Filter
Associated Lab
- Betzig Lab (2) Apply Betzig Lab filter
- Bock Lab (1) Apply Bock Lab filter
- Cardona Lab (1) Apply Cardona Lab filter
- Dickson Lab (3) Apply Dickson Lab filter
- Druckmann Lab (1) Apply Druckmann Lab filter
- Dudman Lab (3) Apply Dudman Lab filter
- Eddy/Rivas Lab (2) Apply Eddy/Rivas Lab filter
- Egnor Lab (1) Apply Egnor Lab filter
- Fetter Lab (2) Apply Fetter Lab filter
- Fitzgerald Lab (2) Apply Fitzgerald Lab filter
- Gonen Lab (2) Apply Gonen Lab filter
- Heberlein Lab (3) Apply Heberlein Lab filter
- Hess Lab (1) Apply Hess Lab filter
- Jayaraman Lab (1) Apply Jayaraman Lab filter
- Kainmueller Lab (1) Apply Kainmueller Lab filter
- Karpova Lab (1) Apply Karpova Lab filter
- Keleman Lab (2) Apply Keleman Lab filter
- Keller Lab (1) Apply Keller Lab filter
- Koay Lab (1) Apply Koay Lab filter
- Lavis Lab (3) Apply Lavis Lab filter
- Lippincott-Schwartz Lab (1) Apply Lippincott-Schwartz Lab filter
- Magee Lab (1) Apply Magee Lab filter
- Pavlopoulos Lab (2) Apply Pavlopoulos Lab filter
- Reiser Lab (2) Apply Reiser Lab filter
- Riddiford Lab (2) Apply Riddiford Lab filter
- Romani Lab (1) Apply Romani Lab filter
- Rubin Lab (2) Apply Rubin Lab filter
- Schreiter Lab (3) Apply Schreiter Lab filter
- Sgro Lab (1) Apply Sgro Lab filter
- Shroff Lab (1) Apply Shroff Lab filter
- Simpson Lab (1) Apply Simpson Lab filter
- Spruston Lab (4) Apply Spruston Lab filter
- Stern Lab (9) Apply Stern Lab filter
- Svoboda Lab (3) Apply Svoboda Lab filter
- Tervo Lab (1) Apply Tervo Lab filter
- Tjian Lab (5) Apply Tjian Lab filter
- Truman Lab (1) Apply Truman Lab filter
- Turaga Lab (1) Apply Turaga Lab filter
Publication Date
- December 2007 (12) Apply December 2007 filter
- November 2007 (9) Apply November 2007 filter
- October 2007 (12) Apply October 2007 filter
- September 2007 (8) Apply September 2007 filter
- August 2007 (7) Apply August 2007 filter
- July 2007 (16) Apply July 2007 filter
- June 2007 (5) Apply June 2007 filter
- May 2007 (7) Apply May 2007 filter
- April 2007 (8) Apply April 2007 filter
- March 2007 (7) Apply March 2007 filter
- February 2007 (3) Apply February 2007 filter
- January 2007 (12) Apply January 2007 filter
- Remove 2007 filter 2007
Type of Publication
106 Publications
Showing 31-40 of 106 resultsThe Methoprene-tolerant (Met) gene in Drosophila melanogaster has been shown to function in juvenile hormone (JH) action. Met homologs were isolated from three mosquito species, Culex pipiens, Aedes aegypti and Anopheles gambiae. Sequence similarity was found to be high in bHLH and PAS conserved domains, and the majority of the 7-9 introns in AaMet and AgMet are located in either identical or similar positions, indicating evolutionary relatedness. Sequence comparison with Met and the similar germ-cell expressed (gce) gene in D. melanogaster showed that the mosquito genes are more similar to gce than to Met. Moreover, the multiple introns in AgMet and AaMet are more similar in number with the 7 introns in Dmgce than to the single intron in DmMet; in fact, six intron positions in AaMet and AgMet are similar to those in Dmgce. Efforts to identify a second homologous gene in mosquitoes were unsuccessful, suggesting a single gene in lower Diptera, consistent with the single gene uncovered in genomic sequencing of Ae. aegypti and An. gambiae. These results suggest that a gene duplication occurred during the evolution of higher Diptera, resulting in Met and gce.
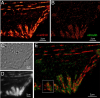
Accurate determination of the relative positions of proteins within localized regions of the cell is essential for understanding their biological function. Although fluorescent fusion proteins are targeted with molecular precision, the position of these genetically expressed reporters is usually known only to the resolution of conventional optics ( approximately 200 nm). Here, we report the use of two-color photoactivated localization microscopy (PALM) to determine the ultrastructural relationship between different proteins fused to spectrally distinct photoactivatable fluorescent proteins (PA-FPs). The nonperturbative incorporation of these endogenous tags facilitates an imaging resolution in whole, fixed cells of approximately 20-30 nm at acquisition times of 5-30 min. We apply the technique to image different pairs of proteins assembled in adhesion complexes, the central attachment points between the cytoskeleton and the substrate in migrating cells. For several pairs, we find that proteins that seem colocalized when viewed by conventional optics are resolved as distinct interlocking nano-aggregates when imaged via PALM. The simplicity, minimal invasiveness, resolution, and speed of the technique all suggest its potential to directly visualize molecular interactions within cellular structures at the nanometer scale.
Commentary: Identifies the photoactivatable fluorescent proteins (PA-FPs) Dronpa and PS-CFP2 as green partners to orange-red PA-FPs such as Kaede and Eos for dual color PALM imaging. Very low crosstalk is demonstrated between the two color channels. Furthermore, since the probes are genetically expressed, they are closely bound to their target proteins and exhibit zero non-specific background. All these properties are essential to unambiguously identify regions of co-localization or separate compartmentalization at the nanoscale, as demonstrated in the examples here.
Optomotor flight control in houseflies shows bandwidth fractionation such that steering responses to an oscillating large-field rotating panorama peak at low frequency, whereas responses to small-field objects peak at high frequency. In fruit flies, steady-state large-field translation generates steering responses that are three times larger than large-field rotation. Here, we examine the optomotor steering reactions to dynamically oscillating visual stimuli consisting of large-field rotation, large-field expansion, and small-field motion. The results show that, like in larger flies, large-field optomotor steering responses peak at low frequency, whereas small-field responses persist under high frequency conditions. However, in fruit flies large-field expansion elicits higher magnitude and tighter phase-locked optomotor responses than rotation throughout the frequency spectrum, which may suggest a further segregation within the large-field pathway. An analysis of wing beat frequency and amplitude reveals that mechanical power output during flight varies according to the spatial organization and motion dynamics of the visual scene. These results suggest that, like in larger flies, the optomotor control system is organized into parallel large-field and small-field pathways, and extends previous analyses to quantify expansion-sensitivity for steering reflexes and flight power output across the frequency spectrum.
Genetically encoded optical indicators hold the promise of enabling non-invasive monitoring of activity in identified neurons in behaving organisms. However, the interpretation of images of brain activity produced using such sensors is not straightforward. Several recent studies of sensory coding used G-CaMP 1.3-a calcium sensor-as an indicator of neural activity; some of these studies characterized the imaged neurons as having narrow tuning curves, a conclusion not always supported by parallel electrophysiological studies. To better understand the possible cause of these conflicting results, we performed simultaneous in vivo 2-photon imaging and electrophysiological recording of G-CaMP 1.3 expressing neurons in the antennal lobe (AL) of intact fruitflies. We find that G-CaMP has a relatively high threshold, that its signal often fails to capture spiking response kinetics, and that it can miss even high instantaneous rates of activity if those are not sustained. While G-CaMP can be misleading, it is clearly useful for the identification of promising neural targets: when electrical activity is well above the sensor’s detection threshold, its signal is fairly well correlated with mean firing rate and G-CaMP does not appear to alter significantly the responses of neurons that express it. The methods we present should enable any genetically encoded sensor, activator, or silencer to be evaluated in an intact neural circuit in vivo in Drosophila.
Comparative analysis of multiple genomes in a phylogenetic framework dramatically improves the precision and sensitivity of evolutionary inference, producing more robust results than single-genome analyses can provide. The genomes of 12 Drosophila species, ten of which are presented here for the first time (sechellia, simulans, yakuba, erecta, ananassae, persimilis, willistoni, mojavensis, virilis and grimshawi), illustrate how rates and patterns of sequence divergence across taxa can illuminate evolutionary processes on a genomic scale. These genome sequences augment the formidable genetic tools that have made Drosophila melanogaster a pre-eminent model for animal genetics, and will further catalyse fundamental research on mechanisms of development, cell biology, genetics, disease, neurobiology, behaviour, physiology and evolution. Despite remarkable similarities among these Drosophila species, we identified many putatively non-neutral changes in protein-coding genes, non-coding RNA genes, and cis-regulatory regions. These may prove to underlie differences in the ecology and behaviour of these diverse species.
On August 1, 2006 the Howard Hughes Medical Institute's first stand-alone research campus opened at Janelia Farm, near Washington DC. Our mission at Janelia is to do exceptional fundamental research. Our two scientific foci are to understand the function of neural circuits and to develop synergistic imaging technologies. To achieve this we have changed many of the conventions of academic and/or industrial science. The founding director at Janelia is the well-known Drosophilist Gerry Rubin, who has been a central figure in fly molecular, developmental and genomic biology in recent decades. Not coincidentally, we at Janelia fully appreciate the potential of flies to contribute to an understanding of neuronal circuits. Our objectives are ambitious, and in the first ten months of operations at Janelia we have made some good beginnings.
Both long-term behavioral memory and synaptic plasticity require protein synthesis, some of which may occur locally at specific synapses. Cytoplasmic polyadenylation element-binding (CPEB) proteins are thought to contribute to the local protein synthesis that underlies long-term changes in synaptic efficacy, but a role has not been established for them in the formation of long-term behavioral memory. We found that the Drosophila melanogaster CPEB protein Orb2 is acutely required for long-term conditioning of male courtship behavior. Deletion of the N-terminal glutamine-rich region of Orb2 resulted in flies that were impaired in their ability to form long-term, but not short-term, memory. Memory was restored by expressing Orb2 selectively in fruitless (fru)-positive gamma neurons of the mushroom bodies and by providing Orb2 function in mushroom bodies only during and shortly after training. Our data thus demonstrate that a CPEB protein is important in long-term memory and map the molecular, spatial and temporal requirements for its function in memory formation.
The pea aphid, Acyrthosiphon pisum, exhibits several environmentally cued, discrete, alternate phenotypes (polyphenisms) during its life cycle. In the wing polyphenism, female progeny develop as either winged or unwinged depending on the extent of crowding or host plant quality experienced by the mother. Males also have the ability to develop as either winged or unwinged, but this is genetically determined by a single locus on the X chromosome and is thus referred to as a wing polymorphism. In order to gain insight into the patterns of gene expression that underlie the wing polyphenism and polymorphism we have used a pea aphid cDNA microarray to examine gene expression in winged and unwinged females and males. Results suggest that winged and unwinged morphs exhibit systemic differences in gene expression and that many of these differences are shared between the wing polyphenism and polymorphism (i.e., between females and males). In addition, adult winged and unwinged males exhibit pronounced differences when compared to adult females and fourth instar males, as well as to each other.