Filter
Associated Lab
- Dudman Lab (1) Apply Dudman Lab filter
- Hermundstad Lab (1) Apply Hermundstad Lab filter
- Lavis Lab (1) Apply Lavis Lab filter
- Lee (Albert) Lab (1) Apply Lee (Albert) Lab filter
- Looger Lab (2) Apply Looger Lab filter
- Pachitariu Lab (1) Apply Pachitariu Lab filter
- Saalfeld Lab (2) Apply Saalfeld Lab filter
- Scheffer Lab (1) Apply Scheffer Lab filter
- Spruston Lab (1) Apply Spruston Lab filter
- Remove Sternson Lab filter Sternson Lab
- Svoboda Lab (4) Apply Svoboda Lab filter
- Tillberg Lab (3) Apply Tillberg Lab filter
- Turaga Lab (1) Apply Turaga Lab filter
Associated Project Team
Associated Support Team
Publication Date
- 2023 (2) Apply 2023 filter
- 2022 (3) Apply 2022 filter
- 2021 (3) Apply 2021 filter
- 2020 (4) Apply 2020 filter
- 2019 (2) Apply 2019 filter
- 2017 (2) Apply 2017 filter
- 2016 (3) Apply 2016 filter
- 2015 (7) Apply 2015 filter
- 2014 (5) Apply 2014 filter
- 2013 (3) Apply 2013 filter
- 2012 (4) Apply 2012 filter
- 2011 (6) Apply 2011 filter
- 2010 (1) Apply 2010 filter
- 2009 (1) Apply 2009 filter
- 2008 (1) Apply 2008 filter
47 Janelia Publications
Showing 11-20 of 47 resultsMolecular and cellular processes in neurons are critical for sensing and responding to energy deficit states, such as during weight-loss. AGRP neurons are a key hypothalamic population that is activated during energy deficit and increases appetite and weight-gain. Cell type-specific transcriptomics can be used to identify pathways that counteract weight-loss, and here we report high-quality gene expression profiles of AGRP neurons from well-fed and food-deprived young adult mice. For comparison, we also analyzed POMC neurons, an intermingled population that suppresses appetite and body weight. We find that AGRP neurons are considerably more sensitive to energy deficit than POMC neurons. Furthermore, we identify cell type-specific pathways involving endoplasmic reticulum-stress, circadian signaling, ion channels, neuropeptides, and receptors. Combined with methods to validate and manipulate these pathways, this resource greatly expands molecular insight into neuronal regulation of body weight, and may be useful for devising therapeutic strategies for obesity and eating disorders.
Chemogenetics is a technique for obtaining selective pharmacological control over a cell population by expressing an engineered receptor that is selectively activated by an exogenously administered ligand. A promising approach for neuronal modulation involves the use of “Pharmacologically Selective Actuator Modules” (PSAMs); these chemogenetic receptors are selectively activated by ultrapotent “Pharmacologically Selective Effector Molecules” (uPSEMs). To extend the use of PSAM/PSEMs to studies in nonhuman primates it is necessary to thoroughly characterize the efficacy and safety of these tools. We describe the time course and brain penetrance in rhesus monkeys of two compounds with promising binding specificity and efficacy profiles in in vitro studies, uPSEM792 and uPSEM817, after systemic administration. Rhesus macaques received subcutaneous (s.c.) or intravenous (i.v.) administration of uPSEM817(0.064 mg/kg) or uPSEM792 (0.87 mg/kg) and plasma and CSF samples were collected over the course of 48 hours. Both compounds exhibited good brain penetrance, relatively slow washout and negligible conversion to potential metabolites - varenicline or hydroxyvarenicline. In addition, we found that neither of these uPSEMs significantly altered heart rate or sleep. Our results indicate that both compounds are suitable candidates for neuroscience studies using PSAMs in nonhuman primates.
Chemogenetics is a technique for obtaining selective pharmacological control over a cell population by expressing an engineered receptor that is selectively activated by an exogenously administered ligand. A promising approach for neuronal modulation involves the use of "Pharmacologically Selective Actuator Modules" (PSAMs); these chemogenetic receptors are selectively activated by ultrapotent "Pharmacologically Selective Effector Molecules" (uPSEMs). To extend the use of PSAM/PSEMs to studies in nonhuman primates, it is necessary to thoroughly characterize the efficacy and safety of these tools. We describe the time course and brain penetrance in rhesus monkeys of two compounds with promising binding specificity and efficacy profiles in studies, uPSEM792 and uPSEM817, after systemic administration. Rhesus monkeys received subcutaneous (s.c.) or intravenous (i.v.) administration of uPSEM817 (0.064 mg/kg) or uPSEM792 (0.87 mg/kg), and plasma and cerebrospinal fluid samples were collected over 48 h. Both compounds exhibited good brain penetrance, relatively slow washout, and negligible conversion to potential metabolites─varenicline or hydroxyvarenicline. In addition, we found that neither of these uPSEMs significantly altered the heart rate or sleep. Our results indicate that both compounds are suitable candidates for neuroscience studies using PSAMs in nonhuman primates.
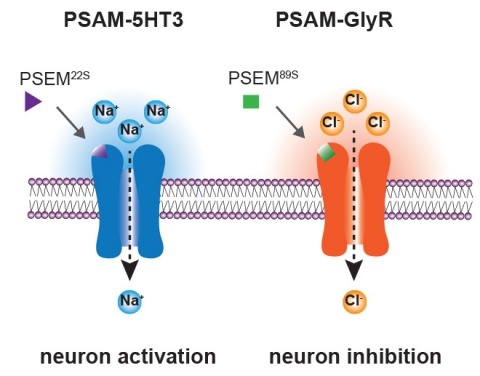
Ionic flux mediates essential physiological and behavioral functions in defined cell populations. Cell type-specific activators of diverse ionic conductances are needed for probing these effects. We combined chemistry and protein engineering to enable the systematic creation of a toolbox of ligand-gated ion channels (LGICs) with orthogonal pharmacologic selectivity and divergent functional properties. The LGICs and their small-molecule effectors were able to activate a range of ionic conductances in genetically specified cell types. LGICs constructed for neuronal perturbation could be used to selectively manipulate neuron activity in mammalian brains in vivo. The diversity of ion channel tools accessible from this approach will be useful for examining the relationship between neuronal activity and animal behavior, as well as for cell biological and physiological applications requiring chemical control of ion conductance.
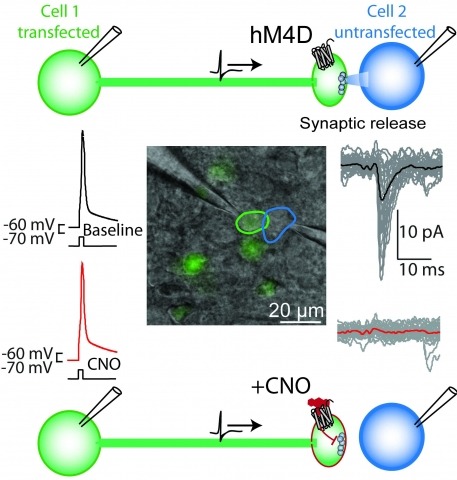
Brain function is mediated by neural circuit connectivity, and elucidating the role of connections is aided by techniques to block their output. We developed cell-type-selective, reversible synaptic inhibition tools for mammalian neural circuits by leveraging G protein signaling pathways to suppress synaptic vesicle release. Here, we find that the pharmacologically selective designer Gi-protein-coupled receptor hM4D is a presynaptic silencer in the presence of its cognate ligand clozapine-N-oxide (CNO). Activation of hM4D signaling sharply reduced synaptic release probability and synaptic current amplitude. To demonstrate the utility of this tool for neural circuit perturbations, we developed an axon-selective hM4D-neurexin variant and used spatially targeted intracranial CNO injections to localize circuit connections from the hypothalamus to the midbrain responsible for feeding behavior. This synaptic silencing approach is broadly applicable for cell-type-specific and axon projection-selective functional analysis of diverse neural circuits.
Chemogenetic technologies enable selective pharmacological control of specific cell populations. An increasing number of approaches have been developed that modulate different signaling pathways. Selective pharmacological control over G protein-coupled receptor signaling, ion channel conductances, protein association, protein stability, and small molecule targeting allows modulation of cellular processes in distinct cell types. Here, we review these chemogenetic technologies and instances of their applications in complex tissues in vivo and ex vivo.
Elucidating the roles of neuronal cell types for physiology and behavior is essential for understanding brain functions. Perturbation of neuron electrical activity can be used to probe the causal relationship between neuronal cell types and behavior. New genetically encoded neuron perturbation tools have been developed for remotely controlling neuron function using small molecules that activate engineered receptors that can be targeted to cell types using genetic methods. Here we describe recent progress for approaches using genetically engineered receptors that selectively interact with small molecules. Called "chemogenetics," receptors with diverse cellular functions have been developed that facilitate the selective pharmacological control over a diverse range of cell-signaling processes, including electrical activity, for molecularly defined cell types. These tools have revealed remarkably specific behavioral physiological influences for molecularly defined cell types that are often intermingled with populations having different or even opposite functions.
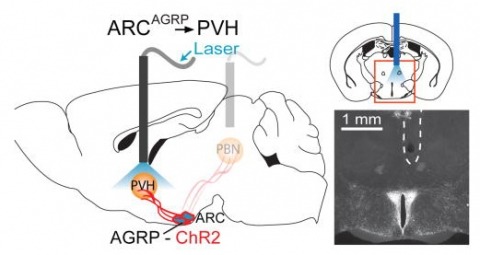
Hunger is a complex behavioural state that elicits intense food seeking and consumption. These behaviours are rapidly recapitulated by activation of starvation-sensitive AGRP neurons, which present an entry point for reverse-engineering neural circuits for hunger. Here we mapped synaptic interactions of AGRP neurons with multiple cell populations in mice and probed the contribution of these distinct circuits to feeding behaviour using optogenetic and pharmacogenetic techniques. An inhibitory circuit with paraventricular hypothalamus (PVH) neurons substantially accounted for acute AGRP neuron-evoked eating, whereas two other prominent circuits were insufficient. Within the PVH, we found that AGRP neurons target and inhibit oxytocin neurons, a small population that is selectively lost in Prader-Willi syndrome, a condition involving insatiable hunger. By developing strategies for evaluating molecularly defined circuits, we show that AGRP neuron suppression of oxytocin neurons is critical for evoked feeding. These experiments reveal a new neural circuit that regulates hunger state and pathways associated with overeating disorders.
Primary aldosteronism (PA) is the most frequent form of secondary hypertension. The identification of germline or somatic mutations in different genes coding for ion channels and defines PA as a channelopathy. These mutations promote activation of calcium signaling, the main trigger for aldosterone biosynthesis.
Determining the spatial organization and morphological characteristics of molecularly defined cell types is a major bottleneck for characterizing the architecture underpinning brain function. We developed Expansion-Assisted Iterative Fluorescence In Situ Hybridization (EASI-FISH) to survey gene expression in brain tissue, as well as a turnkey computational pipeline to rapidly process large EASI-FISH image datasets. EASI-FISH was optimized for thick brain sections (300 μm) to facilitate reconstruction of spatio-molecular domains that generalize across brains. Using the EASI-FISH pipeline, we investigated the spatial distribution of dozens of molecularly defined cell types in the lateral hypothalamic area (LHA), a brain region with poorly defined anatomical organization. Mapping cell types in the LHA revealed nine spatially and molecularly defined subregions. EASI-FISH also facilitates iterative reanalysis of scRNA-seq datasets to determine marker-genes that further dissociated spatial and morphological heterogeneity. The EASI-FISH pipeline democratizes mapping molecularly defined cell types, enabling discoveries about brain organization.