Filter
Associated Lab
- Aso Lab (30) Apply Aso Lab filter
- Betzig Lab (1) Apply Betzig Lab filter
- Bock Lab (2) Apply Bock Lab filter
- Branson Lab (9) Apply Branson Lab filter
- Card Lab (5) Apply Card Lab filter
- Clapham Lab (1) Apply Clapham Lab filter
- Dickson Lab (2) Apply Dickson Lab filter
- Druckmann Lab (1) Apply Druckmann Lab filter
- Fetter Lab (1) Apply Fetter Lab filter
- Funke Lab (2) Apply Funke Lab filter
- Harris Lab (3) Apply Harris Lab filter
- Heberlein Lab (2) Apply Heberlein Lab filter
- Hermundstad Lab (2) Apply Hermundstad Lab filter
- Hess Lab (6) Apply Hess Lab filter
- Jayaraman Lab (7) Apply Jayaraman Lab filter
- Lippincott-Schwartz Lab (1) Apply Lippincott-Schwartz Lab filter
- Looger Lab (2) Apply Looger Lab filter
- O'Shea Lab (1) Apply O'Shea Lab filter
- Otopalik Lab (1) Apply Otopalik Lab filter
- Reiser Lab (17) Apply Reiser Lab filter
- Riddiford Lab (1) Apply Riddiford Lab filter
- Romani Lab (2) Apply Romani Lab filter
- Rubin Lab (149) Apply Rubin Lab filter
- Saalfeld Lab (5) Apply Saalfeld Lab filter
- Scheffer Lab (8) Apply Scheffer Lab filter
- Schreiter Lab (1) Apply Schreiter Lab filter
- Simpson Lab (3) Apply Simpson Lab filter
- Singer Lab (1) Apply Singer Lab filter
- Spruston Lab (1) Apply Spruston Lab filter
- Stern Lab (1) Apply Stern Lab filter
- Svoboda Lab (3) Apply Svoboda Lab filter
- Tjian Lab (1) Apply Tjian Lab filter
- Truman Lab (4) Apply Truman Lab filter
- Turaga Lab (1) Apply Turaga Lab filter
- Turner Lab (5) Apply Turner Lab filter
- Zuker Lab (1) Apply Zuker Lab filter
Associated Project Team
- CellMap (1) Apply CellMap filter
- Fly Functional Connectome (4) Apply Fly Functional Connectome filter
- Fly Olympiad (3) Apply Fly Olympiad filter
- FlyEM (12) Apply FlyEM filter
- FlyLight (20) Apply FlyLight filter
- GENIE (1) Apply GENIE filter
- Transcription Imaging (1) Apply Transcription Imaging filter
Publication Date
- 2026 (2) Apply 2026 filter
- 2025 (8) Apply 2025 filter
- 2024 (4) Apply 2024 filter
- 2023 (5) Apply 2023 filter
- 2022 (1) Apply 2022 filter
- 2021 (4) Apply 2021 filter
- 2020 (9) Apply 2020 filter
- 2019 (6) Apply 2019 filter
- 2018 (7) Apply 2018 filter
- 2017 (15) Apply 2017 filter
- 2016 (3) Apply 2016 filter
- 2015 (16) Apply 2015 filter
- 2014 (9) Apply 2014 filter
- 2013 (5) Apply 2013 filter
- 2012 (8) Apply 2012 filter
- 2011 (4) Apply 2011 filter
- 2010 (4) Apply 2010 filter
- 2009 (2) Apply 2009 filter
- 2008 (4) Apply 2008 filter
- 2007 (2) Apply 2007 filter
- 2006 (1) Apply 2006 filter
- 2002 (1) Apply 2002 filter
- 2000 (2) Apply 2000 filter
- 1999 (1) Apply 1999 filter
- 1997 (1) Apply 1997 filter
- 1995 (2) Apply 1995 filter
- 1994 (2) Apply 1994 filter
- 1993 (2) Apply 1993 filter
- 1992 (1) Apply 1992 filter
- 1991 (2) Apply 1991 filter
- 1990 (3) Apply 1990 filter
- 1989 (2) Apply 1989 filter
- 1987 (2) Apply 1987 filter
- 1986 (1) Apply 1986 filter
- 1985 (1) Apply 1985 filter
- 1984 (1) Apply 1984 filter
- 1983 (1) Apply 1983 filter
- 1982 (2) Apply 1982 filter
- 1981 (1) Apply 1981 filter
- 1979 (1) Apply 1979 filter
- 1973 (1) Apply 1973 filter
Type of Publication
149 Publications
Showing 121-130 of 149 resultsMaking inferences about the computations performed by neuronal circuits from synapse-level connectivity maps is an emerging opportunity in neuroscience. The mushroom body (MB) is well positioned for developing and testing such an approach due to its conserved neuronal architecture, recently completed dense connectome, and extensive prior experimental studies of its roles in learning, memory and activity regulation. Here we identify new components of the MB circuit in , including extensive visual input and MB output neurons (MBONs) with direct connections to descending neurons. We find unexpected structure in sensory inputs, in the transfer of information about different sensory modalities to MBONs, and in the modulation of that transfer by dopaminergic neurons (DANs). We provide insights into the circuitry used to integrate MB outputs, connectivity between the MB and the central complex and inputs to DANs, including feedback from MBONs. Our results provide a foundation for further theoretical and experimental work.
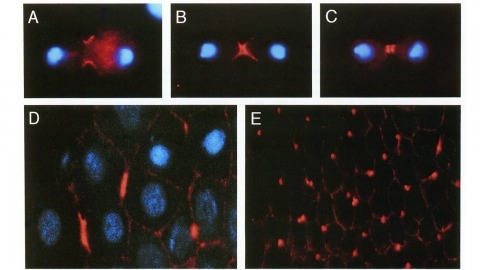
We have identified a Drosophila gene, peanut (pnut), that is related in sequence to the CDC3, CDC10, CDC11, and CDC12 genes of S. cerevisiae. These genes are required for cytokinesis, and their products are present at the bud neck during cell division. We find that pnut is also required for cytokinesis: in pnut mutants, imaginal tissues fail to proliferate and instead develop clusters of large, multinucleate cells. Pnut protein is localized to the cleavage furrow of dividing cells during cytokinesis and to the intercellular bridge connecting postmitotic daughter cells. In addition to its role in cytokinesis, pnut displays genetic interactions with seven in absentia, a gene required for neuronal fate determination in the compound eye, suggesting that pnut may have pleiotropic functions. Our results suggest that this class of proteins is involved in aspects of cytokinesis that have been conserved between flies and yeast.
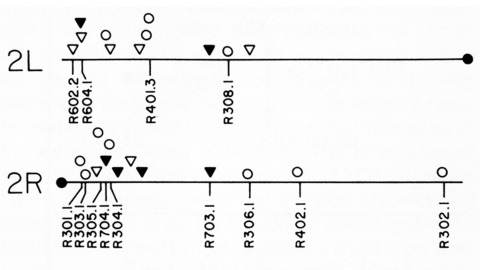
Thirty-six isogenic D. melanogaster strains that differed only in the chromosomal location of a 7.2 or an 8.1 kb DNA segment containing the (autosomal) rosy gene were constructed by P-element-mediated gene transfer. Since the flies were homozygous for a rosy- allele, rosy gene function in these indicated the influence of flanking sequences on gene expression. The tissue distribution of XDH activity in all the strains was normal. Each line exhibited a characteristic level of adult XDH-specific activity. The majority of these values were close to wild-type levels; however, the total variation in specific activity among the lines was nearly fivefold. Thus position effects influence expression of the rosy gene quantitatively but do not detectably alter tissue specificity. X-linked rosy insertions were expressed on average 1.6 times more activity in males than in females. Hence the gene acquires at least partial dosage compensation upon insertion into the X chromosome.
The analysis of genetic mosaics, in which an animal carries populations of cells with differing genotypes, is a powerful tool for understanding developmental and cell biology. In 1990, we set out to improve the methods used to make genetic mosaics in Drosophila by taking advantage of recently developed approaches for genome engineering. These efforts led to the work described in our 1993 Development paper.
The perception of visual motion is critical for animal navigation, and flies are a prominent model system for exploring this neural computation. In Drosophila, the T4 cells of the medulla are directionally selective and necessary for ON motion behavioral responses. To examine the emergence of directional selectivity, we developed genetic driver lines for the neuron types with the most synapses onto T4 cells. Using calcium imaging, we found that these neuron types are not directionally selective and that selectivity arises in the T4 dendrites. By silencing each input neuron type, we identified which neurons are necessary for T4 directional selectivity and ON motion behavioral responses. We then determined the sign of the connections between these neurons and T4 cells using neuronal photoactivation. Our results indicate a computational architecture for motion detection that is a hybrid of classic theoretical models.
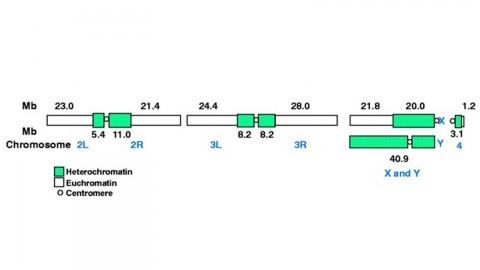
The fly Drosophila melanogaster is one of the most intensively studied organisms in biology and serves as a model system for the investigation of many developmental and cellular processes common to higher eukaryotes, including humans. We have determined the nucleotide sequence of nearly all of the approximately 120-megabase euchromatic portion of the Drosophila genome using a whole-genome shotgun sequencing strategy supported by extensive clone-based sequence and a high-quality bacterial artificial chromosome physical map. Efforts are under way to close the remaining gaps; however, the sequence is of sufficient accuracy and contiguity to be declared substantially complete and to support an initial analysis of genome structure and preliminary gene annotation and interpretation. The genome encodes approximately 13,600 genes, somewhat fewer than the smaller Caenorhabditis elegans genome, but with comparable functional diversity.
Null mutations of glass specifically remove photoreceptor cells, leaving other cell types intact. We have isolated the glass gene and have shown that its transcript encodes a putative protein of 604 amino acids with five zinc-fingers. The glass product may be a transcription factor required for the development of a single neuronal cell type.
Glia play crucial roles in the development and homeostasis of the nervous system. While the GLIA in the Drosophila embryo have been well characterized, their study in the adult nervous system has been limited. Here, we present a detailed description of the glia in the adult nervous system, based on the analysis of some 500 glial drivers we identified within a collection of synthetic GAL4 lines. We find that glia make up ∼10% of the cells in the nervous system and envelop all compartments of neurons (soma, dendrites, axons) as well as the nervous system as a whole. Our morphological analysis suggests a set of simple rules governing the morphogenesis of glia and their interactions with other cells. All glial subtypes minimize contact with their glial neighbors but maximize their contact with neurons and adapt their macromorphology and micromorphology to the neuronal entities they envelop. Finally, glial cells show no obvious spatial organization or registration with neuronal entities. Our detailed description of all glial subtypes and their regional specializations, together with the powerful genetic toolkit we provide, will facilitate the functional analysis of glia in the mature nervous system. GLIA 2017.
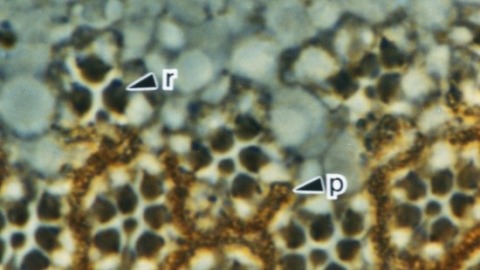
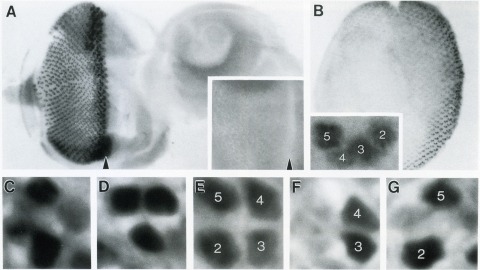
The Drosophila homeo box gene rough is required in photoreceptor cells R2 and R5 for normal eye development. We show here that rough protein expression is limited to a subset of cells in the developing retina where it is transiently expressed for 30-60 hr. The rough protein is first expressed broadly in the morphogenetic furrow but is rapidly restricted to the R2, R3, R4, and R5 precursor cells. Ubiquitous expression of rough under the control of the hsp70 promoter in third-instar larvae suppresses the initial steps of ommatidial assembly. Structures derived from other imaginal discs are not affected. Ectopic expression of rough in the R7 precursor, through the use of the sevenless promoter, causes this cell to develop into an R1-6 photoreceptor subtype; however, this cell still requires sevenless function for its neural differentiation. Taken together with previous analyses of the rough mutant phenotype, these results suggest that the normal role of rough is to establish the unique cell identity of photoreceptors R2 and R5.
Large scientific projects in genomics and astronomy are influential not because they answer any single question but because they enable investigation of continuously arising new questions from the same data-rich sources. Advances in automated mapping of the brain's synaptic connections (connectomics) suggest that the complicated circuits underlying brain function are ripe for analysis. We discuss benefits of mapping a mouse brain at the level of synapses.