Filter
Associated Lab
- Ahrens Lab (3) Apply Ahrens Lab filter
- Harris Lab (4) Apply Harris Lab filter
- Hermundstad Lab (1) Apply Hermundstad Lab filter
- Jayaraman Lab (7) Apply Jayaraman Lab filter
- Karpova Lab (1) Apply Karpova Lab filter
- Leonardo Lab (1) Apply Leonardo Lab filter
- Remove Looger Lab filter Looger Lab
- Podgorski Lab (2) Apply Podgorski Lab filter
- Rubin Lab (1) Apply Rubin Lab filter
- Remove Schreiter Lab filter Schreiter Lab
- Svoboda Lab (8) Apply Svoboda Lab filter
- Tillberg Lab (1) Apply Tillberg Lab filter
- Turner Lab (1) Apply Turner Lab filter
- Zlatic Lab (1) Apply Zlatic Lab filter
Associated Project Team
Associated Support Team
13 Janelia Publications
Showing 1-10 of 13 resultsEndoplasmic reticulum calcium homeostasis is critical for cellular functions and is disrupted in diverse pathologies including neurodegeneration and cardiovascular disease. Owing to the high concentration of calcium within the ER, studying this subcellular compartment requires tools that are optimized for these conditions. To develop a single-fluorophore genetically encoded calcium indicator for this organelle, we targeted a low affinity variant of GCaMP3 to the ER lumen (GCaMPer (10.19)). A set of viral vectors was constructed to express GCaMPer in human neuroblastoma cells, rat primary cortical neurons, and human induced pluripotent stem cell-derived cardiomyocytes. We observed dynamic changes in GCaMPer (10.19) fluorescence in response to pharmacologic manipulations of the ER calcium store. Additionally, periodic calcium efflux from the ER was observed during spontaneous beating of cardiomyocytes. GCaMPer (10.19) has utility in imaging ER calcium in living cells and providing insight into luminal calcium dynamics under physiologic and pathologic states.
Fluorescent protein-based sensors for detecting neuronal activity have been developed largely based on non-neuronal screening systems. However, the dynamics of neuronal state variables (e.g., voltage, calcium, etc.) are typically very rapid compared to those of non-excitable cells. We developed an electrical stimulation and fluorescence imaging platform based on dissociated rat primary neuronal cultures. We describe its use in testing genetically-encoded calcium indicators (GECIs). Efficient neuronal GECI expression was achieved using lentiviruses containing a neuronal-selective gene promoter. Action potentials (APs) and thus neuronal calcium levels were quantitatively controlled by electrical field stimulation, and fluorescence images were recorded. Images were segmented to extract fluorescence signals corresponding to individual GECI-expressing neurons, which improved sensitivity over full-field measurements. We demonstrate the superiority of screening GECIs in neurons compared with solution measurements. Neuronal screening was useful for efficient identification of variants with both improved response kinetics and high signal amplitudes. This platform can be used to screen many types of sensors with cellular resolution under realistic conditions where neuronal state variables are in relevant ranges with respect to timing and amplitude.
We describe an intensity-based glutamate-sensing fluorescent reporter (iGluSnFR) with signal-to-noise ratio and kinetics appropriate for in vivo imaging. We engineered iGluSnFR in vitro to maximize its fluorescence change, and we validated its utility for visualizing glutamate release by neurons and astrocytes in increasingly intact neurological systems. In hippocampal culture, iGluSnFR detected single field stimulus-evoked glutamate release events. In pyramidal neurons in acute brain slices, glutamate uncaging at single spines showed that iGluSnFR responds robustly and specifically to glutamate in situ, and responses correlate with voltage changes. In mouse retina, iGluSnFR-expressing neurons showed intact light-evoked excitatory currents, and the sensor revealed tonic glutamate signaling in response to light stimuli. In worms, glutamate signals preceded and predicted postsynaptic calcium transients. In zebrafish, iGluSnFR revealed spatial organization of direction-selective synaptic activity in the optic tectum. Finally, in mouse forelimb motor cortex, iGluSnFR expression in layer V pyramidal neurons revealed task-dependent single-spine activity during running.
Calcium imaging with protein-based indicators is widely used to follow neural activity in intact nervous systems, but current protein sensors report neural activity at timescales much slower than electrical signalling and are limited by trade-offs between sensitivity and kinetics. Here we used large-scale screening and structure-guided mutagenesis to develop and optimize several fast and sensitive GCaMP-type indicators. The resulting 'jGCaMP8' sensors, based on the calcium-binding protein calmodulin and a fragment of endothelial nitric oxide synthase, have ultra-fast kinetics (half-rise times of 2 ms) and the highest sensitivity for neural activity reported for a protein-based calcium sensor. jGCaMP8 sensors will allow tracking of large populations of neurons on timescales relevant to neural computation.
Genetically encoded calcium indicators (GECIs) are powerful tools for systems neuroscience. Here we describe red, single-wavelength GECIs, "RCaMPs," engineered from circular permutation of the thermostable red fluorescent protein mRuby. High-resolution crystal structures of mRuby, the red sensor RCaMP, and the recently published red GECI R-GECO1 give insight into the chromophore environments of the Ca(2+)-bound state of the sensors and the engineered protein domain interfaces of the different indicators. We characterized the biophysical properties and performance of RCaMP sensors in vitro and in vivo in Caenorhabditis elegans, Drosophila larvae, and larval zebrafish. Further, we demonstrate 2-color calcium imaging both within the same cell (registering mitochondrial and somatic [Ca(2+)]) and between two populations of cells: neurons and astrocytes. Finally, we perform integrated optogenetics experiments, wherein neural activation via channelrhodopsin-2 (ChR2) or a red-shifted variant, and activity imaging via RCaMP or GCaMP, are conducted simultaneously, with the ChR2/RCaMP pair providing independently addressable spectral channels. Using this paradigm, we measure calcium responses of naturalistic and ChR2-evoked muscle contractions in vivo in crawling C. elegans. We systematically compare the RCaMP sensors to R-GECO1, in terms of action potential-evoked fluorescence increases in neurons, photobleaching, and photoswitching. R-GECO1 displays higher Ca(2+) affinity and larger dynamic range than RCaMP, but exhibits significant photoactivation with blue and green light, suggesting that integrated channelrhodopsin-based optogenetics using R-GECO1 may be subject to artifact. Finally, we create and test blue, cyan, and yellow variants engineered from GCaMP by rational design. This engineered set of chromatic variants facilitates new experiments in functional imaging and optogenetics.
The fluorescent glutamate indicator iGluSnFR enables imaging of neurotransmission with genetic and molecular specificity. However, existing iGluSnFR variants exhibit low in vivo signal-to-noise ratios, saturating activation kinetics and exclusion from postsynaptic densities. Using a multiassay screen in bacteria, soluble protein and cultured neurons, we generated variants with improved signal-to-noise ratios and kinetics. We developed surface display constructs that improve iGluSnFR's nanoscopic localization to postsynapses. The resulting indicator iGluSnFR3 exhibits rapid nonsaturating activation kinetics and reports synaptic glutamate release with decreased saturation and increased specificity versus extrasynaptic signals in cultured neurons. Simultaneous imaging and electrophysiology at individual boutons in mouse visual cortex showed that iGluSnFR3 transients report single action potentials with high specificity. In vibrissal sensory cortex layer 4, we used iGluSnFR3 to characterize distinct patterns of touch-evoked feedforward input from thalamocortical boutons and both feedforward and recurrent input onto L4 cortical neuron dendritic spines.
Calcium imaging with genetically encoded calcium indicators (GECIs) is routinely used to measure neural activity in intact nervous systems. GECIs are frequently used in one of two different modes: to track activity in large populations of neuronal cell bodies, or to follow dynamics in subcellular compartments such as axons, dendrites and individual synaptic compartments. Despite major advances, calcium imaging is still limited by the biophysical properties of existing GECIs, including affinity, signal-to-noise ratio, rise and decay kinetics, and dynamic range. Using structure-guided mutagenesis and neuron-based screening, we optimized the green fluorescent protein-based GECI GCaMP6 for different modes of in vivo imaging. The jGCaMP7 sensors provide improved detection of individual spikes (jGCaMP7s,f), imaging in neurites and neuropil (jGCaMP7b), and tracking large populations of neurons using 2-photon (jGCaMP7s,f) or wide-field (jGCaMP7c) imaging.
Femtosecond lasers at fixed wavelengths above 1,000 nm are powerful, stable and inexpensive, making them promising sources for two-photon microscopy. Biosensors optimized for these wavelengths are needed for both next-generation microscopes and affordable turn-key systems. Here we report jYCaMP1, a yellow variant of the calcium indicator jGCaMP7 that outperforms its parent in mice and flies at excitation wavelengths above 1,000 nm and enables improved two-color calcium imaging with red fluorescent protein-based indicators.
The identification of active neurons and circuits in vivo is a fundamental challenge in understanding the neural basis of behavior. Genetically encoded calcium (Ca(2+)) indicators (GECIs) enable quantitative monitoring of cellular-resolution activity during behavior. However, such indicators require online monitoring within a limited field of view. Alternatively, post hoc staining of immediate early genes (IEGs) indicates highly active cells within the entire brain, albeit with poor temporal resolution. We designed a fluorescent sensor, CaMPARI, that combines the genetic targetability and quantitative link to neural activity of GECIs with the permanent, large-scale labeling of IEGs, allowing a temporally precise "activity snapshot" of a large tissue volume. CaMPARI undergoes efficient and irreversible green-to-red conversion only when elevated intracellular Ca(2+) and experimenter-controlled illumination coincide. We demonstrate the utility of CaMPARI in freely moving larvae of zebrafish and flies, and in head-fixed mice and adult flies.
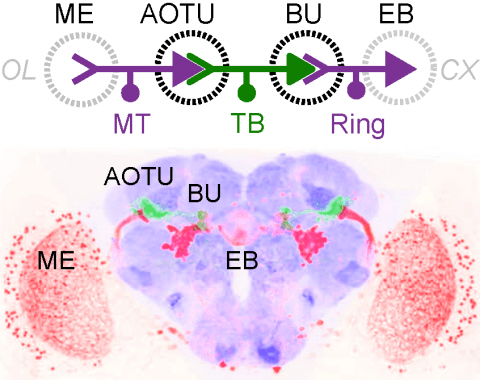
Many animals orient using visual cues, but how a single cue is selected from among many is poorly understood. Here we show that Drosophila ring neurons—central brain neurons implicated in navigation—display visual stimulus selection. Using in vivo two-color two-photon imaging with genetically encoded calcium indicators, we demonstrate that individual ring neurons inherit simple-cell-like receptive fields from their upstream partners. Stimuli in the contralateral visual field suppressed responses to ipsilateral stimuli in both populations. Suppression strength depended on when and where the contralateral stimulus was presented, an effect stronger in ring neurons than in their upstream inputs. This history-dependent effect on the temporal structure of visual responses, which was well modeled by a simple biphasic filter, may determine how visual references are selected for the fly's internal compass. Our approach highlights how two-color calcium imaging can help identify and localize the origins of sensory transformations across synaptically connected neural populations.