neuPrint and NeuronBridge enable scientists worldwide to develop and test hypotheses about brain connections.
Annika Barber knew that 2020 was going to be a big year for her.
“I started my faculty position in January of 2020, the connectome of the fly hemibrain dropped that same month, and I was like, this is going to be a great year,” says Barber, a behavioral neurobiologist at Rutgers University. “It was not a great year.”
In March, the pandemic hit, and like researchers around the world, Barber and her team, who study the circadian clock in fruit flies, couldn’t go into their lab. But they did have access to the fly connectome and two key software tools developed at Janelia – neuPrint and NeuronBridge.
Using neuPrint, the team was able to virtually explore the wiring of the fly brain. Like using Google Maps™ to navigate a city, neuPrint allowed the researchers to chart paths through the brain to see where clock neurons, which control circadian rhythms, connect to non-clock neurons that send signals to other parts of the fly. This enabled the team to create a list of connections – down to the level of single neurons – that they could test once the lab reopened.
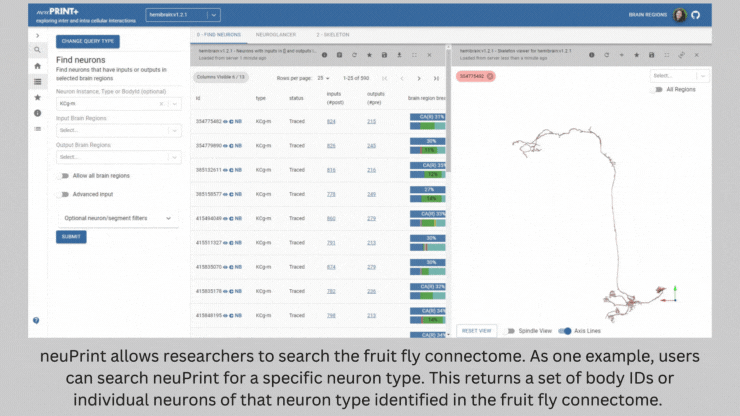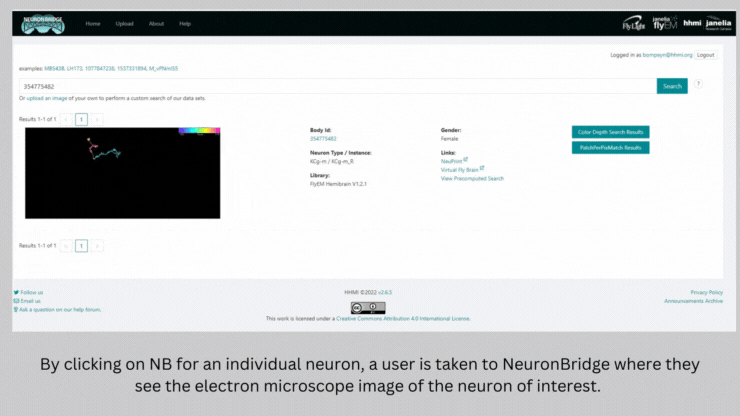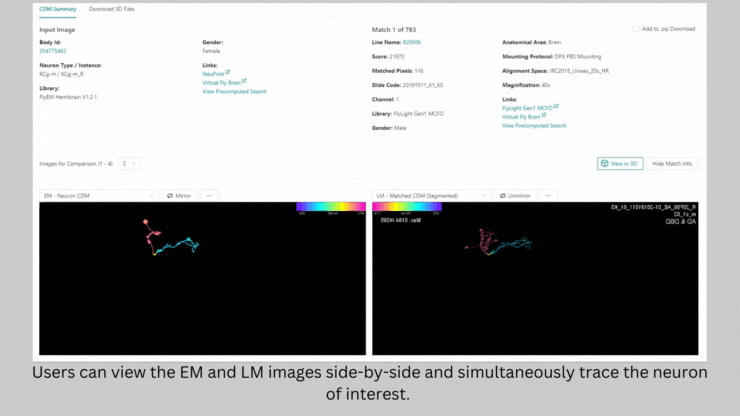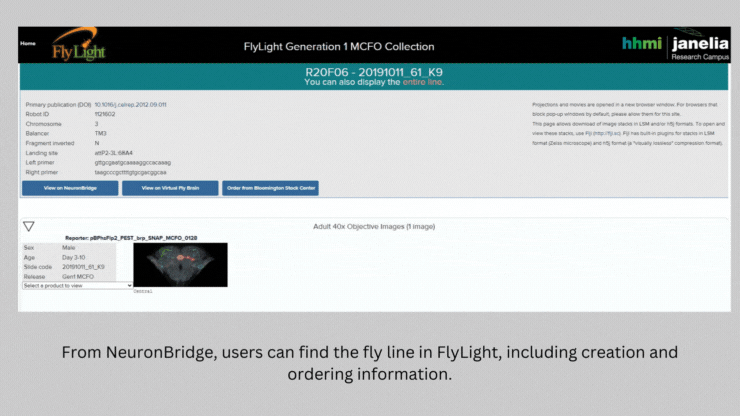
NeuronBridge helped Barber and her team select the genetically modified strains of fly they needed to test the connections they identified. The software serves as a bridge between two types of imaging – light microscopy and electron microscopy – that are used to image flies expressing markers in certain neurons. NeuronBridge searches across these two imaging modalities, enabling the researcher to locate specific cell types and home in on a fly strain to use in their experiments.
For Barber, a new PI, the software tools were critical for her ability to continue her research and apply for grants during the pandemic.
“They are just such a gift to the community,” Barber says of the tools. “It really is like giving us all a huge stack of preliminary data for free, which is just amazing.”
neuPrint and NeuronBridge were created by Janelia’s Scientific Computing team in collaboration with the FlyLight and FlyEM Project Teams. neuPrint allows researchers to use data the FlyEM Project Team generated while assembling the fly connectome. NeuronBridge gives scientists access to data from FlyLight, which produces data sets and collections of flies for visualization and manipulation of individual cell types in the fly brain. Together, these tools allow researchers like Barber to develop and test new ideas about connections in the fly brain from anywhere.
Creating easy-to-use products such as neuPrint and NeuronBridge that are freely available to the entire scientific community is part of Janelia’s commitment to open science. Such tools are also key to the success of Janelia’s Project Teams, which aim to tackle challenging, trans-disciplinary problems that can’t be accomplished in a traditional scientific environment.
For Project Teams to fulfill their goals and serve the scientific community, they need mechanisms for sharing their results in a usable form, allowing anyone with an internet connection to access them, says Janelia Senior Group Leader Gerry Rubin. That’s where Janelia’s Scientific Computing team comes in, building freely available, user-friendly software tools to complement the work of the Project Teams.
“It democratizes the data,” says Rubin, who started the Project Teams as the research campus’s first director.
Since they were released, neuPrint and NeuronBridge, which are fully integrated with each other, have been embraced by the fly research community. To date, neuPrint has 3,200 registered users. NeuronBridge had more than 1,000 registered users as of July 2022. Details about neuPrint can be found in a new paper in Frontiers in Neuroinformatics, while the construction and functionality of NeuronBridge is explained in a new preprint on bioRxiv.
“Open science is about making scientific research accessible to as many people as possible,” says Konrad Rokicki, manager of software engineering at Janelia. “The FlyEM connectome and FlyLight driver line resources are large and complex data sets and would not be accessible to nearly as many people without the tools needed to browse and search them.”
For Barber, who has been studying clock neurons in one area of the brain, the two tools and the data behind them are powerful, unbiased hypothesis generators. They have allowed her and her team – even those without extensive computation experience – to start looking at other areas of the brain to build a bigger, more comprehensive map of clock neurons.
Barber loves the tools so much she wrote a sonnet about them as part of pandemic project to read and tweet about scientific papers.
“It really exemplifies the kind of community that I’ve found in fly biology, where people are very open and very sharing of tools and reagents and data,” Barber says. “That helps us build a better scientific community, and I love being a part of a community that shares this way.”
###
Citation:
Stephen M. Plaza, Jody Clements, Tom Dolafi, Lowell Umayam, Nicole N. Neubarth, Louis K. Scheffer and Stuart Berg. “neuPrint: An open access tool for EM connectomics.” Frontiers in Neuroinformatics, published July 20, 2022. DOI: 10.3389/fninf.2022.896292
Jody Clements, Cristian Goina, Philip M. Hubbard, Takashi Kawase, Donald J. Olbris, Hideo Otsuna, Robert Svirskas and Konrad Rokicki. “NeuronBridge: an intuitive web application for neuronal morphology search across large data sets.” bioRxiv, posted July 21, 2022. DOI: 10.1101/2022.07.20.500311
Media Contacts
Nanci Bompey












