by Stephen Plaza, Manager of FlyEM & Connectome Research Team Leader
July 10, 2019
Before recent advances in automated segmentation, comprehensively or densely reconstructing all neurons and (most) connections in a dataset was infeasible except for relatively small datasets. Instead, biologists sparsely traced the neurons most relevant to their work. But sparse is not enough.
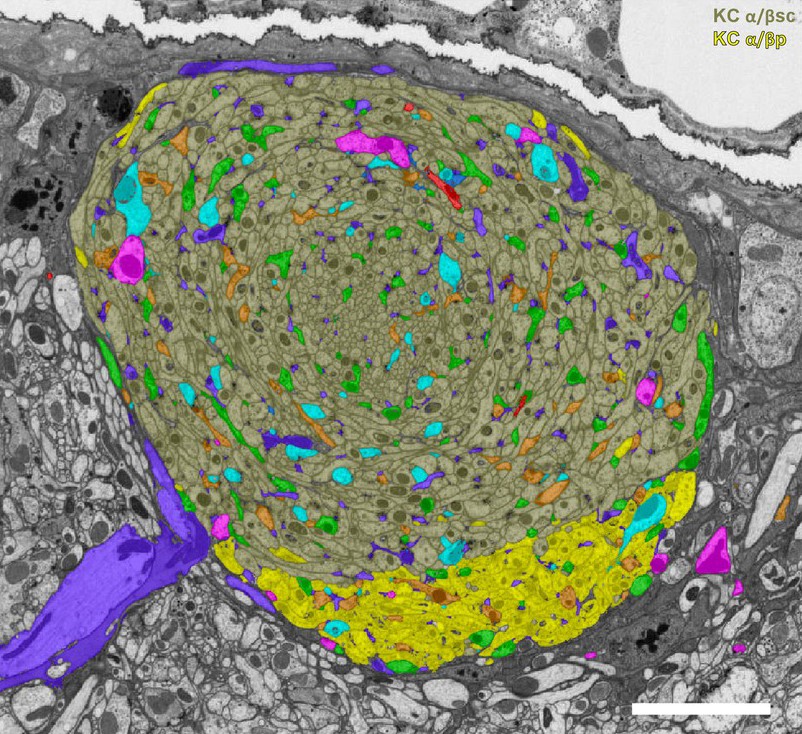
In our current hemibrain dataset, it is often expedient to find a neuron and trace its partners. This expediency can motivate a sparse-centric tracing philosophy, whereby connectomics is treated as an on-demand byproduct of explicit human inquiry, biased by one's preconceived hypotheses. In practice, we find that the scientists’ desire for information rarely stops at the local connectivity neighborhood. With the ability to trace neurons in minutes, the thirst for more information is great. Given the often small world properties connectomes can display, it is not long before the scientist requires thousands of neurons to answer an initially simple question (in fact we even find neurons with 1000 different direct partners). In addition, in some cases, the relevant questions demand analysis of system-wide connectivity patterns.
While the complexity of the brain wiring highlights the limits of narrow analysis, we also observe that even a small amount of tracing is an impediment for some scientists. Some EM purists might argue that this is the “barrier of entry” is necessary to truly appreciate this data. Indeed, it is often still very valuable when analyzing data. But, in practice, it hinders exploration of the data by a more diverse and broader mindset. We have many collaborators who only want to look at a summarized matrix of the data (which in turn makes faithful data representation a major challenge).
To be fair, a dense connectome, even with technological improvements, still requires extensive proofreading and logistics. Also, notably, not every brain region will motivate biological inquiry meaning some proofreading must come from full-time proofreaders, requiring different sources of motivation.
To mitigate the proofreading required, we re-define (or clarify) the definition of a dense connectome as one where all neuron morphologies are extracted and their connections are traced to ‘some level’ of accuracy. ‘Some level’ provides a dial whereby we trade off completeness and effort. We observe that the size distribution of automatically predicted segments resembles a long tail distribution, whereby there is a disproportionately small number of segments encompassing most of the dataset. We can systematically proofread these bigger segments, ideally identifying neural pathways with many connections. Furthermore, we note several properties to help ensure reconstruction accuracy, such as each neuron requires a nucleus and small segments should be merged to something bigger.
The advantages of top-down proofreading is that many scientists will still find the less complete reconstruction very useful, and as technology gets better, we can produce more complete results faster. Also, presumably, a centralized and systematic tracing effort can ensure more consistency and quality control in reconstruction than possible with just supporting sparse tracing efforts. Though, in practice we can still support sparse tracing, with careful management, in conjunction with our dense tracing efforts for the hemibrain. Having confidence in our error intervals and the technical and management logistics required to produce a dense connectome is a subject of many future blog entries.
