Filter
Associated Lab
- Ahrens Lab (1) Apply Ahrens Lab filter
- Aso Lab (2) Apply Aso Lab filter
- Baker Lab (2) Apply Baker Lab filter
- Betzig Lab (5) Apply Betzig Lab filter
- Bock Lab (1) Apply Bock Lab filter
- Branson Lab (3) Apply Branson Lab filter
- Card Lab (1) Apply Card Lab filter
- Cardona Lab (6) Apply Cardona Lab filter
- Chklovskii Lab (1) Apply Chklovskii Lab filter
- Cui Lab (6) Apply Cui Lab filter
- Dickson Lab (3) Apply Dickson Lab filter
- Druckmann Lab (3) Apply Druckmann Lab filter
- Eddy/Rivas Lab (1) Apply Eddy/Rivas Lab filter
- Fetter Lab (1) Apply Fetter Lab filter
- Fitzgerald Lab (1) Apply Fitzgerald Lab filter
- Gonen Lab (7) Apply Gonen Lab filter
- Grigorieff Lab (7) Apply Grigorieff Lab filter
- Harris Lab (3) Apply Harris Lab filter
- Heberlein Lab (9) Apply Heberlein Lab filter
- Hess Lab (3) Apply Hess Lab filter
- Jayaraman Lab (2) Apply Jayaraman Lab filter
- Ji Lab (2) Apply Ji Lab filter
- Kainmueller Lab (2) Apply Kainmueller Lab filter
- Karpova Lab (1) Apply Karpova Lab filter
- Keleman Lab (2) Apply Keleman Lab filter
- Keller Lab (3) Apply Keller Lab filter
- Koay Lab (1) Apply Koay Lab filter
- Lavis Lab (4) Apply Lavis Lab filter
- Lee (Albert) Lab (2) Apply Lee (Albert) Lab filter
- Leonardo Lab (2) Apply Leonardo Lab filter
- Lippincott-Schwartz Lab (12) Apply Lippincott-Schwartz Lab filter
- Looger Lab (13) Apply Looger Lab filter
- Magee Lab (6) Apply Magee Lab filter
- Otopalik Lab (1) Apply Otopalik Lab filter
- Pachitariu Lab (1) Apply Pachitariu Lab filter
- Pastalkova Lab (1) Apply Pastalkova Lab filter
- Pavlopoulos Lab (1) Apply Pavlopoulos Lab filter
- Pedram Lab (1) Apply Pedram Lab filter
- Reiser Lab (1) Apply Reiser Lab filter
- Riddiford Lab (1) Apply Riddiford Lab filter
- Rubin Lab (8) Apply Rubin Lab filter
- Saalfeld Lab (7) Apply Saalfeld Lab filter
- Satou Lab (2) Apply Satou Lab filter
- Scheffer Lab (3) Apply Scheffer Lab filter
- Schreiter Lab (2) Apply Schreiter Lab filter
- Sgro Lab (1) Apply Sgro Lab filter
- Simpson Lab (1) Apply Simpson Lab filter
- Singer Lab (11) Apply Singer Lab filter
- Spruston Lab (4) Apply Spruston Lab filter
- Stern Lab (5) Apply Stern Lab filter
- Sternson Lab (4) Apply Sternson Lab filter
- Svoboda Lab (9) Apply Svoboda Lab filter
- Tervo Lab (1) Apply Tervo Lab filter
- Tjian Lab (1) Apply Tjian Lab filter
- Truman Lab (3) Apply Truman Lab filter
Associated Project Team
Publication Date
- December 2012 (16) Apply December 2012 filter
- November 2012 (16) Apply November 2012 filter
- October 2012 (23) Apply October 2012 filter
- September 2012 (6) Apply September 2012 filter
- August 2012 (13) Apply August 2012 filter
- July 2012 (9) Apply July 2012 filter
- June 2012 (15) Apply June 2012 filter
- May 2012 (13) Apply May 2012 filter
- April 2012 (14) Apply April 2012 filter
- March 2012 (10) Apply March 2012 filter
- February 2012 (19) Apply February 2012 filter
- January 2012 (36) Apply January 2012 filter
- Remove 2012 filter 2012
Type of Publication
190 Publications
Showing 1-10 of 190 resultsThis paper presents a digital neural/EMG telemetry system small enough and lightweight enough to permit recording from insects in flight. It has a measured flight package mass of only 38 mg. This system includes a single-chip telemetry integrated circuit (IC) employing RF power harvesting for battery-free operation, with communication via modulated backscatter in the UHF (902-928 MHz) band. An on-chip 11-bit ADC digitizes 10 neural channels with a sampling rate of 26.1 kSps and 4 EMG channels at 1.63 kSps, and telemeters this data wirelessly to a base station. The companion base station transceiver includes an RF transmitter of +36 dBm (4 W) output power to wirelessly power the telemetry IC, and a digital receiver with a sensitivity of -70 dBm for 10⁻⁵ BER at 5.0 Mbps to receive the data stream from the telemetry IC. The telemetry chip was fabricated in a commercial 0.35 μ m 4M1P (4 metal, 1 poly) CMOS process. The die measures 2.36 × 1.88 mm, is 250 μm thick, and is wire bonded into a flex circuit assembly measuring 4.6 × 6.8 mm.
Fluorescent calcium indicator proteins, such as GCaMP3, allow imaging of activity in genetically defined neuronal populations. GCaMP3 can be expressed using various gene delivery methods, such as viral infection or electroporation. However, these methods are invasive and provide inhomogeneous and nonstationary expression. Here, we developed a genetic reporter mouse, Ai38, which expresses GCaMP3 in a Cre-dependent manner from the ROSA26 locus, driven by a strong CAG promoter. Crossing Ai38 with appropriate Cre mice produced robust GCaMP3 expression in defined cell populations in the retina, cortex, and cerebellum. In the primary visual cortex, visually evoked GCaMP3 signals showed normal orientation and direction selectivity. GCaMP3 signals were rapid, compared with virally expressed GCaMP3 and synthetic calcium indicators. In the retina, Ai38 allowed imaging spontaneous calcium waves in starburst amacrine cells during development, and light-evoked responses in ganglion cells in adult tissue. Our results show that the Ai38 reporter mouse provides a flexible method for targeted expression of GCaMP3.
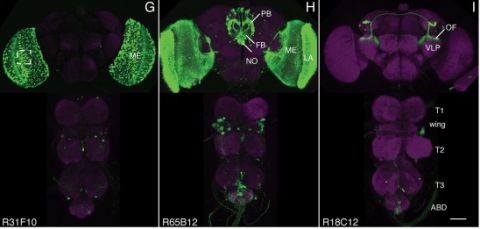
We established a collection of 7,000 transgenic lines of Drosophila melanogaster. Expression of GAL4 in each line is controlled by a different, defined fragment of genomic DNA that serves as a transcriptional enhancer. We used confocal microscopy of dissected nervous systems to determine the expression patterns driven by each fragment in the adult brain and ventral nerve cord. We present image data on 6,650 lines. Using both manual and machine-assisted annotation, we describe the expression patterns in the most useful lines. We illustrate the utility of these data for identifying novel neuronal cell types, revealing brain asymmetry, and describing the nature and extent of neuronal shape stereotypy. The GAL4 lines allow expression of exogenous genes in distinct, small subsets of the adult nervous system. The set of DNA fragments, each driving a documented expression pattern, will facilitate the generation of additional constructs for manipulating neuronal function. synapse was substantially elevated, at the endocytic zone there was no enhanced polymerization activity. We conclude that actin subserves spatially diverse, independently regulated processes throughout spines. Perisynaptic actin forms a uniquely dynamic structure well suited for direct, active regulation of the synapse.
For the overall strategy and methods used to produce the GAL4 lines:
Pfeiffer, B.D., Jenett, A., Hammonds, A.S., Ngo, T.T., Misra, S., Murphy, C., Scully, A., Carlson, J.W., Wan, K.H., Laverty, T.R., Mungall, C., Svirskas, R., Kadonaga, J.T., Doe, C.Q., Eisen, M.B., Celniker, S.E., Rubin, G.M. (2008). Tools for neuroanatomy and neurogenetics in Drosophila. Proc. Natl. Acad. Sci. USA 105, 9715-9720. http://www.pnas.org/content/105/28/9715.full.pdf+html synapse was substantially elevated, at the endocytic zone there was no enhanced polymerization activity. We conclude that actin subserves spatially diverse, independently regulated processes throughout spines. Perisynaptic actin forms a uniquely dynamic structure well suited for direct, active regulation of the synapse.
For data on expression in the embryo:
Manning, L., Purice, M.D., Roberts, J., Pollard, J.L., Bennett, A.L., Kroll, J.R., Dyukareva, A.V., Doan, P.N., Lupton, J.R., Strader, M.E., Tanner, S., Bauer, D., Wilbur, A., Tran, K.D., Laverty, T.R., Pearson, J.C., Crews, S.T., Rubin, G.M., and Doe, C.Q. (2012) Annotated embryonic CNS expression patterns of 5000 GMR GAL4 lines: a resource for manipulating gene expression and analyzing cis-regulatory motifs. Cell Reports (2012) Doi: 10.1016/j.celrep.2012.09.009 http://www.cell.com/cell-reports/fulltext/S2211-1247(12)00290-2 synapse was substantially elevated, at the endocytic zone there was no enhanced polymerization activity. We conclude that actin subserves spatially diverse, independently regulated processes throughout spines. Perisynaptic actin forms a uniquely dynamic structure well suited for direct, active regulation of the synapse.
For data on expression in imaginal discs:
Jory, A., Estella, C., Giorgianni, M.W., Slattery, M., Laverty, T.R., Rubin, G.M., and Mann, R.S. (2012) A survey of 6300 genomic fragments for cis-regulatory activity in the imaginal discs of Drosophila melanogaster. Cell Reports (2012) Doi: 10.1016/j.celrep.2012.09.010 http://www.cell.com/cell-reports/fulltext/S2211-1247(12)00291-4 synapse was substantially elevated, at the endocytic zone there was no enhanced polymerization activity. We conclude that actin subserves spatially diverse, independently regulated processes throughout spines. Perisynaptic actin forms a uniquely dynamic structure well suited for direct, active regulation of the synapse.
For data on expression in the larval nervous system:
Li, H.-H., Kroll, J.R., Lennox, S., Ogundeyi, O., Jeter, J., Depasquale, G., and Truman, J.W. (2013) A GAL4 driver resource for developmental and behavioral studies on the larval CNS of Drosophila. Cell Reports (submitted).
Habituation is a form of non-associative learning that enables animals to reduce their reaction to repeated harmless stimuli. When exposed to ethanol vapor, Drosophila show an olfactory-mediated startle response characterized by a transient increase in locomotor activity. Upon repeated exposures, this olfactory startle attenuates with the characteristics of habituation. Here we describe the results of a genetic screen to identify olfactory startle habituation (OSH) mutants. One mutation is a transcript specific allele of foraging (for) encoding a cGMP-dependent kinase. We show this allele of for reduces expression of a for-T1 isoform expressed in the head and functions normally to inhibit OSH. We localize for-T1 function to a limited set of neurons that include olfactory receptor neurons (ORNs) and the mushroom body (MB). Overexpression of for-T1 in ORNs inhibits OSH, an effect also seen upon synaptic silencing of the ORNs; for-T1 may therefore function in ORNs to decrease synaptic release upon repeated exposure to ethanol vapor. Overall, this work contributes to our understanding of the genes and neurons underlying olfactory habituation in Drosophila.
Since the original identification of GFP from jellyfish and corals, the genetically encoded fluorescent proteins have become mainstream indicators for imaging. Functionally homologous candidates exist in more highly evolved bioluminescent invertebrates, including echinoderms. For example, in brittlestars, stimulus-evoked bioluminescence is transient, lasting seconds, and emanates from specialized cells (photocytes). Prior to light emission, we observe little or no green fluorescence. However, concurrent with light emission, an intense green, calcium-dependent fluorescence develops that persists indefinitely.
Although the diversity of cortical interneuron electrical properties is well recognized, the number of distinct electrical types (e-types) is still a matter of debate. Recently, descriptions of interneuron variability were standardized by multiple laboratories on the basis of a subjective classification scheme as set out by the Petilla convention (Petilla Interneuron Nomenclature Group, PING). Here, we present a quantitative, statistical analysis of a database of nearly five hundred neurons manually annotated according to the PING nomenclature. For each cell, 38 features were extracted from responses to suprathreshold current stimuli and statistically analyzed to examine whether cortical interneurons subdivide into e-types. We showed that the partitioning into different e-types is indeed the major component of data variability. The analysis suggests refining the PING e-type classification to be hierarchical, whereby most variability is first captured within a coarse subpartition, and then subsequently divided into finer subpartitions. The coarse partition matches the well-known partitioning of interneurons into fast spiking and adapting cells. Finer subpartitions match the burst, continuous, and delayed subtypes. Additionally, our analysis enabled the ranking of features according to their ability to differentiate among e-types. We showed that our quantitative e-type assignment is more than 90% accurate and manages to catch several human errors.
Early stages of visual processing are thought to decorrelate, or whiten, the incoming temporally varying signals. Because the typical correlation time of natural stimuli, as well as the extent of temporal receptive fields of lateral geniculate nucleus (LGN) neurons, is much greater than neuronal time constants, such decorrelation must be done in stages combining contributions of multiple neurons. We propose to model temporal decorrelation in the visual pathway with the lattice filter, a signal processing device for stage-wise decorrelation of temporal signals. The stage-wise architecture of the lattice filter maps naturally onto the visual pathway (photoreceptors -> bipolar cells -> retinal ganglion cells -> LGN) and its filter weights can be learned using Hebbian rules in a stage-wise sequential manner. Moreover, predictions of neural activity from the lattice filter model are consistent with physiological measurements in LGN neurons and fruit fly second-order visual neurons. Therefore, the lattice filter model is a useful abstraction that may help unravel visual system function.
Early stages of sensory systems face the challenge of compressing information from numerous receptors onto a much smaller number of projection neurons, a so called communication bottleneck. To make more efficient use of limited bandwidth, compression may be achieved using predictive coding, whereby predictable, or redundant, components of the stimulus are removed. In the case of the retina, Srinivasan et al. (1982) suggested that feedforward inhibitory connections subtracting a linear prediction generated from nearby receptors implement such compression, resulting in biphasic center-surround receptive fields. However, feedback inhibitory circuits are common in early sensory circuits and furthermore their dynamics may be nonlinear. Can such circuits implement predictive coding as well? Here, solving the transient dynamics of nonlinear reciprocal feedback circuits through analogy to a signal-processing algorithm called linearized Bregman iteration we show that nonlinear predictive coding can be implemented in an inhibitory feedback circuit. In response to a step stimulus, interneuron activity in time constructs progressively less sparse but more accurate representations of the stimulus, a temporally evolving prediction. This analysis provides a powerful theoretical framework to interpret and understand the dynamics of early sensory processing in a variety of physiological experiments and yields novel predictions regarding the relation between activity and stimulus statistics.
Computing sparse redundant representations is an important problem in both applied mathematics and neuroscience. In many applications, this problem must be solved in an energy-efficient way. Here, we propose a hybrid distributed algorithm (HDA), which solves this problem on a network of simple nodes communicating by low-bandwidth channels. HDA nodes perform both gradient-descent-like steps on analog internal variables and coordinate-descent-like steps via quantized external variables communicated to each other. Interestingly, the operation is equivalent to a network of integrate-and-fire neurons, suggesting that HDA may serve as a model of neural computation. We show that the numerical performance of HDA is on par with existing algorithms. In the asymptotic regime, the representation error of HDA decays with time, t, as 1/t. HDA is stable against time-varying noise; specifically, the representation error decays as 1/√t for gaussian white noise.
View Publication PageThe standard approach for single-sequence RNA secondary structure prediction uses a nearest-neighbor thermodynamic model with several thousand experimentally determined energy parameters. An attractive alternative is to use statistical approaches with parameters estimated from growing databases of structural RNAs. Good results have been reported for discriminative statistical methods using complex nearest-neighbor models, including CONTRAfold, Simfold, and ContextFold. Little work has been reported on generative probabilistic models (stochastic context-free grammars [SCFGs]) of comparable complexity, although probabilistic models are generally easier to train and to use. To explore a range of probabilistic models of increasing complexity, and to directly compare probabilistic, thermodynamic, and discriminative approaches, we created TORNADO, a computational tool that can parse a wide spectrum of RNA grammar architectures (including the standard nearest-neighbor model and more) using a generalized super-grammar that can be parameterized with probabilities, energies, or arbitrary scores. By using TORNADO, we find that probabilistic nearest-neighbor models perform comparably to (but not significantly better than) discriminative methods. We find that complex statistical models are prone to overfitting RNA structure and that evaluations should use structurally nonhomologous training and test data sets. Overfitting has affected at least one published method (ContextFold). The most important barrier to improving statistical approaches for RNA secondary structure prediction is the lack of diversity of well-curated single-sequence RNA secondary structures in current RNA databases.