Druckmann Lab filter
- Dudman Lab (4) Apply Dudman Lab filter
- Eddy/Rivas Lab (3) Apply Eddy/Rivas Lab filter
- Egnor Lab (1) Apply Egnor Lab filter
- Fetter Lab (1) Apply Fetter Lab filter
- Freeman Lab (3) Apply Freeman Lab filter
- Gonen Lab (10) Apply Gonen Lab filter
- Grigorieff Lab (3) Apply Grigorieff Lab filter
- Harris Lab (2) Apply Harris Lab filter
- Heberlein Lab (1) Apply Heberlein Lab filter
- Hess Lab (5) Apply Hess Lab filter
- Jayaraman Lab (1) Apply Jayaraman Lab filter
- Ji Lab (3) Apply Ji Lab filter
- Karpova Lab (1) Apply Karpova Lab filter
- Keller Lab (7) Apply Keller Lab filter
- Lavis Lab (7) Apply Lavis Lab filter
- Lee (Albert) Lab (4) Apply Lee (Albert) Lab filter
- Leonardo Lab (4) Apply Leonardo Lab filter
- Liu (Zhe) Lab (4) Apply Liu (Zhe) Lab filter
- Looger Lab (11) Apply Looger Lab filter
- Magee Lab (1) Apply Magee Lab filter
- Menon Lab (3) Apply Menon Lab filter
- Murphy Lab (1) Apply Murphy Lab filter
- Reiser Lab (2) Apply Reiser Lab filter
- Riddiford Lab (4) Apply Riddiford Lab filter
- Rubin Lab (8) Apply Rubin Lab filter
- Saalfeld Lab (1) Apply Saalfeld Lab filter
- Scheffer Lab (7) Apply Scheffer Lab filter
- Simpson Lab (2) Apply Simpson Lab filter
- Singer Lab (4) Apply Singer Lab filter
- Spruston Lab (1) Apply Spruston Lab filter
- Stern Lab (6) Apply Stern Lab filter
- Sternson Lab (5) Apply Sternson Lab filter
- Svoboda Lab (7) Apply Svoboda Lab filter
- Tervo Lab (1) Apply Tervo Lab filter
- Tjian Lab (4) Apply Tjian Lab filter
- Truman Lab (1) Apply Truman Lab filter
- Wu Lab (2) Apply Wu Lab filter
- Zlatic Lab (1) Apply Zlatic Lab filter
- Zuker Lab (1) Apply Zuker Lab filter
Associated Project Team
Associated Support Team
Publication Date
- December 2014 (31) Apply December 2014 filter
- November 2014 (7) Apply November 2014 filter
- October 2014 (14) Apply October 2014 filter
- September 2014 (14) Apply September 2014 filter
- August 2014 (11) Apply August 2014 filter
- July 2014 (23) Apply July 2014 filter
- June 2014 (13) Apply June 2014 filter
- May 2014 (10) Apply May 2014 filter
- April 2014 (18) Apply April 2014 filter
- March 2014 (13) Apply March 2014 filter
- February 2014 (11) Apply February 2014 filter
- January 2014 (25) Apply January 2014 filter
- Remove 2014 filter 2014
190 Janelia Publications
Showing 41-50 of 190 resultsDrosophila melanogaster females respond to male courtship by either rejecting the male or allowing copulation. The neural mechanisms underlying these female behaviors likely involve the integration of sensory information in the brain. Because doublesex (dsx) controls other aspects of female differentiation, we asked whether dsx-expressing neurons mediate virgin female receptivity to courting males. Using intersectional techniques to manipulate the activities of defined subsets of dsx-expressing neurons, we found that activation of neurons in either the pCd or pC1 clusters promotes receptivity, while silencing these neurons makes females unreceptive. Furthermore, pCd and pC1 neurons physiologically respond to the male-specific pheromone cis-vaccenyl acetate (cVA), while pC1 neurons also respond to male courtship song. The pCd and pC1 neurons expressing dsx in females do not express transcripts from the fruitless (fru) P1 promoter. Thus, virgin female receptivity is controlled at least in part by neurons that are distinct from those governing male courtship.
To gain insights into coordinated lineage-specification and morphogenetic processes during early embryogenesis, here we report a systematic identification of transcriptional programs mediated by a key developmental regulator-Brachyury. High-resolution chromosomal localization mapping of Brachyury by ChIP sequencing and ChIP-exonuclease revealed distinct sequence signatures enriched in Brachyury-bound enhancers. A combination of genome-wide in vitro and in vivo perturbation analysis and cross-species evolutionary comparison unveiled a detailed Brachyury-dependent gene-regulatory network that directly links the function of Brachyury to diverse developmental pathways and cellular housekeeping programs. We also show that Brachyury functions primarily as a transcriptional activator genome-wide and that an unexpected gene-regulatory feedback loop consisting of Brachyury, Foxa2, and Sox17 directs proper stem-cell lineage commitment during streak formation. Target gene and mRNA-sequencing correlation analysis of the T(c) mouse model supports a crucial role of Brachyury in up-regulating multiple key hematopoietic and muscle-fate regulators. Our results thus chart a comprehensive map of the Brachyury-mediated gene-regulatory network and how it influences in vivo developmental homeostasis and coordination.
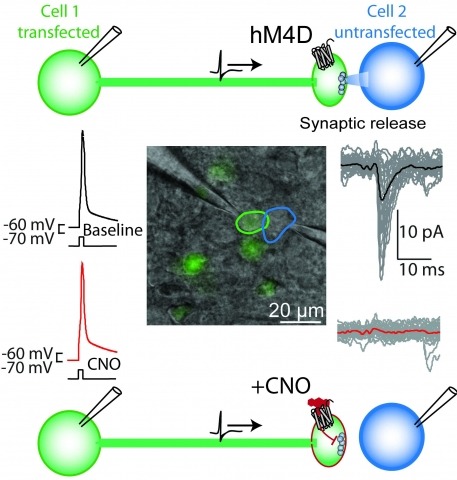
Brain function is mediated by neural circuit connectivity, and elucidating the role of connections is aided by techniques to block their output. We developed cell-type-selective, reversible synaptic inhibition tools for mammalian neural circuits by leveraging G protein signaling pathways to suppress synaptic vesicle release. Here, we find that the pharmacologically selective designer Gi-protein-coupled receptor hM4D is a presynaptic silencer in the presence of its cognate ligand clozapine-N-oxide (CNO). Activation of hM4D signaling sharply reduced synaptic release probability and synaptic current amplitude. To demonstrate the utility of this tool for neural circuit perturbations, we developed an axon-selective hM4D-neurexin variant and used spatially targeted intracranial CNO injections to localize circuit connections from the hypothalamus to the midbrain responsible for feeding behavior. This synaptic silencing approach is broadly applicable for cell-type-specific and axon projection-selective functional analysis of diverse neural circuits.
Elucidating the roles of neuronal cell types for physiology and behavior is essential for understanding brain functions. Perturbation of neuron electrical activity can be used to probe the causal relationship between neuronal cell types and behavior. New genetically encoded neuron perturbation tools have been developed for remotely controlling neuron function using small molecules that activate engineered receptors that can be targeted to cell types using genetic methods. Here we describe recent progress for approaches using genetically engineered receptors that selectively interact with small molecules. Called "chemogenetics," receptors with diverse cellular functions have been developed that facilitate the selective pharmacological control over a diverse range of cell-signaling processes, including electrical activity, for molecularly defined cell types. These tools have revealed remarkably specific behavioral physiological influences for molecularly defined cell types that are often intermingled with populations having different or even opposite functions.
Chromatin immunoprecipitation (ChIP) is a technique that reveals in vivo location of a protein bound to DNA. ChIP coupled with DNA microarrays (ChIP-chip) or next-generation sequencing (ChIP-seq) allows for identification of binding sites of transcription factors on a global scale. Here we describe a protocol for ChIP to identify binding of the Ultrabithorax (Ubx) Hox transcription factors from imaginal discs of Drosophila larvae. The protocol can be extended to other model organisms and transcription factors.
Integrin alpha M (ITGAM; CD11b) is a component of the macrophage-1 antigen complex, which mediates leukocyte adhesion, migration and phagocytosis as part of the immune system. We previously identified a missense polymorphism, rs1143679 (R77H), strongly associated with systemic lupus erythematosus (SLE). However, the molecular mechanisms of this variant are incompletely understood. A meta-analysis of published and novel data on 28 439 individuals with European, African, Hispanic and Asian ancestries reinforces genetic association between rs1143679 and SLE [Pmeta = 3.60 × 10(-90), odds ratio (OR) = 1.76]. Since rs1143679 is in the most active region of chromatin regulation and transcription factor binding in ITGAM, we quantitated ITGAM RNA and surface protein levels in monocytes from patients with each rs1143679 genotype. We observed that transcript levels significantly decreased for the risk allele ('A') relative to the non-risk allele ('G'), in a dose-dependent fashion: ('AA' < 'AG' < 'GG'). CD11b protein levels in patients' monocytes were directly correlated with RNA levels. Strikingly, heterozygous individuals express much lower (average 10- to 15-fold reduction) amounts of the 'A' transcript than 'G' transcript. We found that the non-risk sequence surrounding rs1143679 exhibits transcriptional enhancer activity in vivo and binds to Ku70/80, NFKB1 and EBF1 in vitro, functions that are significantly reduced with the risk allele. Mutant CD11b protein shows significantly reduced binding to fibrinogen and vitronectin, relative to non-risk, both in purified protein and in cellular models. This two-pronged contribution (nucleic acid- and protein-level) of the rs1143679 risk allele to decreasing ITGAM activity provides insight into the molecular mechanisms of its potent association with SLE.
The hemoglobinopathies, such as β-thalassemia and sickle cell anemia (SCA), are characterized by mutations of the β-globin gene resulting in either decreased or functionally abnormal hemoglobin (Hb) production. As bone marrow transplant is the only curative option for these patients, there is a strong need for new therapeutic approaches. Both β-thalassemia and SCA represent ideal targets for gene therapy since introduction of a normal β-globin gene can ameliorate the phenotype, as we and others have shown previously. Overcoming the developmental silencing of the fetal γ-globin gene represents an additional approach for the treatment of hemoglobinopathies. Here, we directly compare a recently established approach to activate the γ-globin gene using forced chromatin looping with pharmacologic approaches to raise γ-globin expression. The β-type globin genes are activated through dynamic interactions with a distal upstream enhancer, the locus control region (LCR). The LCR physically contacts the developmental stage appropriate globin gene via chromatin looping, a process partially dependent on the protein Ldb1. Previously, we have shown that tethering Ldb1 to the murine β-globin promoter with a custom designed zinc finger protein (ZF-Ldb1) can induce loop formation and β-globin transcription in an erythroid cell line (Deng et al., 2012). Further work showed that forced chromatin looping can be exploited to potently reactivate fetal globin gene expression in adult human erythroid cells (Deng et al., 2014). Here we compared the efficacy and toxicity of ZF-Ldb1 to pharmacologic compounds that induce HbF in cultured hematopoietic stem progenitor cell-derived erythroid cultures from normal and SCA donors. ZF-Ldb1 increased HbF synthesis in SCA erythroid cells (N=8) up to 86% and, concurrently, reduced sickle Hb (HbS) below 15%, consistent with previous studies of erythroid cells from normal probands. Preliminary results obtained from treating SCA specimens (N=3) show that the induction of HbF in cells treated with ZF-Ldb1 is twice as high (+35.55% ± 8.34%, at a dose of ~ one ZF-Ldb1 transgene copy per cell) as that observed using pomalidomide (+16.50% ± 14.57%, 20μM) and decitabine (+15.60% ± 12.36%, 0.5μM). Tranylcypromine and hydroxyurea showed the lowest HbF increase (+9.67% ± 3.26% and +5.06 ± 2.82%, 1.5μM and 150μM respectively). Importantly, decitabine and pomalidomide treatment lowered cell viability to 39% and 26%, respectively, while ZF-Ldb1 expressing cells retained normal viability similar to control populations. In related experiments, we are comparing the expression of a battery of genes known to regulate HbF levels (BCL11A, SOX6, KLF1 and C-Myb) in normal and SCA derived erythroid cells treated with ZF-Ldb1 or HbF inducers and compared to controls. Preliminary analyses indicate altered expression of KLF1 in SCA versus normal cells, consistent with a superior response of SCA cells to HbF induction. In conclusion, lentiviral-mediated ZF-Ldb1 gene transfer appears superior to pharmacologic compounds in terms of efficacy and cell viability further supporting suitability for the reactivation of HbF in SCA erythroid cells.
BACKGROUND: Recording of physiological parameters in behaving mice has seen an immense increase over recent years driven by, for example, increased miniaturization of recording devices. One parameter particularly important for odorant-driven behaviors is the breathing frequency, since the latter dictates the rate of odorant delivery to the nasal cavity and the olfactory receptor neurons located therein. NEW METHOD: Typically, breathing patterns are monitored by either measuring the breathing-induced temperature or pressure changes in the nasal cavity. Both require the implantation of a nasal cannula and tethering of the mouse to either a cable or tubing. To avoid these limitations we used an implanted pressure sensor which reads the thoracic pressure and transmits the data telemetrically, thus making it suitable for experiments which require a freely moving animal. RESULTS: Mice performed a Go/NoGo odorant-driven behavioral task with the implanted pressure sensor, which proved to work reliably to allow recording of breathing signals over several weeks from a given animal. COMPARISON TO EXISTING METHOD(S): We simultaneously recorded the thoracic and nasal pressure changes and found that measuring the thoracic pressure change yielded similar results compared to measurements of nasal pressure changes. CONCLUSION: Telemetrically recorded breathing signals are a feasible method to monitor odorant-guided behavioral changes in breathing rates. Its advantages are most significant when recording from a freely moving animal over several weeks. The advantages and disadvantages of different methods to record breathing patterns are discussed.
Transcriptomics experiments and computational predictions both enable systematic discovery of new functional RNAs. However, many putative noncoding transcripts arise instead from artifacts and biological noise, and current computational prediction methods have high false positive rates. I discuss prospects for improving computational methods for analyzing and identifying functional RNAs, with a focus on detecting signatures of conserved RNA secondary structure. An interesting new front is the application of chemical and enzymatic experiments that probe RNA structure on a transcriptome-wide scale. I review several proposed approaches for incorporating structure probing data into the computational prediction of RNA secondary structure. Using probabilistic inference formalisms, I show how all these approaches can be unified in a well-principled framework, which in turn allows RNA probing data to be easily integrated into a wide range of analyses that depend on RNA secondary structure inference. Such analyses include homology search and genome-wide detection of new structural RNAs.
The spatiotemporal activities of astrocyte Ca(2+) signaling in mature neuronal circuits remain unclear. We used genetically encoded Ca(2+) and glutamate indicators as well as pharmacogenetic and electrical control of neurotransmitter release to explore astrocyte activity in the hippocampal mossy fiber pathway. Our data revealed numerous localized, spontaneous Ca(2+) signals in astrocyte branches and territories, but these were not driven by neuronal activity or glutamate. Moreover, evoked astrocyte Ca(2+) signaling changed linearly with the number of mossy fiber action potentials. Under these settings, astrocyte responses were global, suppressed by neurotransmitter clearance, and mediated by glutamate and GABA. Thus, astrocyte engagement in the fully developed mossy fiber pathway was slow and territorial, contrary to that frequently proposed for astrocytes within microcircuits. We show that astrocyte Ca(2+) signaling functionally segregates large volumes of neuropil and that these transients are not suited for responding to, or regulating, single synapses in the mossy fiber pathway.