Filter
Associated Lab
- Aso Lab (1) Apply Aso Lab filter
- Baker Lab (3) Apply Baker Lab filter
- Betzig Lab (7) Apply Betzig Lab filter
- Bock Lab (1) Apply Bock Lab filter
- Cui Lab (3) Apply Cui Lab filter
- Dickson Lab (1) Apply Dickson Lab filter
- Dudman Lab (1) Apply Dudman Lab filter
- Eddy/Rivas Lab (5) Apply Eddy/Rivas Lab filter
- Fetter Lab (2) Apply Fetter Lab filter
- Gonen Lab (1) Apply Gonen Lab filter
- Hess Lab (2) Apply Hess Lab filter
- Jayaraman Lab (2) Apply Jayaraman Lab filter
- Ji Lab (2) Apply Ji Lab filter
- Keller Lab (2) Apply Keller Lab filter
- Lavis Lab (5) Apply Lavis Lab filter
- Lee (Albert) Lab (1) Apply Lee (Albert) Lab filter
- Leonardo Lab (2) Apply Leonardo Lab filter
- Looger Lab (7) Apply Looger Lab filter
- Magee Lab (1) Apply Magee Lab filter
- Menon Lab (3) Apply Menon Lab filter
- Murphy Lab (1) Apply Murphy Lab filter
- Reiser Lab (2) Apply Reiser Lab filter
- Riddiford Lab (1) Apply Riddiford Lab filter
- Rubin Lab (3) Apply Rubin Lab filter
- Scheffer Lab (2) Apply Scheffer Lab filter
- Schreiter Lab (2) Apply Schreiter Lab filter
- Simpson Lab (3) Apply Simpson Lab filter
- Singer Lab (1) Apply Singer Lab filter
- Sternson Lab (6) Apply Sternson Lab filter
- Svoboda Lab (7) Apply Svoboda Lab filter
- Tjian Lab (2) Apply Tjian Lab filter
- Truman Lab (1) Apply Truman Lab filter
- Zlatic Lab (1) Apply Zlatic Lab filter
- Zuker Lab (2) Apply Zuker Lab filter
Associated Project Team
Publication Date
- December 2011 (10) Apply December 2011 filter
- November 2011 (8) Apply November 2011 filter
- October 2011 (8) Apply October 2011 filter
- September 2011 (8) Apply September 2011 filter
- August 2011 (9) Apply August 2011 filter
- July 2011 (5) Apply July 2011 filter
- June 2011 (10) Apply June 2011 filter
- May 2011 (6) Apply May 2011 filter
- April 2011 (5) Apply April 2011 filter
- March 2011 (6) Apply March 2011 filter
- February 2011 (8) Apply February 2011 filter
- January 2011 (15) Apply January 2011 filter
- Remove 2011 filter 2011
Type of Publication
- Remove Janelia filter Janelia
98 Publications
Showing 81-90 of 98 resultsThe phnD gene of Escherichia coli encodes the periplasmic binding protein of the phosphonate (Pn) uptake and utilization pathway. We have crystallized and determined structures of E. coli PhnD (EcPhnD) in the absence of ligand and in complex with the environmentally abundant 2-aminoethylphosphonate (2AEP). Similar to other bacterial periplasmic binding proteins, 2AEP binds near the center of mass of EcPhnD in a cleft formed between two lobes. Comparison of the open, unliganded structure with the closed 2AEP-bound structure shows that the two lobes pivot around a hinge by \~{}70° between the two states. Extensive hydrogen bonding and electrostatic interactions stabilize 2AEP, which binds to EcPhnD with low nanomolar affinity. These structures provide insight into Pn uptake by bacteria and facilitated the rational design of high signal-to-noise Pn biosensors based on both coupled small-molecule dyes and autocatalytic fluorescent proteins.
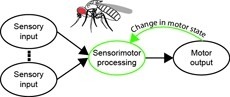
Sensorimotor integration is a field rich in theory backed by a large body of psychophysical evidence. Relating the underlying neural circuitry to these theories has, however, been more challenging. With a wide array of complex behaviors coordinated by their small brains, insects provide powerful model systems to study key features of sensorimotor integration at a mechanistic level. Insect neural circuits perform both hard-wired and learned sensorimotor transformations. They modulate their neural processing based on both internal variables, such as the animal’s behavioral state, and external ones, such as the time of day. Here we present some studies using insect model systems that have produced insights, at the level of individual neurons, about sensorimotor integration and the various ways in which it can be modified by context.
The neural underpinnings of sensorimotor integration are best studied in the context of well-characterized behavior. A rich trove of Drosophila behavioral genetics research offers a variety of well-studied behaviors and candidate brain regions that can form the bases of such studies. The development of tools to perform in vivo physiology from the Drosophila brain has made it possible to monitor activity in defined neurons in response to sensory stimuli. More recently still, it has become possible to perform recordings from identified neurons in the brain of head-fixed flies during walking or flight behaviors. In this chapter, we discuss how experiments that simultaneously monitor behavior and physiology in Drosophila can be combined with other techniques to produce testable models of sensorimotor circuit function.
Recent findings implicate alternate core promoter recognition complexes in regulating cellular differentiation. Here we report a spatial segregation of the alternative core factor TAF3, but not canonical TFIID subunits, away from the nuclear periphery, where the key myogenic gene MyoD is preferentially localized in myoblasts. This segregation is correlated with the differential occupancy of TAF3 versus TFIID at the MyoD promoter. Loss of this segregation by modulating either the intranuclear location of the MyoD gene or TAF3 protein leads to altered TAF3 occupancy at the MyoD promoter. Intriguingly, in differentiated myotubes, the MyoD gene is repositioned to the nuclear interior, where TAF3 resides. The specific high-affinity recognition of H3K4Me3 by the TAF3 PHD (plant homeodomain) finger appears to be required for the sequestration of TAF3 to the nuclear interior. We suggest that intranuclear sequestration of core transcription components and their target genes provides an additional mechanism for promoter selectivity during differentiation.
Commentary: Jie Yao in Bob Tijan’s lab used a combination of confocal microscopy and dual label PALM in thin sections cut from resin-embedded cells to show that certain core transcription components and their target genes are spatially segregated in myoblasts, but not in differentiated myotubes, suggesting that such spatial segregation may play a role in guiding cellular differentiation.
Three-dimensional (3D) structured-illumination microscopy (SIM) can double the lateral and axial resolution of a wide-field fluorescence microscope but has been too slow for live imaging. Here we apply 3D SIM to living samples and record whole cells at up to 5 s per volume for >50 time points with 120-nm lateral and 360-nm axial resolution. We demonstrate the technique by imaging microtubules in S2 cells and mitochondria in HeLa cells.
A fundamental objective in molecular biology is to understand how DNA is organized in concert with various proteins, RNA, and biological membranes. Mitochondria maintain and express their own DNA (mtDNA), which is arranged within structures called nucleoids. Their functions, dimensions, composition, and precise locations relative to other mitochondrial structures are poorly defined. Superresolution fluorescence microscopy techniques that exceed the previous limits of imaging within the small and highly compartmentalized mitochondria have been recently developed. We have improved and employed both two- and three-dimensional applications of photoactivated localization microscopy (PALM and iPALM, respectively) to visualize the core dimensions and relative locations of mitochondrial nucleoids at an unprecedented resolution. PALM reveals that nucleoids differ greatly in size and shape. Three-dimensional volumetric analysis indicates that, on average, the mtDNA within ellipsoidal nucleoids is extraordinarily condensed. Two-color PALM shows that the freely diffusible mitochondrial matrix protein is largely excluded from the nucleoid. In contrast, nucleoids are closely associated with the inner membrane and often appear to be wrapped around cristae or crista-like inner membrane invaginations. Determinations revealing high packing density, separation from the matrix, and tight association with the inner membrane underscore the role of mechanisms that regulate access to mtDNA and that remain largely unknown.
Phenolic fluorophores such as fluorescein, Tokyo Green, resorufin, and their derivatives are workhorses of biological science. Acylating the phenolic hydroxyl group(s) in these fluorophores masks their fluorescence. The ensuing ester is a substrate for cellular esterases, which can restore fluorescence. These esters are, however, notoriously unstable to hydrolysis, severely compromising their utility. The acetoxymethyl (AM) group is an esterase-sensitive motif that can mask polar functionalities in small molecules. Here, we report on the use of AM ether groups to mask phenolic fluorophores. The resulting profluorophores have a desirable combination of low background fluorescence, high chemical stability, and high enzymatic reactivity, both in vitro and in cellulo. These simple phenyl ether-based profluorophores could supplement or supplant the use of phenyl esters for imaging biochemical and biological systems.
A unified, convenient, and efficient strategy for the preparation of rhodamines and N,N’-diacylated rhodamines has been developed. Fluorescein ditriflates were found to undergo palladium-catalyzed C-N cross-coupling with amines, amides, carbamates, and other nitrogen nucleophiles to provide direct access to known and novel rhodamine derivatives, including fluorescent dyes, quenchers, and latent fluorophores.
Various members of the family of BTB/POZ zinc-finger transcription factors influence patterns of dendritic branching. One such member, Broad, is notable because its BrZ3 isoform is widely expressed in Drosophila in immature neurons around the time of arbor outgrowth. We used the metamorphic remodeling of an identified sensory neuron, the dorsal bipolar dendrite sensory neuron (dbd), to examine the effects of BrZ3 expression on the extent and pattern of dendrite growth during metamorphosis.