Filter
Associated Lab
- Aso Lab (1) Apply Aso Lab filter
- Bock Lab (1) Apply Bock Lab filter
- Card Lab (3) Apply Card Lab filter
- Cardona Lab (1) Apply Cardona Lab filter
- Chklovskii Lab (1) Apply Chklovskii Lab filter
- Dickson Lab (1) Apply Dickson Lab filter
- Fetter Lab (3) Apply Fetter Lab filter
- Funke Lab (2) Apply Funke Lab filter
- Harris Lab (1) Apply Harris Lab filter
- Hess Lab (9) Apply Hess Lab filter
- Jayaraman Lab (1) Apply Jayaraman Lab filter
- Reiser Lab (1) Apply Reiser Lab filter
- Rubin Lab (10) Apply Rubin Lab filter
- Saalfeld Lab (2) Apply Saalfeld Lab filter
- Scheffer Lab (20) Apply Scheffer Lab filter
- Stern Lab (1) Apply Stern Lab filter
- Truman Lab (1) Apply Truman Lab filter
- Turner Lab (1) Apply Turner Lab filter
Associated Project Team
Associated Support Team
- Janelia Experimental Technology (1) Apply Janelia Experimental Technology filter
- Project Technical Resources (3) Apply Project Technical Resources filter
- Scientific Computing Software (13) Apply Scientific Computing Software filter
- Scientific Computing Systems (1) Apply Scientific Computing Systems filter
Publication Date
- 2024 (5) Apply 2024 filter
- 2023 (4) Apply 2023 filter
- 2020 (3) Apply 2020 filter
- 2019 (3) Apply 2019 filter
- 2018 (4) Apply 2018 filter
- 2017 (4) Apply 2017 filter
- 2016 (1) Apply 2016 filter
- 2015 (7) Apply 2015 filter
- 2014 (11) Apply 2014 filter
- 2013 (3) Apply 2013 filter
- 2012 (2) Apply 2012 filter
- 2010 (4) Apply 2010 filter
51 Janelia Publications
Showing 31-40 of 51 resultsMany motor control systems generate multiple movements using a common set of muscles. How are premotor circuits able to flexibly generate diverse movement patterns? Here, we characterize the neuronal circuits that drive the distinct courtship songs of Drosophila melanogaster. Male flies vibrate their wings towards females to produce two different song modes – pulse and sine song – which signal species identity and male quality. Using cell-type specific genetic reagents and the connectome, we provide a cellular and synaptic map of the circuits in the male ventral nerve cord that generate these songs and examine how activating or inhibiting each cell type within these circuits affects the song. Our data reveal that the song circuit is organized into two nested feed-forward pathways, with extensive reciprocal and feed-back connections. The larger network produces pulse song, the more complex and ancestral song form. A subset of this network produces sine song, the simpler and more recent form. Such nested organization may be a common feature of motor control circuits in which evolution has layered increasing flexibility on to a basic movement pattern.
Neuroscience research in Drosophila is benefiting from large-scale connectomics efforts using electron microscopy (EM) to reveal all the neurons in a brain and their connections. To exploit this knowledge base, researchers relate a connectome’s structure to neuronal function, often by studying individual neuron cell types. Vast libraries of fly driver lines expressing fluorescent reporter genes in sets of neurons have been created and imaged using confocal light microscopy (LM), enabling the targeting of neurons for experimentation. However, creating a fly line for driving gene expression within a single neuron found in an EM connectome remains a challenge, as it typically requires identifying a pair of driver lines where only the neuron of interest is expressed in both. This task and other emerging scientific workflows require finding similar neurons across large data sets imaged using different modalities. Here, we present NeuronBridge, a web application for easily and rapidly finding putative morphological matches between large data sets of neurons imaged using different modalities. We describe the functionality and construction of the NeuronBridge service, including its user-friendly graphical user interface (GUI), extensible data model, serverless cloud architecture, and massively parallel image search engine. NeuronBridge fills a critical gap in the Drosophila research workflow and is used by hundreds of neuroscience researchers around the world. We offer our software code, open APIs, and processed data sets for integration and reuse, and provide the application as a service at http://neuronbridge.janelia.org.Background
Results
Conclusions
Reconstructing a connectome from an EM dataset often requires a large effort of proofreading automatically generated segmentations. While many tools exist to enable tracing or proofreading, recent advances in EM imaging and segmentation quality suggest new strategies and pose unique challenges for tool design to accelerate proofreading. Namely, we now have access to very large multi-TB EM datasets where (1) many segments are largely correct, (2) segments can be very large (several GigaVoxels), and where (3) several proofreaders and scientists are expected to collaborate simultaneously. In this paper, we introduce NeuTu as a solution to efficiently proofread large, high-quality segmentation in a collaborative setting. NeuTu is a client program of our high-performance, scalable image database called DVID so that it can easily be scaled up. Besides common features of typical proofreading software, NeuTu tames unprecedentedly large data with its distinguishing functions, including: (1) low-latency 3D visualization of large mutable segmentations; (2) interactive splitting of very large false merges with highly optimized semi-automatic segmentation; (3) intuitive user operations for investigating or marking interesting points in 3D visualization; (4) visualizing proofreading history of a segmentation; and (5) real-time collaborative proofreading with lock-based concurrency control. These unique features have allowed us to manage the workflow of proofreading a large dataset smoothly without dividing them into subsets as in other segmentation-based tools. Most importantly, NeuTu has enabled some of the largest connectome reconstructions as well as interesting discoveries in the fly brain.
The brain is a network of neurons and its biological output is behaviour. This is an exciting age, with a growing acknowledgement that the comprehensive compilation of synaptic circuits densely reconstructed in the brains of model species is now both technologically feasible and a scientifically enabling possibility in neurobiology, much as 30 years ago genomics was in molecular biology and genetics. Implemented by huge advances in electron microscope technology, especially focused ion beam-scanning electron microscope (FIB-SEM) milling (see Glossary), image capture and alignment, and computer-aided reconstruction of neuron morphologies, enormous progress has been made in the last decade in the detailed knowledge of the actual synaptic circuits formed by real neurons, in various brain regions of the fly It is useful to distinguish synaptic pathways that are major, with 100 or more presynaptic contacts, from those that are minor, with fewer than about 10; most neurites are both presynaptic and postsynaptic, and all synaptic sites have multiple postsynaptic dendrites. Work on has spearheaded these advances because cell numbers are manageable, and neuron classes are morphologically discrete and genetically identifiable, many confirmed by reporters. Recent advances are destined within the next few years to reveal the complete connectome in an adult fly, paralleling advances in the larval brain that offer the same prospect possibly within an even shorter time frame. The final amendment and validation of segmented bodies by human proof-readers remains the most time-consuming step, however. The value of a complete connectome in is that, by targeting to specific neurons transgenes that either silence or activate morphologically identified circuits, and then identifying the resulting behavioural outcome, we can determine the causal mechanism for behaviour from its loss or gain. More importantly, the connectome reveals hitherto unsuspected pathways, leading us to seek novel behaviours for these. Circuit information will eventually be required to understand how differences between brains underlie differences in behaviour, and especially to herald yet more advanced connectomic strategies for the vertebrate brain, with an eventual prospect of understanding cognitive disorders having a connectomic basis. Connectomes also help us to identify common synaptic circuits in different species and thus to reveal an evolutionary progression in candidate pathways.
The increasing IC manufacturing cost encourages a business model where design houses outsource IC fabrication to remote foundries. Despite cost savings, this model exposes design houses to IC piracy as remote foundries can manufacture in excess to sell on the black market. Recent efforts in digital hardware security aim to thwart piracy by using XOR-based chip locking, cryptography, and active metering. To counter direct attacks and lower the exposure of unlocked circuits to the foundry, we introduce a multiplexor-based locking strategy that preserves test response allowing IC testing by an untrusted party before activation. We demonstrate a simple yet effective attack against a locked circuit that does not preserve test response, and validate the effectiveness of our locking strategy on IWLS 2005 benchmarks.
View Publication PageConnectomics-the study of how neurons wire together in the brain-is at the forefront of modern neuroscience research. However, many connectomics studies are limited by the time and precision needed to correctly segment large volumes of electron microscopy (EM) image data. We present here a semi-automated segmentation pipeline using freely available software that can significantly decrease segmentation time for extracting both nuclei and cell bodies from EM image volumes.
The brain of fruit fly Drosophila melanogaster has been used as a model system for functional analysis of neuronal circuits, including connectomics research, due to its modest size (~700 μm) and availability of abundant molecular genetics tools for visualizing neurons. Three-dimensional (3D) reconstruction of high-resolution images of neurons or circuits visualized with appropriate methods is a critical step for obtaining information such as morphology and connectivity patterns of neuronal circuits. In this chapter, we introduce methods for generating 3D reconstructed images with data acquired from confocal laser scanning microscopy (CLSM) or electron microscopy (EM) to analyze neuronal circuits found in the central nervous system (CNS) of the fruit fly. Comparisons of different algorithms and strategies for reconstructing neuronal circuits, using actual studies as references, will be discussed within this chapter.
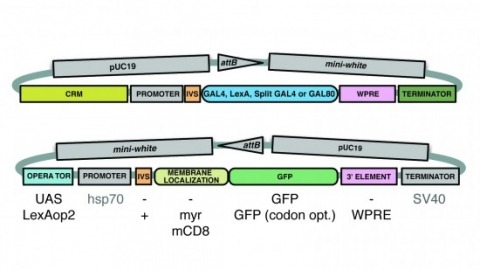
A wide variety of biological experiments rely on the ability to express an exogenous gene in a transgenic animal at a defined level and in a spatially and temporally controlled pattern. We describe major improvements of the methods available for achieving this objective in Drosophila melanogaster. We have systematically varied core promoters, UTRs, operator sequences, and transcriptional activating domains used to direct gene expression with the GAL4, LexA, and Split GAL4 transcription factors and the GAL80 transcriptional repressor. The use of site-specific integration allowed us to make quantitative comparisons between different constructs inserted at the same genomic location. We also characterized a set of PhiC31 integration sites for their ability to support transgene expression of both drivers and responders in the nervous system. The increased strength and reliability of these optimized reagents overcome many of the previous limitations of these methods and will facilitate genetic manipulations of greater complexity and sophistication.
Reconstructing neuronal circuits at the level of synapses is a central problem in neuroscience, and the focus of the nascent field of connectomics. Previously used to reconstruct the C. elegans wiring diagram, serial-section transmission electron microscopy (ssTEM) is a proven technique for the task. However, to reconstruct more complex circuits, ssTEM will require the automation of image processing. We review progress in the processing of electron microscopy images and, in particular, a semi-automated reconstruction pipeline deployed at Janelia. Drosophila circuits underlying identified behaviors are being reconstructed in the pipeline with the goal of generating a complete Drosophila connectome.