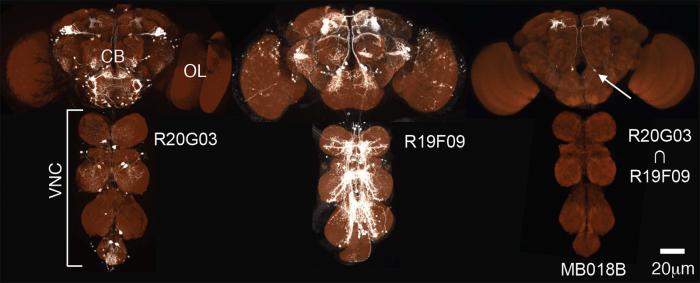
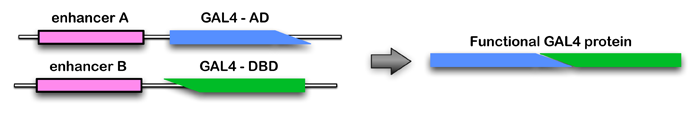
The Generation 1 GAL4 lines almost always express in more than one cell type. However, we found that it is possible to achieve cell-type specificity if expression is limited to the overlap in the expression patterns of two Generation 1 lines. The approach we prefer for carrying out such positive intersections is the split-GAL4 method (Luan et al. 2006; Pfeiffer et al, 2010). In this method, the coding region of the GAL4 gene is split and one enhancer is used to drive the expression of the GAL4 activating domain (AD) and a second enhancer is used to drive the expression of the DNA-binding domain (DBD). The DBD and AD proteins are not able to promote gene expression; only cells where both enhancers are active produce functional GAL4 protein (see diagram and example).
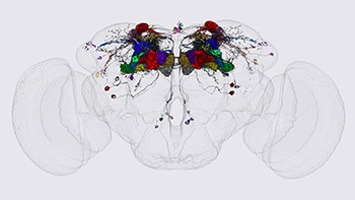
An example of use of the split-GAL4 method to generate a driver liver line specific for one cell type of Mushroom Body output neurons is shown below.
We believe we will be able to generate split-GAL4 lines that are specific for the vast majority of cell types in the fly brain. We have already made several hundred such lines and this effort is a major ongoing project of the FlyLight Project Team in collaboration with a number of laboratories from around the world who have anatomical expertise in particular brain areas. Data on split-GAL4 lines are being added to a database on a regular basis.
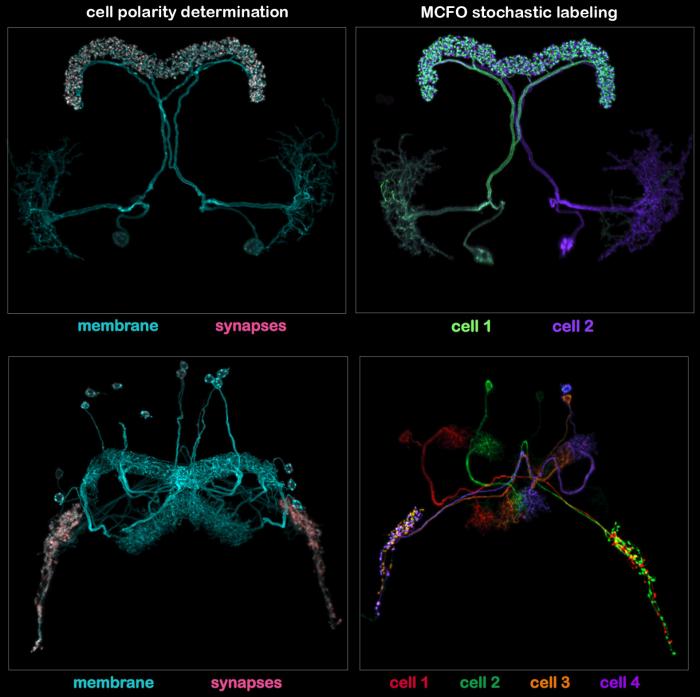
Once split-GAL4 lines are generated they are characterized by two imaging protocol. In the first protocol (illustrated in the left panels of the figure shown below), two UAS-constructs are expressed. One is myristolated and labels the cell membrane; the other is targeted to synapses by fusion to a synaptic protein (Synaptotagmin). In the second protocol (illustrated in the right panels) individual cells are labeled in different colors using the MCFO stochastic labeling method. The imaging protocol also includes a reference marker (the anti-Brp monoclonal nc82) not shown in these examples.
Images are of split-GAL4 lines for central complex cell types that innervate the protocerebral bridge (Tanya Wolff and FlyLight); see text for explanation.
Pfeiffer BD, et al