Filter
Associated Lab
- Aso Lab (30) Apply Aso Lab filter
- Betzig Lab (1) Apply Betzig Lab filter
- Bock Lab (2) Apply Bock Lab filter
- Branson Lab (9) Apply Branson Lab filter
- Card Lab (5) Apply Card Lab filter
- Clapham Lab (1) Apply Clapham Lab filter
- Dickson Lab (2) Apply Dickson Lab filter
- Druckmann Lab (1) Apply Druckmann Lab filter
- Fetter Lab (1) Apply Fetter Lab filter
- Funke Lab (2) Apply Funke Lab filter
- Harris Lab (3) Apply Harris Lab filter
- Heberlein Lab (2) Apply Heberlein Lab filter
- Hermundstad Lab (2) Apply Hermundstad Lab filter
- Hess Lab (6) Apply Hess Lab filter
- Jayaraman Lab (7) Apply Jayaraman Lab filter
- Lippincott-Schwartz Lab (1) Apply Lippincott-Schwartz Lab filter
- Looger Lab (2) Apply Looger Lab filter
- O'Shea Lab (1) Apply O'Shea Lab filter
- Otopalik Lab (1) Apply Otopalik Lab filter
- Reiser Lab (17) Apply Reiser Lab filter
- Riddiford Lab (1) Apply Riddiford Lab filter
- Romani Lab (2) Apply Romani Lab filter
- Rubin Lab (149) Apply Rubin Lab filter
- Saalfeld Lab (5) Apply Saalfeld Lab filter
- Scheffer Lab (8) Apply Scheffer Lab filter
- Schreiter Lab (1) Apply Schreiter Lab filter
- Simpson Lab (3) Apply Simpson Lab filter
- Singer Lab (1) Apply Singer Lab filter
- Spruston Lab (1) Apply Spruston Lab filter
- Stern Lab (1) Apply Stern Lab filter
- Svoboda Lab (3) Apply Svoboda Lab filter
- Tjian Lab (1) Apply Tjian Lab filter
- Truman Lab (4) Apply Truman Lab filter
- Turaga Lab (1) Apply Turaga Lab filter
- Turner Lab (5) Apply Turner Lab filter
- Zuker Lab (1) Apply Zuker Lab filter
Associated Project Team
- CellMap (1) Apply CellMap filter
- Fly Functional Connectome (4) Apply Fly Functional Connectome filter
- Fly Olympiad (3) Apply Fly Olympiad filter
- FlyEM (12) Apply FlyEM filter
- FlyLight (20) Apply FlyLight filter
- GENIE (1) Apply GENIE filter
- Transcription Imaging (1) Apply Transcription Imaging filter
Publication Date
- 2026 (2) Apply 2026 filter
- 2025 (8) Apply 2025 filter
- 2024 (4) Apply 2024 filter
- 2023 (5) Apply 2023 filter
- 2022 (1) Apply 2022 filter
- 2021 (4) Apply 2021 filter
- 2020 (9) Apply 2020 filter
- 2019 (6) Apply 2019 filter
- 2018 (7) Apply 2018 filter
- 2017 (15) Apply 2017 filter
- 2016 (3) Apply 2016 filter
- 2015 (16) Apply 2015 filter
- 2014 (9) Apply 2014 filter
- 2013 (5) Apply 2013 filter
- 2012 (8) Apply 2012 filter
- 2011 (4) Apply 2011 filter
- 2010 (4) Apply 2010 filter
- 2009 (2) Apply 2009 filter
- 2008 (4) Apply 2008 filter
- 2007 (2) Apply 2007 filter
- 2006 (1) Apply 2006 filter
- 2002 (1) Apply 2002 filter
- 2000 (2) Apply 2000 filter
- 1999 (1) Apply 1999 filter
- 1997 (1) Apply 1997 filter
- 1995 (2) Apply 1995 filter
- 1994 (2) Apply 1994 filter
- 1993 (2) Apply 1993 filter
- 1992 (1) Apply 1992 filter
- 1991 (2) Apply 1991 filter
- 1990 (3) Apply 1990 filter
- 1989 (2) Apply 1989 filter
- 1987 (2) Apply 1987 filter
- 1986 (1) Apply 1986 filter
- 1985 (1) Apply 1985 filter
- 1984 (1) Apply 1984 filter
- 1983 (1) Apply 1983 filter
- 1982 (2) Apply 1982 filter
- 1981 (1) Apply 1981 filter
- 1979 (1) Apply 1979 filter
- 1973 (1) Apply 1973 filter
Type of Publication
149 Publications
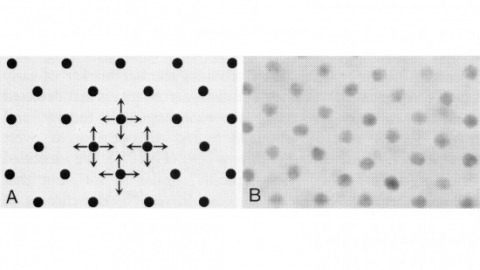
Showing 111-120 of 149 resultsIn the development of multicellular organisms a diversity of cell types differentiate at specific positions. Spacing patterns, in which an array of two or more cell types forms from a uniform field of cells, are a common feature of development. Identical precursor cells may adopt different fates because of competition and inhibition between them. Such a pattern in the developing Drosophila eye is the evenly spaced array of R8 cells, around which other cell types are subsequently recruited. Genetic studies suggest that the scabrous mutation disrupts a signal produced by R8 cells that inhibits other cells from also becoming R8 cells. The scabrous locus was cloned, and it appears to encode a secreted protein partly related to the beta and gamma chains of fibrinogen. It is proposed that the sca locus encodes a lateral inhibitor of R8 differentiation. The roles of the Drosophila EGF-receptor homologue (DER) and Notch genes in this process were also investigated.
Visual systems can exploit spatial correlations in the visual scene by using retinotopy. However, retinotopy is often lost, such as when visual pathways are integrated with other sensory modalities. How is spatial information processed outside of strictly visual brain areas? Here, we focused on visual looming responsive LC6 cells in , a population whose dendrites collectively cover the visual field, but whose axons form a single glomerulus-a structure without obvious retinotopic organization-in the central brain. We identified multiple cell types downstream of LC6 in the glomerulus and found that they more strongly respond to looming in different portions of the visual field, unexpectedly preserving spatial information. Through EM reconstruction of all LC6 synaptic inputs to the glomerulus, we found that LC6 and downstream cell types form circuits within the glomerulus that enable spatial readout of visual features and contralateral suppression-mechanisms that transform visual information for behavioral control.
Building a sizable, complex brain requires both cellular expansion and diversification. One mechanism to achieve these goals is production of multiple transiently amplifying intermediate neural progenitors (INPs) from a single neural stem cell. Like mammalian neural stem cells, Drosophila type II neuroblasts utilize INPs to produce neurons and glia. Within a given lineage, the consecutively born INPs produce morphologically distinct progeny, presumably due to differential inheritance of temporal factors. To uncover the underlying temporal fating mechanisms, we profiled type II neuroblasts' transcriptome across time. Our results reveal opposing temporal gradients of Imp and Syp RNA-binding proteins (descending and ascending, respectively). Maintaining high Imp throughout serial INP production expands the number of neurons and glia with early temporal fate at the expense of cells with late fate. Conversely, precocious upregulation of Syp reduces the number of cells with early fate. Furthermore, we reveal that the transcription factor Seven-up initiates progression of the Imp/Syp gradients. Interestingly, neuroblasts that maintain initial Imp/Syp levels can still yield progeny with a small range of early fates. We therefore propose that the Seven-up-initiated Imp/Syp gradients create coarse temporal windows within type II neuroblasts to pattern INPs, which subsequently undergo fine-tuned subtemporal patterning.
Neuronal connectivity in the circadian clock network is essential for robust endogenous timekeeping. In the circadian clock network, the small ventral lateral neurons (sLNs) serve as critical pacemakers. Peptidergic communication mediated by the neuropeptide (PDF), released by sLNs, has been well characterized. In contrast, little is known about the role of the synaptic connections that sLNs form with downstream neurons. Connectomic analyses revealed that the sLNs form strong synaptic connections with previously uncharacterized neurons called superior lateral protocerebrum 316 (SLP316). Here, we show that silencing the synaptic output from the SLP316 neurons via tetanus toxin expression shortened the free-running period, whereas hyperexciting them by expressing the bacterial voltage-gated sodium channel resulted in period lengthening. Under light-dark cycles, silencing SLP316 neurons caused lower daytime activity and higher daytime sleep. Our results reveal that the main postsynaptic partners of key pacemaker neurons are a nonclock neuronal cell type that regulates the timing of sleep and activity.
Color and polarization provide complementary information about the world and are detected by specialized photoreceptors. However, the downstream neural circuits that process these distinct modalities are incompletely understood in any animal. Using electron microscopy, we have systematically reconstructed the synaptic targets of the photoreceptors specialized to detect color and skylight polarization in Drosophila, and we have used light microscopy to confirm many of our findings. We identified known and novel downstream targets that are selective for different wavelengths or polarized light, and followed their projections to other areas in the optic lobes and the central brain. Our results revealed many synapses along the photoreceptor axons between brain regions, new pathways in the optic lobes, and spatially segregated projections to central brain regions. Strikingly, photoreceptors in the polarization-sensitive dorsal rim area target fewer cell types, and lack strong connections to the lobula, a neuropil involved in color processing. Our reconstruction identifies shared wiring and modality-specific specializations for color and polarization vision, and provides a comprehensive view of the first steps of the pathways processing color and polarized light inputs.
Neuronal connectivity in the circadian clock network is essential for robust endogenous timekeeping. In the Drosophila circadian clock network, the four pairs of small ventral lateral neurons (sLNvs) serve as main pacemakers. Peptidergic communication via sLNv, which release the key output neuropeptide, Pigment Dispersing Factor (PDF), has been well characterized. In the absence of PDF, flies become largely arrhythmic, similar to the phenotype associated with the loss of the mammalian circadian peptide, VIP. In contrast, little is known about the role of the synaptic connections that sLNvs form with downstream neurons. Connectomic analyses revealed that despite their role as key pacemaker neurons within the clock network, the sLNvs form few connections with other clock neurons. However, they form strong synaptic connections with a small group of previously uncharacterized neurons, SLP316, which in turn synapse onto dorsal clock neurons. Here, we show that silencing SLP316 neurons via tetanus toxin (TNT) expression shortens the free-running period, whereas hyper-exciting them by expressing the constitutively open Na[+] channel, NaChBac, results in period lengthening. Under light-dark cycles, silencing SLP316 neurons also causes lower daytime activity and higher daytime sleep. Our results revealed that the main postsynaptic partners of the Drosophila pacemaker neurons are a non-clock neuronal cell type that regulates the timing of sleep and activity.
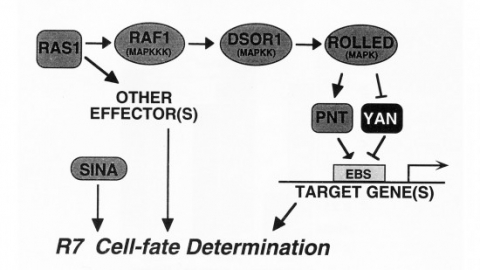
We show that the activities of two Ets-related transcription factors required for normal eye development in Drosophila, pointed and yan, are regulated by the Ras1/MAPK pathway. The pointed gene codes for two related proteins, and we show that one form is a constitutive activator of transcription, while the activity of the other form is stimulated by the Ras1/MAPK pathway. Mutation of the single consensus MAPK phosphorylation site in the second form abrogates this responsiveness. yan is a negative regulator of photoreceptor determination, and genetic data suggest that it acts as an antagonist of Ras1. We demonstrate that yan can repress transcription and that this repression activity is negatively regulated by the Ras1/MAPK signal, most likely through direct phosphorylation of yan by MAPK.
The argos gene encodes a protein that is required for viability and that regulates the determination of cells in the Drosophila eye. A developmental analysis of argos mutant eyes indicates that the mystery cells, which are usually nonneuronal, are transformed into extra photoreceptors, and that supernumerary cone cells and pigment cells are also recruited. Clonal analysis indicates that argos acts nonautonomously and can diffuse over the range of several cell diameters. Conceptual translation of the argos gene suggests that it encodes a secreted protein.
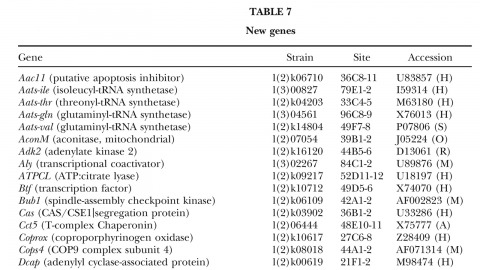
A fundamental goal of genetics and functional genomics is to identify and mutate every gene in model organisms such as Drosophila melanogaster. The Berkeley Drosophila Genome Project (BDGP) gene disruption project generates single P-element insertion strains that each mutate unique genomic open reading frames. Such strains strongly facilitate further genetic and molecular studies of the disrupted loci, but it has remained unclear if P elements can be used to mutate all Drosophila genes. We now report that the primary collection has grown to contain 1045 strains that disrupt more than 25% of the estimated 3600 Drosophila genes that are essential for adult viability. Of these P insertions, 67% have been verified by genetic tests to cause the associated recessive mutant phenotypes, and the validity of most of the remaining lines is predicted on statistical grounds. Sequences flanking >920 insertions have been determined to exactly position them in the genome and to identify 376 potentially affected transcripts from collections of EST sequences. Strains in the BDGP collection are available from the Bloomington Stock Center and have already assisted the research community in characterizing >250 Drosophila genes. The likely identity of 131 additional genes in the collection is reported here. Our results show that Drosophila genes have a wide range of sensitivity to inactivation by P elements, and provide a rationale for greatly expanding the BDGP primary collection based entirely on insertion site sequencing. We predict that this approach can bring >85% of all Drosophila open reading frames under experimental control.
Analysing computations in neural circuits often uses simplified models because the actual neuronal implementation is not known. For example, a problem in vision, how the eye detects image motion, has long been analysed using Hassenstein-Reichardt (HR) detector or Barlow-Levick (BL) models. These both simulate motion detection well, but the exact neuronal circuits undertaking these tasks remain elusive. We reconstructed a comprehensive connectome of the circuits of Drosophila's motion-sensing T4 cells using a novel EM technique. We uncover complex T4 inputs and reveal that putative excitatory inputs cluster at T4's dendrite shafts, while inhibitory inputs localize to the bases. Consistent with our previous study, we reveal that Mi1 and Tm3 cells provide most synaptic contacts onto T4. We are, however, unable to reproduce the spatial offset between these cells reported previously. Our comprehensive connectome reveals complex circuits that include candidate anatomical substrates for both HR and BL types of motion detectors.