Filter
Associated Lab
- Aso Lab (29) Apply Aso Lab filter
- Betzig Lab (1) Apply Betzig Lab filter
- Bock Lab (2) Apply Bock Lab filter
- Branson Lab (7) Apply Branson Lab filter
- Card Lab (5) Apply Card Lab filter
- Clapham Lab (1) Apply Clapham Lab filter
- Dickson Lab (2) Apply Dickson Lab filter
- Druckmann Lab (1) Apply Druckmann Lab filter
- Fetter Lab (1) Apply Fetter Lab filter
- Funke Lab (1) Apply Funke Lab filter
- Harris Lab (3) Apply Harris Lab filter
- Heberlein Lab (1) Apply Heberlein Lab filter
- Hermundstad Lab (2) Apply Hermundstad Lab filter
- Hess Lab (5) Apply Hess Lab filter
- Jayaraman Lab (5) Apply Jayaraman Lab filter
- Lippincott-Schwartz Lab (1) Apply Lippincott-Schwartz Lab filter
- Looger Lab (2) Apply Looger Lab filter
- O'Shea Lab (1) Apply O'Shea Lab filter
- Otopalik Lab (1) Apply Otopalik Lab filter
- Reiser Lab (15) Apply Reiser Lab filter
- Riddiford Lab (1) Apply Riddiford Lab filter
- Romani Lab (1) Apply Romani Lab filter
- Remove Rubin Lab filter Rubin Lab
- Saalfeld Lab (4) Apply Saalfeld Lab filter
- Scheffer Lab (7) Apply Scheffer Lab filter
- Schreiter Lab (1) Apply Schreiter Lab filter
- Simpson Lab (3) Apply Simpson Lab filter
- Singer Lab (1) Apply Singer Lab filter
- Spruston Lab (1) Apply Spruston Lab filter
- Svoboda Lab (3) Apply Svoboda Lab filter
- Truman Lab (4) Apply Truman Lab filter
- Turaga Lab (1) Apply Turaga Lab filter
- Turner Lab (5) Apply Turner Lab filter
Associated Project Team
Associated Support Team
- Electron Microscopy (3) Apply Electron Microscopy filter
- Fly Facility (8) Apply Fly Facility filter
- Janelia Experimental Technology (1) Apply Janelia Experimental Technology filter
- Management Team (1) Apply Management Team filter
- Primary & iPS Cell Culture (1) Apply Primary & iPS Cell Culture filter
- Project Technical Resources (8) Apply Project Technical Resources filter
- Quantitative Genomics (2) Apply Quantitative Genomics filter
- Scientific Computing Software (9) Apply Scientific Computing Software filter
- Scientific Computing Systems (2) Apply Scientific Computing Systems filter
Publication Date
- 2024 (3) Apply 2024 filter
- 2023 (6) Apply 2023 filter
- 2022 (1) Apply 2022 filter
- 2021 (4) Apply 2021 filter
- 2020 (9) Apply 2020 filter
- 2019 (6) Apply 2019 filter
- 2018 (7) Apply 2018 filter
- 2017 (15) Apply 2017 filter
- 2016 (3) Apply 2016 filter
- 2015 (16) Apply 2015 filter
- 2014 (8) Apply 2014 filter
- 2013 (5) Apply 2013 filter
- 2012 (7) Apply 2012 filter
- 2011 (3) Apply 2011 filter
- 2010 (2) Apply 2010 filter
- 2009 (1) Apply 2009 filter
- 2008 (2) Apply 2008 filter
- 2007 (2) Apply 2007 filter
- 2006 (1) Apply 2006 filter
101 Janelia Publications
Showing 81-90 of 101 resultsMotion detection is a fundamental neural computation performed by many sensory systems. In the fly, local motion computation is thought to occur within the first two layers of the visual system, the lamina and medulla. We constructed specific genetic driver lines for each of the 12 neuron classes in the lamina. We then depolarized and hyperpolarized each neuron type and quantified fly behavioral responses to a diverse set of motion stimuli. We found that only a small number of lamina output neurons are essential for motion detection, while most neurons serve to sculpt and enhance these feedforward pathways. Two classes of feedback neurons (C2 and C3), and lamina output neurons (L2 and L4), are required for normal detection of directional motion stimuli. Our results reveal a prominent role for feedback and lateral interactions in motion processing and demonstrate that motion-dependent behaviors rely on contributions from nearly all lamina neuron classes.
Internal state as well as environmental conditions influence choice behavior. The neural circuits underpinning state-dependent behavior remain largely unknown. Carbon dioxide (CO2) is an important olfactory cue for many insects, including mosquitoes, flies, moths, and honeybees [1]. Concentrations of CO2 higher than 0.02% above atmospheric level trigger a strong innate avoidance in the fly Drosophila melanogaster [2, 3]. Here, we show that the mushroom body (MB), a brain center essential for olfactory associative memories [4-6] but thought to be dispensable for innate odor processing [7], is essential for CO2 avoidance behavior only in the context of starvation or in the context of a food-related odor. Consistent with this, CO2 stimulation elicits Ca(2+) influx into the MB intrinsic cells (Kenyon cells: KCs) in vivo. We identify an atypical projection neuron (bilateral ventral projection neuron, biVPN) that connects CO2 sensory input bilaterally to the MB calyx. Blocking synaptic output of the biVPN completely abolishes CO2 avoidance in food-deprived flies, but not in fed flies. These findings show that two alternative neural pathways control innate choice behavior, and they are dependent on the animal’s internal state. In addition, they suggest that, during innate choice behavior, the MB serves as an integration site for internal state and olfactory input.
How neurons form synapses within specific layers remains poorly understood. In the Drosophila medulla, neurons target to discrete layers in a precise fashion. Here we demonstrate that the targeting of L3 neurons to a specific layer occurs in two steps. Initially, L3 growth cones project to a common domain in the outer medulla, overlapping with the growth cones of other neurons destined for a different layer through the redundant functions of N-Cadherin (CadN) and Semaphorin-1a (Sema-1a). CadN mediates adhesion within the domain and Sema-1a mediates repulsion through Plexin A (PlexA) expressed in an adjacent region. Subsequently, L3 growth cones segregate from the domain into their target layer in part through Sema-1a/PlexA-dependent remodeling. Together, our results and recent studies argue that the early medulla is organized into common domains, comprising processes bound for different layers, and that discrete layers later emerge through successive interactions between processes within domains and developing layers.
The analysis of genetic mosaics, in which an animal carries populations of cells with differing genotypes, is a powerful tool for understanding developmental and cell biology. In 1990, we set out to improve the methods used to make genetic mosaics in Drosophila by taking advantage of recently developed approaches for genome engineering. These efforts led to the work described in our 1993 Development paper.
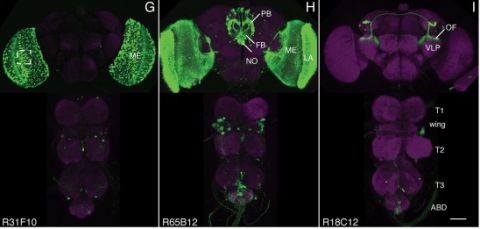
We established a collection of 7,000 transgenic lines of Drosophila melanogaster. Expression of GAL4 in each line is controlled by a different, defined fragment of genomic DNA that serves as a transcriptional enhancer. We used confocal microscopy of dissected nervous systems to determine the expression patterns driven by each fragment in the adult brain and ventral nerve cord. We present image data on 6,650 lines. Using both manual and machine-assisted annotation, we describe the expression patterns in the most useful lines. We illustrate the utility of these data for identifying novel neuronal cell types, revealing brain asymmetry, and describing the nature and extent of neuronal shape stereotypy. The GAL4 lines allow expression of exogenous genes in distinct, small subsets of the adult nervous system. The set of DNA fragments, each driving a documented expression pattern, will facilitate the generation of additional constructs for manipulating neuronal function. synapse was substantially elevated, at the endocytic zone there was no enhanced polymerization activity. We conclude that actin subserves spatially diverse, independently regulated processes throughout spines. Perisynaptic actin forms a uniquely dynamic structure well suited for direct, active regulation of the synapse.
For the overall strategy and methods used to produce the GAL4 lines:
Pfeiffer, B.D., Jenett, A., Hammonds, A.S., Ngo, T.T., Misra, S., Murphy, C., Scully, A., Carlson, J.W., Wan, K.H., Laverty, T.R., Mungall, C., Svirskas, R., Kadonaga, J.T., Doe, C.Q., Eisen, M.B., Celniker, S.E., Rubin, G.M. (2008). Tools for neuroanatomy and neurogenetics in Drosophila. Proc. Natl. Acad. Sci. USA 105, 9715-9720. http://www.pnas.org/content/105/28/9715.full.pdf+html synapse was substantially elevated, at the endocytic zone there was no enhanced polymerization activity. We conclude that actin subserves spatially diverse, independently regulated processes throughout spines. Perisynaptic actin forms a uniquely dynamic structure well suited for direct, active regulation of the synapse.
For data on expression in the embryo:
Manning, L., Purice, M.D., Roberts, J., Pollard, J.L., Bennett, A.L., Kroll, J.R., Dyukareva, A.V., Doan, P.N., Lupton, J.R., Strader, M.E., Tanner, S., Bauer, D., Wilbur, A., Tran, K.D., Laverty, T.R., Pearson, J.C., Crews, S.T., Rubin, G.M., and Doe, C.Q. (2012) Annotated embryonic CNS expression patterns of 5000 GMR GAL4 lines: a resource for manipulating gene expression and analyzing cis-regulatory motifs. Cell Reports (2012) Doi: 10.1016/j.celrep.2012.09.009 http://www.cell.com/cell-reports/fulltext/S2211-1247(12)00290-2 synapse was substantially elevated, at the endocytic zone there was no enhanced polymerization activity. We conclude that actin subserves spatially diverse, independently regulated processes throughout spines. Perisynaptic actin forms a uniquely dynamic structure well suited for direct, active regulation of the synapse.
For data on expression in imaginal discs:
Jory, A., Estella, C., Giorgianni, M.W., Slattery, M., Laverty, T.R., Rubin, G.M., and Mann, R.S. (2012) A survey of 6300 genomic fragments for cis-regulatory activity in the imaginal discs of Drosophila melanogaster. Cell Reports (2012) Doi: 10.1016/j.celrep.2012.09.010 http://www.cell.com/cell-reports/fulltext/S2211-1247(12)00291-4 synapse was substantially elevated, at the endocytic zone there was no enhanced polymerization activity. We conclude that actin subserves spatially diverse, independently regulated processes throughout spines. Perisynaptic actin forms a uniquely dynamic structure well suited for direct, active regulation of the synapse.
For data on expression in the larval nervous system:
Li, H.-H., Kroll, J.R., Lennox, S., Ogundeyi, O., Jeter, J., Depasquale, G., and Truman, J.W. (2013) A GAL4 driver resource for developmental and behavioral studies on the larval CNS of Drosophila. Cell Reports (submitted).
Here, we describe the embryonic central nervous system expression of 5,000 GAL4 lines made using molecularly defined cis-regulatory DNA inserted into a single attP genomic location. We document and annotate the patterns in early embryos when neurogenesis is at its peak, and in older embryos where there is maximal neuronal diversity and the first neural circuits are established. We note expression in other tissues, such as the lateral body wall (muscle, sensory neurons, and trachea) and viscera. Companion papers report on the adult brain and larval imaginal discs, and the integrated data sets are available online (http://www.janelia.org/gal4-gen1). This collection of embryonically expressed GAL4 lines will be valuable for determining neuronal morphology and function. The 1,862 lines expressed in small subsets of neurons (<20/segment) will be especially valuable for characterizing interneuronal diversity and function, because although interneurons comprise the majority of all central nervous system neurons, their gene expression profile and function remain virtually unexplored.
Over 6,000 fragments from the genome of Drosophila melanogaster were analyzed for their ability to drive expression of GAL4 reporter genes in the third-instar larval imaginal discs. About 1,200 reporter genes drove expression in the eye, antenna, leg, wing, haltere, or genital imaginal discs. The patterns ranged from large regions to individual cells. About 75% of the active fragments drove expression in multiple discs; 20% were expressed in ventral, but not dorsal, discs (legs, genital, and antenna), whereas \~{}23% were expressed in dorsal but not ventral discs (wing, haltere, and eye). Several patterns, for example, within the leg chordotonal organ, appeared a surprisingly large number of times. Unbiased searches for DNA sequence motifs suggest candidate transcription factors that may regulate enhancers with shared activities. Together, these expression patterns provide a valuable resource to the community and offer a broad overview of how transcriptional regulatory information is distributed in the Drosophila genome.
Animals approach stimuli that predict a pleasant outcome. After the paired presentation of an odour and a reward, Drosophila melanogaster can develop a conditioned approach towards that odour. Despite recent advances in understanding the neural circuits for associative memory and appetitive motivation, the cellular mechanisms for reward processing in the fly brain are unknown. Here we show that a group of dopamine neurons in the protocerebral anterior medial (PAM) cluster signals sugar reward by transient activation and inactivation of target neurons in intact behaving flies. These dopamine neurons are selectively required for the reinforcing property of, but not a reflexive response to, the sugar stimulus. In vivo calcium imaging revealed that these neurons are activated by sugar ingestion and the activation is increased on starvation. The output sites of the PAM neurons are mainly localized to the medial lobes of the mushroom bodies (MBs), where appetitive olfactory associative memory is formed. We therefore propose that the PAM cluster neurons endow a positive predictive value to the odour in the MBs. Dopamine in insects is known to mediate aversive reinforcement signals. Our results highlight the cellular specificity underlying the various roles of dopamine and the importance of spatially segregated local circuits within the MBs.
The visual system of Drosophila is an excellent model for determining the interactions that direct the differentiation of the nervous system’s many unique cell types. Glia are essential not only in the development of the nervous system, but also in the function of those neurons with which they become associated in the adult. Given their role in visual system development and adult function we need to both accurately and reliably identify the different subtypes of glia, and to relate the glial subtypes in the larval brain to those previously described for the adult. We viewed driver expression in subsets of larval eye disc glia through the earliest stages of pupal development to reveal the counterparts of these cells in the adult. Two populations of glia exist in the lamina, the first neuropil of the adult optic lobe: those that arise from precursors in the eye-disc/optic stalk and those that arise from precursors in the brain. In both cases, a single larval source gives rise to at least three different types of adult glia. Furthermore, analysis of glial cell types in the second neuropil, the medulla, has identified at least four types of astrocyte-like (reticular) glia. Our clarification of the lamina’s adult glia and identification of their larval origins, particularly the respective eye disc and larval brain contributions, begin to define developmental interactions which establish the different subtypes of glia.
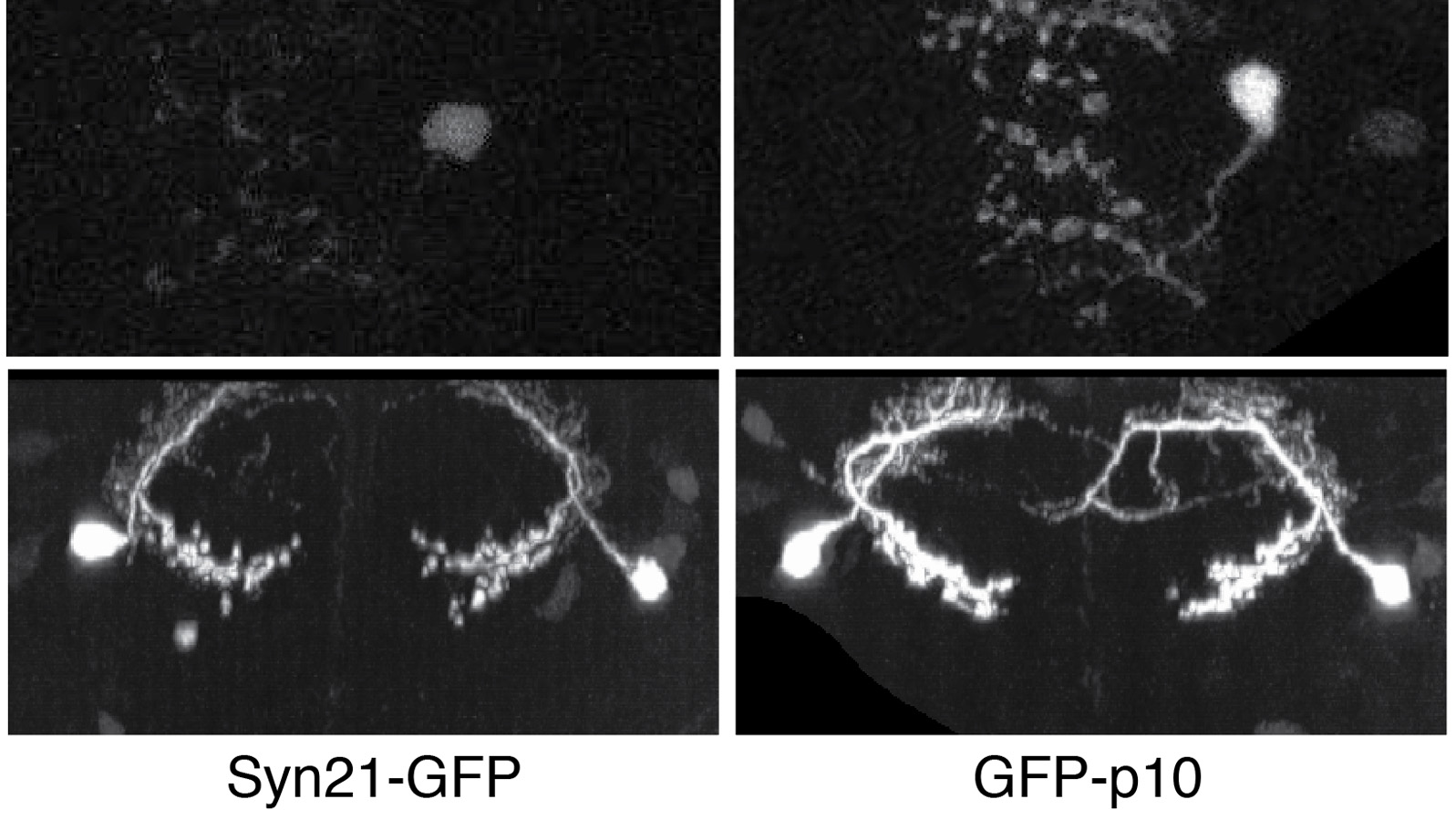
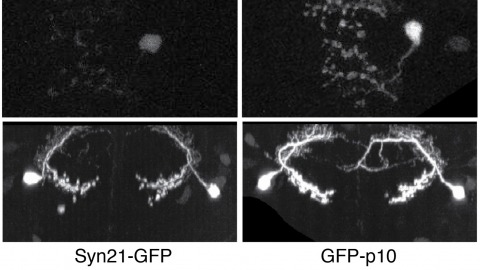
The ability to specify the expression levels of exogenous genes inserted in the genomes of transgenic animals is critical for the success of a wide variety of experimental manipulations. Protein production can be regulated at the level of transcription, mRNA transport, mRNA half-life, or translation efficiency. In this report, we show that several well-characterized sequence elements derived from plant and insect viruses are able to function in Drosophila to increase the apparent translational efficiency of mRNAs by as much as 20-fold. These increases render expression levels sufficient for genetic constructs previously requiring multiple copies to be effective in single copy, including constructs expressing the temperature-sensitive inactivator of neuronal function Shibire(ts1), and for the use of cytoplasmic GFP to image the fine processes of neurons.