Filter
Associated Lab
- Ahrens Lab (5) Apply Ahrens Lab filter
- Aso Lab (3) Apply Aso Lab filter
- Betzig Lab (7) Apply Betzig Lab filter
- Bock Lab (5) Apply Bock Lab filter
- Branson Lab (3) Apply Branson Lab filter
- Card Lab (2) Apply Card Lab filter
- Cardona Lab (4) Apply Cardona Lab filter
- Clapham Lab (2) Apply Clapham Lab filter
- Darshan Lab (1) Apply Darshan Lab filter
- Dickson Lab (5) Apply Dickson Lab filter
- Druckmann Lab (3) Apply Druckmann Lab filter
- Dudman Lab (4) Apply Dudman Lab filter
- Espinosa Medina Lab (3) Apply Espinosa Medina Lab filter
- Feliciano Lab (1) Apply Feliciano Lab filter
- Fitzgerald Lab (2) Apply Fitzgerald Lab filter
- Funke Lab (1) Apply Funke Lab filter
- Gonen Lab (2) Apply Gonen Lab filter
- Grigorieff Lab (4) Apply Grigorieff Lab filter
- Harris Lab (4) Apply Harris Lab filter
- Heberlein Lab (2) Apply Heberlein Lab filter
- Hermundstad Lab (1) Apply Hermundstad Lab filter
- Hess Lab (5) Apply Hess Lab filter
- Jayaraman Lab (4) Apply Jayaraman Lab filter
- Ji Lab (1) Apply Ji Lab filter
- Johnson Lab (1) Apply Johnson Lab filter
- Keleman Lab (2) Apply Keleman Lab filter
- Keller Lab (6) Apply Keller Lab filter
- Lavis Lab (6) Apply Lavis Lab filter
- Lee (Albert) Lab (1) Apply Lee (Albert) Lab filter
- Lippincott-Schwartz Lab (12) Apply Lippincott-Schwartz Lab filter
- Liu (Zhe) Lab (7) Apply Liu (Zhe) Lab filter
- Looger Lab (15) Apply Looger Lab filter
- O'Shea Lab (1) Apply O'Shea Lab filter
- Pachitariu Lab (4) Apply Pachitariu Lab filter
- Pavlopoulos Lab (1) Apply Pavlopoulos Lab filter
- Podgorski Lab (4) Apply Podgorski Lab filter
- Reiser Lab (2) Apply Reiser Lab filter
- Romani Lab (3) Apply Romani Lab filter
- Rubin Lab (6) Apply Rubin Lab filter
- Saalfeld Lab (3) Apply Saalfeld Lab filter
- Scheffer Lab (2) Apply Scheffer Lab filter
- Schreiter Lab (4) Apply Schreiter Lab filter
- Simpson Lab (1) Apply Simpson Lab filter
- Singer Lab (4) Apply Singer Lab filter
- Spruston Lab (6) Apply Spruston Lab filter
- Stern Lab (5) Apply Stern Lab filter
- Sternson Lab (2) Apply Sternson Lab filter
- Stringer Lab (3) Apply Stringer Lab filter
- Svoboda Lab (14) Apply Svoboda Lab filter
- Tillberg Lab (2) Apply Tillberg Lab filter
- Truman Lab (4) Apply Truman Lab filter
- Turaga Lab (2) Apply Turaga Lab filter
- Turner Lab (2) Apply Turner Lab filter
- Zlatic Lab (1) Apply Zlatic Lab filter
Associated Project Team
Associated Support Team
- Cryo-Electron Microscopy (4) Apply Cryo-Electron Microscopy filter
- Electron Microscopy (4) Apply Electron Microscopy filter
- Fly Facility (5) Apply Fly Facility filter
- Gene Targeting and Transgenics (2) Apply Gene Targeting and Transgenics filter
- Integrative Imaging (1) Apply Integrative Imaging filter
- Janelia Experimental Technology (6) Apply Janelia Experimental Technology filter
- Molecular Genomics (1) Apply Molecular Genomics filter
- Primary & iPS Cell Culture (1) Apply Primary & iPS Cell Culture filter
- Project Technical Resources (4) Apply Project Technical Resources filter
- Quantitative Genomics (4) Apply Quantitative Genomics filter
- Scientific Computing Software (5) Apply Scientific Computing Software filter
- Scientific Computing Systems (1) Apply Scientific Computing Systems filter
Publication Date
- December 2019 (9) Apply December 2019 filter
- November 2019 (11) Apply November 2019 filter
- October 2019 (18) Apply October 2019 filter
- September 2019 (15) Apply September 2019 filter
- August 2019 (14) Apply August 2019 filter
- July 2019 (12) Apply July 2019 filter
- June 2019 (18) Apply June 2019 filter
- May 2019 (12) Apply May 2019 filter
- April 2019 (16) Apply April 2019 filter
- March 2019 (17) Apply March 2019 filter
- February 2019 (18) Apply February 2019 filter
- January 2019 (17) Apply January 2019 filter
- Remove 2019 filter 2019
177 Janelia Publications
Showing 171-177 of 177 resultsIn the current model of endothelial barrier regulation, the tyrosine kinase SRC is purported to induce disassembly of endothelial adherens junctions (AJs) via phosphorylation of VE cadherin, and thereby increase junctional permeability. Here, using a chemical biology approach to temporally control SRC activation, we show that SRC exerts distinct time-variant effects on the endothelial barrier. We discovered that the immediate effect of SRC activation was to transiently enhance endothelial barrier function as the result of accumulation of VE cadherin at AJs and formation of morphologically distinct reticular AJs. Endothelial barrier enhancement via SRC required phosphorylation of VE cadherin at Y731. In contrast, prolonged SRC activation induced VE cadherin phosphorylation at Y685, resulting in increased endothelial permeability. Thus, time-variant SRC activation differentially phosphorylates VE cadherin and shapes AJs to fine-tune endothelial barrier function. Our work demonstrates important advantages of synthetic biology tools in dissecting complex signaling systems.
We compared performance of recently developed silicon photomultipliers (SiPMs) to GaAsP photomultiplier tubes (PMTs) for two-photon imaging of neural activity. Despite higher dark counts, SiPMs match or exceed the signal-to-noise ratio of PMTs at photon rates encountered in typical calcium imaging experiments due to their low pulse height variability. At higher photon rates encountered during high-speed voltage imaging, SiPMs substantially outperform PMTs.
External cues, including touch, enable walking animals to flexibly maneuver around obstacles and extricate themselves from dead-ends (for reviews, see [1-3]). In a screen for neurons that enable Drosophila melanogaster to retreat when it encounters a dead-end, we identified a pair of ascending neurons, the TwoLumps Ascending (TLA) neurons. Silencing TLA activity impairs backward locomotion, whereas optogenetic activation triggers backward walking. TLA-induced reversal is mediated in part by the Moonwalker Descending Neurons (MDNs) [4], which receive excitatory input from the TLAs. Silencing the TLAs decreases the extent to which freely walking flies back up upon encountering a physical barrier in the dark, and TLAs show calcium responses to optogenetic activation of neurons expressing the mechanosensory channel NOMPC. We infer that TLAs convey feedforward mechanosensory stimuli to transiently activate MDNs in response to anterior body touch.
Chemogenetics enables non-invasive chemical control over cell populations in behaving animals. However, existing small molecule agonists show insufficient potency or selectivity. There is also need for chemogenetic systems compatible with both research and human therapeutic applications. We developed a new ion channel-based platform for cell activation and silencing that is controlled by low doses of the anti-smoking drug varenicline. We then synthesized novel sub-nanomolar potency agonists, called uPSEMs, with high selectivity for the chemogenetic receptors. uPSEMs and their receptors were characterized in brains of mice and a rhesus monkey by in vivo electrophysiology, calcium imaging, positron emission tomography, behavioral efficacy testing, and receptor counterscreening. This platform of receptors and selective ultrapotent agonists enables potential research and clinical applications of chemogenetics.
Gaining independent genetic access to discrete cell types is critical to interrogate their biological functions as well as to deliver precise gene therapy. Transcriptomics has allowed us to profile cell populations with extraordinary precision, revealing that cell types are typically defined by a unique combination of genetic markers. Given the lack of adequate tools to target cell types based on multiple markers, most cell types remain inaccessible to genetic manipulation. Here we present CaSSA, a platform to create unlimited genetic switches based on CRISPR/Cas9 (Ca) and the DNA repair mechanism known as single-strand annealing (SSA). CaSSA allows engineering of independent genetic switches, each responding to a specific gRNA. Expressing multiple gRNAs in specific patterns enables multiplex cell-type-specific manipulations and combinatorial genetic targeting. CaSSA is a new genetic tool that conceptually works as an unlimited number of recombinases and will facilitate genetic access to cell types in diverse organisms.
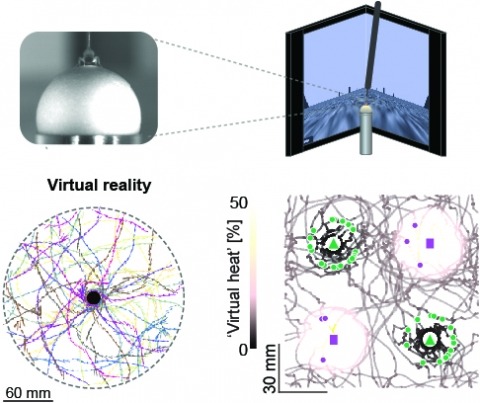
Studying the intertwined roles of sensation, experience, and directed action in navigation has been facilitated by the development of virtual reality (VR) environments for head-fixed animals, allowing for quantitative measurements of behavior in well-controlled conditions. VR has long featured in studies of Drosophila melanogaster, but these experiments have typically allowed the fly to change only its heading in a visual scene and not its position. Here we explore how flies move in two dimensions (2D) using a visual VR environment that more closely captures an animal's experience during free behavior. We show that flies' 2D interaction with landmarks cannot be automatically derived from their orienting behavior under simpler one-dimensional (1D) conditions. Using novel paradigms, we then demonstrate that flies in 2D VR adapt their behavior in response to optogenetically delivered appetitive and aversive stimuli. Much like free-walking flies after encounters with food, head-fixed flies exploring a 2D VR respond to optogenetic activation of sugar-sensing neurons by initiating a local search, which appears not to rely on visual landmarks. Visual landmarks can, however, help flies to avoid areas in VR where they experience an aversive, optogenetically generated heat stimulus. By coupling aversive virtual heat to the flies' presence near visual landmarks of specific shapes, we elicit selective learned avoidance of those landmarks. Thus, we demonstrate that head-fixed flies adaptively navigate in 2D virtual environments, but their reliance on visual landmarks is context dependent. These behavioral paradigms set the stage for interrogation of the fly brain circuitry underlying flexible navigation in complex multisensory environments.
Brains are notoriously hard to understand, and neuroscientists need all the tools they can get their hands on to have a realistic shot at it. Advances in machine learning are proving instrumental, illustrated by their recent use to shed light on navigational strategies implemented by zebrafish brains.