Filter
Associated Lab
- Aso Lab (1) Apply Aso Lab filter
- Dudman Lab (1) Apply Dudman Lab filter
- Gonen Lab (1) Apply Gonen Lab filter
- Ji Lab (1) Apply Ji Lab filter
- Keller Lab (1) Apply Keller Lab filter
- Looger Lab (1) Apply Looger Lab filter
- Rubin Lab (1) Apply Rubin Lab filter
- Scheffer Lab (1) Apply Scheffer Lab filter
- Simpson Lab (1) Apply Simpson Lab filter
- Svoboda Lab (1) Apply Svoboda Lab filter
Associated Project Team
Associated Support Team
Publication Date
- September 26, 2014 (4) Apply September 26, 2014 filter
- September 24, 2014 (1) Apply September 24, 2014 filter
- September 19, 2014 (1) Apply September 19, 2014 filter
- September 14, 2014 (1) Apply September 14, 2014 filter
- September 12, 2014 (1) Apply September 12, 2014 filter
- September 5, 2014 (3) Apply September 5, 2014 filter
- September 3, 2014 (2) Apply September 3, 2014 filter
- September 1, 2014 (1) Apply September 1, 2014 filter
- Remove September 2014 filter September 2014
- Remove 2014 filter 2014
14 Janelia Publications
Showing 1-10 of 14 resultsMotor sequences are formed through the serial execution of different movements, but how nervous systems implement this process remains largely unknown. We determined the organizational principles governing how dirty fruit flies groom their bodies with sequential movements. Using genetically targeted activation of neural subsets, we drove distinct motor programs that clean individual body parts. This enabled competition experiments revealing that the motor programs are organized into a suppression hierarchy; motor programs that occur first suppress those that occur later. Cleaning one body part reduces the sensory drive to its motor program, which relieves suppression of the next movement, allowing the grooming sequence to progress down the hierarchy. A model featuring independently evoked cleaning movements activated in parallel, but selected serially through hierarchical suppression, was successful in reproducing the grooming sequence. This provides the first example of an innate motor sequence implemented by the prevailing model for generating human action sequences.
Reconstructing neuronal circuits at the level of synapses is a central problem in neuroscience and becoming a focus of the emerging field of connectomics. To date, electron microscopy (EM) is the most proven technique for identifying and quantifying synaptic connections. As advances in EM make acquiring larger datasets possible, subsequent manual synapse identification ({\em i.e.}, proofreading) for deciphering a connectome becomes a major time bottleneck. Here we introduce a large-scale, high-throughput, and semi-automated methodology to efficiently identify synapses. We successfully applied our methodology to the Drosophila medulla optic lobe, annotating many more synapses than previous connectome efforts. Our approaches are extensible and will make the often complicated process of synapse identification accessible to a wider-community of potential proofreaders.
A major limitation in understanding embryonic development is the lack of cell type-specific markers. Existing gene expression and marker atlases provide valuable tools, but they typically have one or more limitations: a lack of single-cell resolution; an inability to register multiple expression patterns to determine their precise relationship; an inability to be upgraded by users; an inability to compare novel patterns with the database patterns; and a lack of three-dimensional images. Here, we develop new 'atlas-builder' software that overcomes each of these limitations. A newly generated atlas is three-dimensional, allows the precise registration of an infinite number of cell type-specific markers, is searchable and is open-ended. Our software can be used to create an atlas of any tissue in any organism that contains stereotyped cell positions. We used the software to generate an 'eNeuro' atlas of the Drosophila embryonic CNS containing eight transcription factors that mark the major CNS cell types (motor neurons, glia, neurosecretory cells and interneurons). We found neuronal, but not glial, nuclei occupied stereotyped locations. We added 75 new Gal4 markers to the atlas to identify over 50% of all interneurons in the ventral CNS, and these lines allowed functional access to those interneurons for the first time. We expect the atlas-builder software to benefit a large proportion of the developmental biology community, and the eNeuro atlas to serve as a publicly accessible hub for integrating neuronal attributes - cell lineage, gene expression patterns, axon/dendrite projections, neurotransmitters--and linking them to individual neurons.
Mapping the connectivity of neurons in the brain (i.e., connectomics) is a challenging problem due to both the number of connections in even the smallest organisms and the nanometer resolution required to resolve them. Because of this, previous connectomes contain only hundreds of neurons, such as in the C.elegans connectome. Recent technological advances will unlock the mysteries of increasingly large connectomes (or partial connectomes). However, the value of these maps is limited by our ability to reason with this data and understand any underlying motifs. To aid connectome analysis, we introduce algorithms to cluster similarly-shaped neurons, where 3D neuronal shapes are represented as skeletons. In particular, we propose a novel location-sensitive clustering algorithm. We show clustering results on neurons reconstructed from the Drosophila medulla that show high-accuracy.
The origin of chordates has been debated for more than a century, with one key issue being the emergence of the notochord. In vertebrates, the notochord develops by convergence and extension of the chordamesoderm, a population of midline cells of unique molecular identity. We identify a population of mesodermal cells in a developing invertebrate, the marine annelid Platynereis dumerilii, that converges and extends toward the midline and expresses a notochord-specific combination of genes. These cells differentiate into a longitudinal muscle, the axochord, that is positioned between central nervous system and axial blood vessel and secretes a strong collagenous extracellular matrix. Ancestral state reconstruction suggests that contractile mesodermal midline cells existed in bilaterian ancestors. We propose that these cells, via vacuolization and stiffening, gave rise to the chordate notochord.
Pixel and superpixel classifiers have become essential tools for EM segmentation algorithms. Training these classifiers remains a major bottleneck primarily due to the requirement of completely annotating the dataset which is tedious, error-prone and costly. In this paper, we propose an interactive learning scheme for the superpixel classifier for EM segmentation. Our algorithm is "active semi-supervised" because it requests the labels of a small number of examples from user and applies label propagation technique to generate these queries. Using only a small set (<20%) of all datapoints, the proposed algorithm consistently generates a classifier almost as accurate as that estimated from a complete groundtruth. We provide segmentation results on multiple datasets to show the strength of these classifiers.
MicroED uses very small three-dimensional protein crystals and electron diffraction for structure determination. We present an improved data collection protocol for MicroED called 'continuous rotation'. Microcrystals are continuously rotated during data collection, yielding more accurate data. The method enables data processing with the crystallographic software tool MOSFLM, which resulted in improved resolution for the model protein lysozyme. These improvements are paving the way for the broad implementation and application of MicroED in structural biology.
In this work, we propose a learning framework for identifying synapses using a deep and wide multi-scale recursive (DAWMR) network, previously considered in image segmentation applications. We apply this approach on electron microscopy data from invertebrate fly brain tissue. By learning features directly from the data, we are able to achieve considerable improvements over existing techniques that rely on a small set of hand-designed features. We show that this system can reduce the amount of manual annotation required, in both acquisition of training data as well as verification of inferred detections.
Distal enhancers commonly contact target promoters via chromatin looping. In erythroid cells, the locus control region (LCR) contacts β-type globin genes in a developmental stage-specific manner to stimulate transcription. Previously, we induced LCR-promoter looping by tethering the self-association domain (SA) of Ldb1 to the β-globin promoter via artificial zinc fingers. Here, we show that targeting the SA to a developmentally silenced embryonic globin gene in adult murine erythroblasts triggers its transcriptional reactivation. This activity depends on the LCR, consistent with an LCR-promoter looping mechanism. Strikingly, targeting the SA to the fetal γ-globin promoter in primary adult human erythroblasts increases γ-globin promoter-LCR contacts, stimulating transcription to approximately 85% of total β-globin synthesis, with a reciprocal reduction in adult β-globin expression. Our findings demonstrate that forced chromatin looping can override a stringent developmental gene expression program and suggest a novel approach to control the balance of globin gene transcription for therapeutic applications.
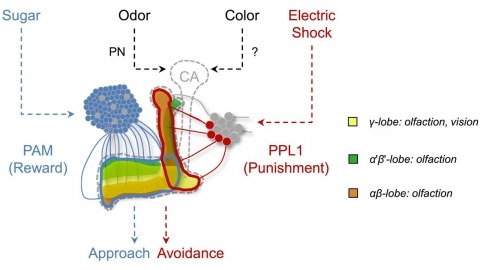
In nature, animals form memories associating reward or punishment with stimuli from different sensory modalities, such as smells and colors. It is unclear, however, how distinct sensory memories are processed in the brain. We established appetitive and aversive visual learning assays for Drosophila that are comparable to the widely used olfactory learning assays. These assays share critical features, such as reinforcing stimuli (sugar reward and electric shock punishment), and allow direct comparison of the cellular requirements for visual and olfactory memories. We found that the same subsets of dopamine neurons drive formation of both sensory memories. Furthermore, distinct yet partially overlapping subsets of mushroom body intrinsic neurons are required for visual and olfactory memories. Thus, our results suggest that distinct sensory memories are processed in a common brain center. Such centralization of related brain functions is an economical design that avoids the repetition of similar circuit motifs.