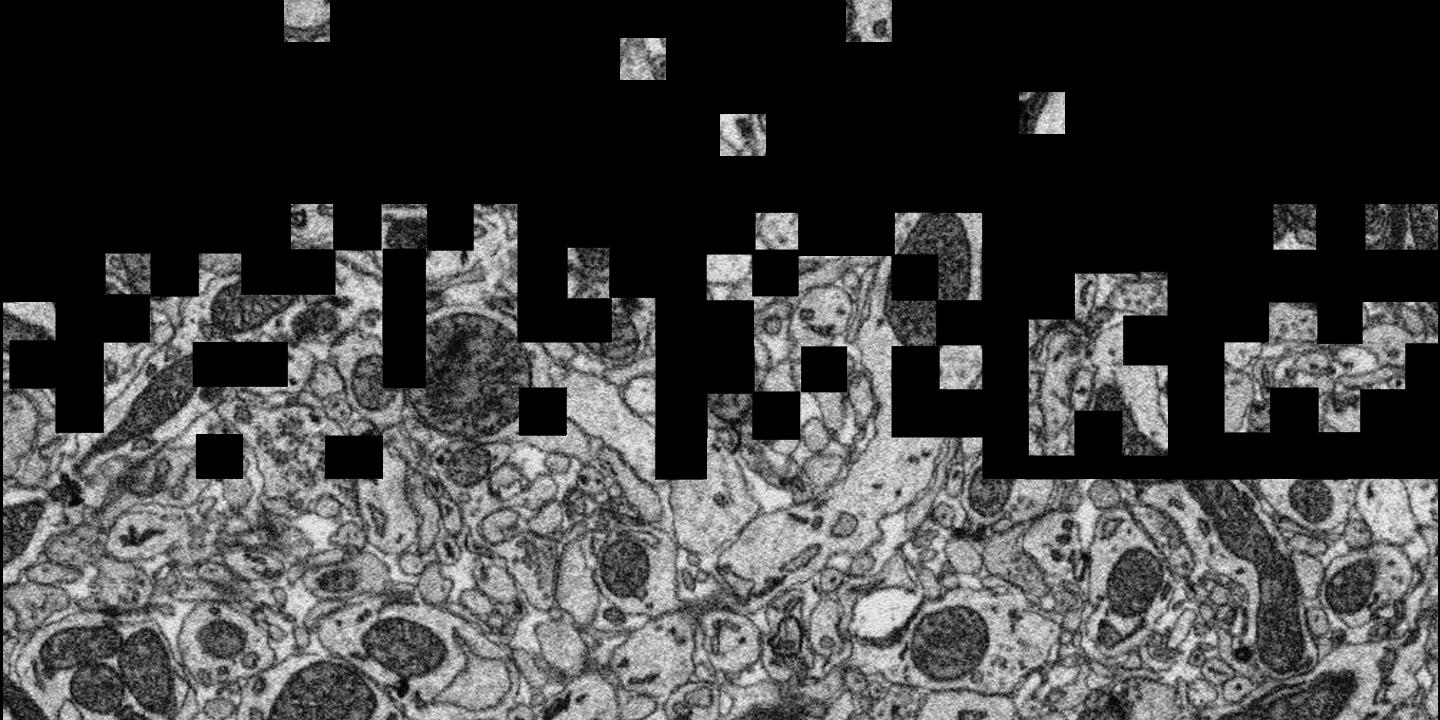
Briefly, serial section tomography entails slicing a block of tissue into many ultra-thin sections, and then tiling each slice with many overlapping microscope pictures. A serial section aligner is a software process for stitching back together all of those images into a seamless digital representation of the entire 3-D sample. It often takes many millions of individual pictures to cover just a small fraction of a brain. With very large numbers of images, inevitably some are lost, damaged or otherwise distorted making the reassembly process quite challenging. It's also a very large problem best solved on supercomputers such as Janelia's.
APIG undertook its alignment software effort as an adjunct to our Optical Array Tomography Tools project, for which it was decided that we should create maximum utility for our client/customers by creating the means to acquire high quality serial section data, and deliver the result as a fully aligned optical stack. The best available starting point was an EM aligner written here by Louis Scheffer, an excellent solution to a one-off problem Lou had been working on, not immediately applicable to other data.
Bill Karsh’s efforts over the last four years have been to produce a general purpose aligner for serial TEM, block-face SEM and optical data sets, generally anything we might produce here at Janelia. The result is one of the fastest and largest scale aligners in existence, capable of aligning modest data sets in hours and billions of images in just days. This is currently the de facto aligner at Janelia. It resides in the public domain (link here: https://github.com/billkarsh), and has been used at several other academic institutions (Harvard, Max Planck).
This aligner is now serving as a key building block for a new project within Janelia’s Scientific Computing group called 'The Stitcher'. That effort will integrate alignment with a larger workflow including database management, visualization tools and automated neuron tracing.
